Cellular communication Biomolecular Processes as Concurrent Computation Aviv

Cellular communication: Biomolecular Processes as Concurrent Computation Aviv Regev March 2000
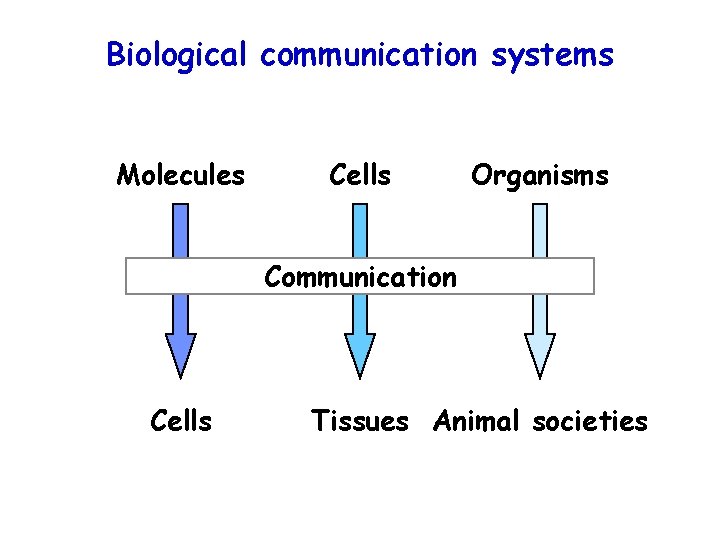
Biological communication systems Molecules Cells Organisms Communication Cells Tissues Animal societies
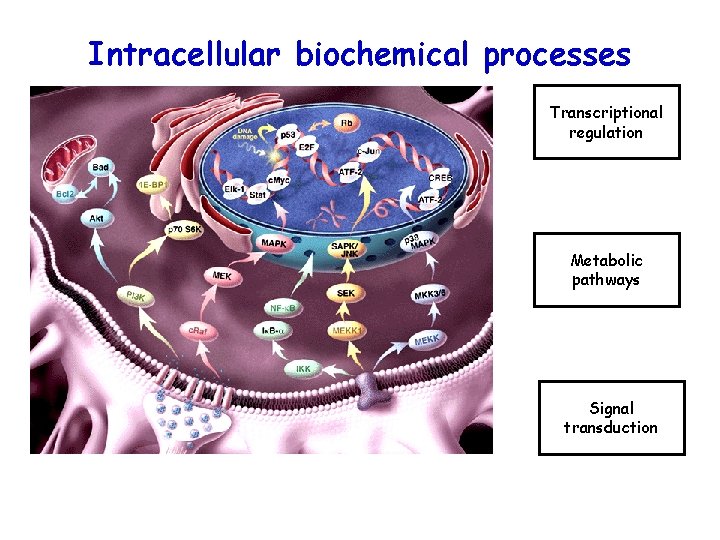
Intracellular biochemical processes Transcriptional regulation Metabolic pathways Signal transduction
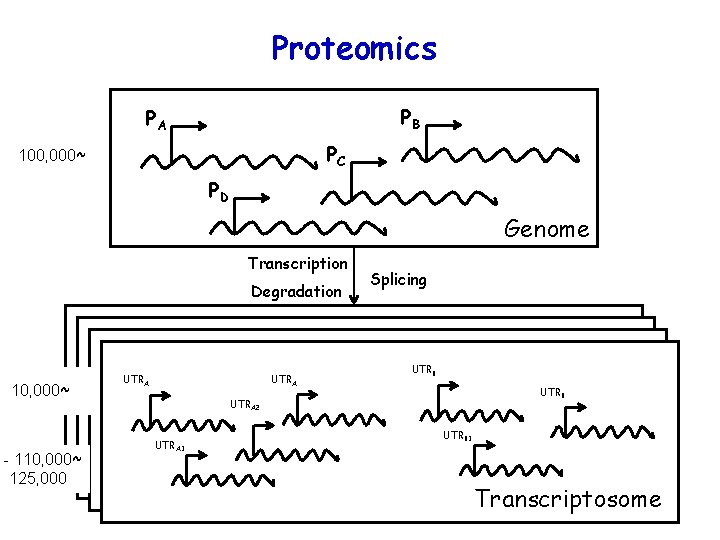
Proteomics PB PA PC 100, 000~ PD Genome Transcription Degradation 10, 000~ - 110, 000~ 125, 000 UTRA Splicing UTRB UTRA 2 UTRA 1 UTRB 1 Transcriptosome
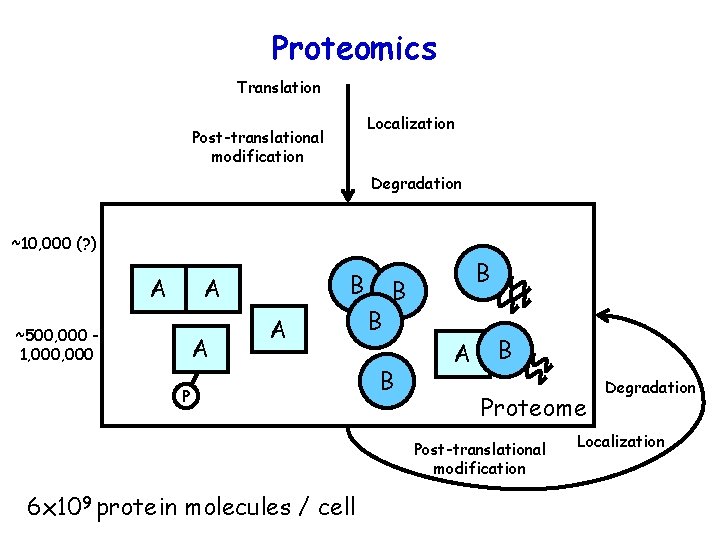
Proteomics Translation Localization Post-translational modification Degradation ~10, 000 (? ) A B A ~500, 000 1, 000 A A P B B A B Proteome Post-translational modification 6 x 109 protein molecules / cell Degradation Localization

Signal transduction (ST) pathways Pathways of molecular interactions that provide communication between the cell membrane and intracellular end-points, leading to some change in the cell.
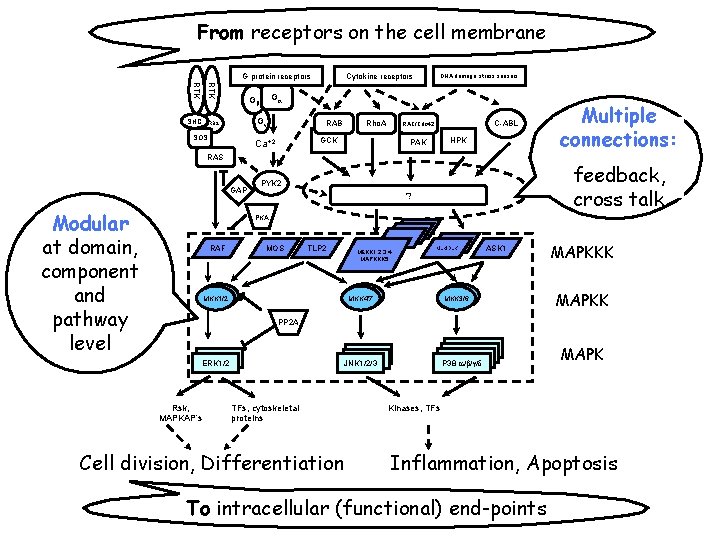
From receptors on the cell membrane G protein receptors RTK SHC Gg SOS DNA damage, stress sensors Ga Gb GRB 2 Cytokine receptors RAB Rho. A GCK Ca+2 C-ABL RAC/Cdc 42 HPK PAK RAS GAP Modular at domain, component and pathway level Multiple connections: feedback, cross talk PYK 2 ? PKA RAF MOS TLP 2 MKK 1/2 MEKK 1, 2, 3, 4 MAPKKK 5 MLK/DLK MKK 4/7 ASK 1 MKK 3/6 MAPKKK MAPKK PP 2 A ERK 1/2 Rsk, MAPKAP’s P 38 a/b/g/d JNK 1/2/3 TFs, cytoskeletal proteins Cell division, Differentiation MAPK Kinases, TFs Inflammation, Apoptosis To intracellular (functional) end-points
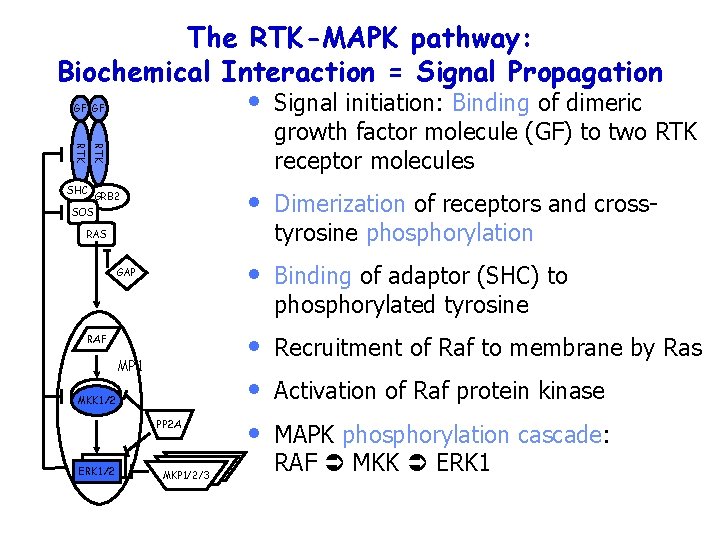
The RTK-MAPK pathway: Biochemical Interaction = Signal Propagation GF GF Signal initiation: Binding of dimeric growth factor molecule (GF) to two RTK receptor molecules • Dimerization of receptors and crosstyrosine phosphorylation • Binding of adaptor (SHC) to phosphorylated tyrosine • • • Recruitment of Raf to membrane by Ras RTK • SHC GRB 2 SOS RAS GAP RAF MP 1 MKK 1/2 PP 2 A ERK 1/2 MKP 1/2/3 Activation of Raf protein kinase MAPK phosphorylation cascade: RAF MKK ERK 1

What is missing from the picture? Information about § Dynamics Formal semantics The Power to § simulate § Molecular structure § analyze § Biochemical detail of interaction § compare Script: Characters +Plot Movie

Outline • Our approach: ST as concurrent computation • Process algebra: The p-calculus • Principles of modeling ST in p-calculus (characters) • Benefits of the approach: § full modeling (plot) § simulation (movie) § comparative analysis (the homology of process)

Our approach • Goal: Find an appropriate model for § molecular structure (characters) § and behavior (plot) § within a formal semantics (movie) • Computer Science analogy: Process algebra as a formalism for modeling of distributed computer systems

Our approach: Biological processes as concurrent computation • We suggest § The molecule as a computational process § Biochemical interaction as communication § Use process algebra to model ST • Benefits § Unified view § Simulation and analysis § Comparative power and scalability

The molecule as a computational process • Represent a structure by its potential behavior = by the process in which it can participate • Example: An enzyme (protein molecule) as the enzymatic reaction process, in which it may participate
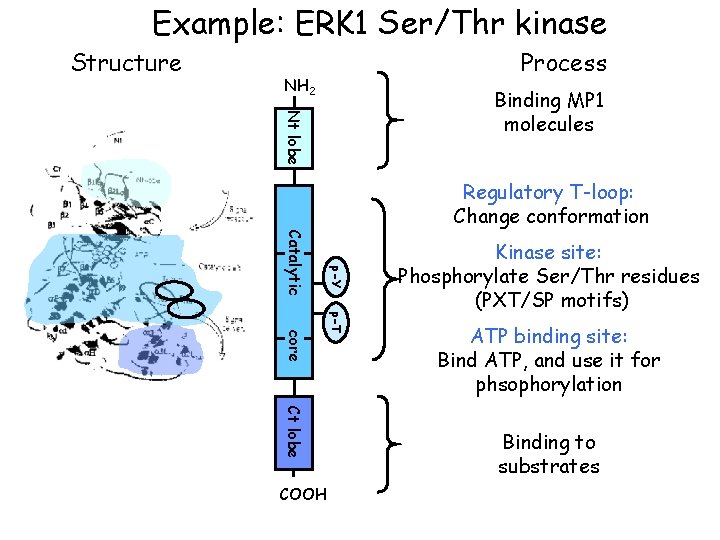
Example: ERK 1 Ser/Thr kinase Structure Process NH 2 Nt lobe Binding MP 1 molecules Regulatory T-loop: Change conformation p-Y Catalytic p-T core Ct lobe COOH Kinase site: Phosphorylate Ser/Thr residues (PXT/SP motifs) ATP binding site: Bind ATP, and use it for phsophorylation Binding to substrates

Interaction as communication • Each interaction enables or disables other interactions • Example: § Proteins A, B, and C § Proteins A and B interact § Protein A phosphorylates a residue on B § Protein C can bind only to the phosphorylated protein B
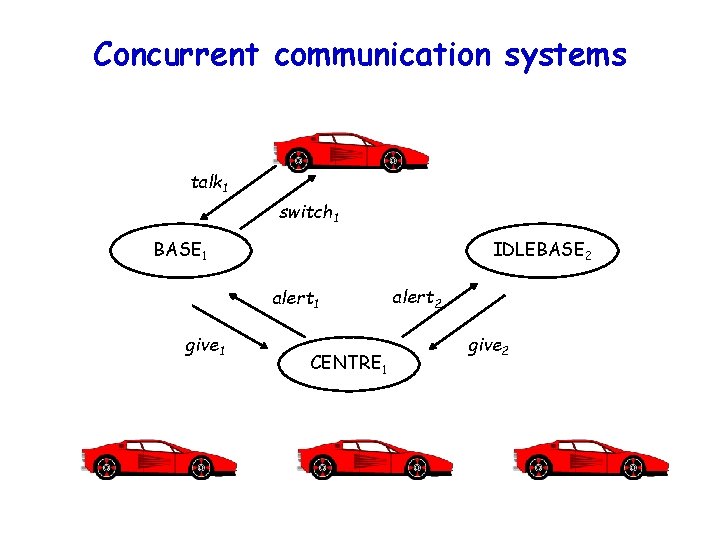
Concurrent communication systems talk 1 switch 1 BASE 1 IDLEBASE 2 alert 1 give 1 CENTRE 1 alert 2 give 2

ST as concurrent computation

An example • • A system: Proteins A, B, and C • Message: Protein A phosphorylates a residue on B • Meaning of message: This enables Protein B to bind to C Communication: Protein A and B can interact

Process algebras (calculi) Small formal languages capable of expressing the essential mechanism of concurrent computation

The p-calculus )Milner, Walker and Parrow( • • A community of interacting processes • Communication occurs via channels, defined by names • Communication content: Change of channel names (mobility) Processes are defined by their potential communication activities

The p-calculus: Formal structure • Syntax • Congruence laws • Reaction rules How to formally write a specification? When are two specifications the same? How does communication occur?

Syntax: Channels All communication events, input or output, occur on channels

Syntax: Processes are composed of communication events and of other processes

Principles for mapping ST to p-calculus ERK 1 Domain = Process SYSTEM : : = ERK 1 | … ERK 1 : : = (new internal_channels) (Nt_LOBE |CATALYTIC_LOBE |Ct_LOBE) Residues = Global (free) channel names and co-names Y T_LOOP (tyr ): : = tyr ? (tyr’ ). PHOSPH_SITE(tyr’)

The p-calculus: Reduction rules COMM: Ready to send z on x Ready to receive y on x Actions consumed; Alternative choices discarded + … )x ! z. Q ) | (… + x ? y. P) Q | P {z/y} z replaces y in P
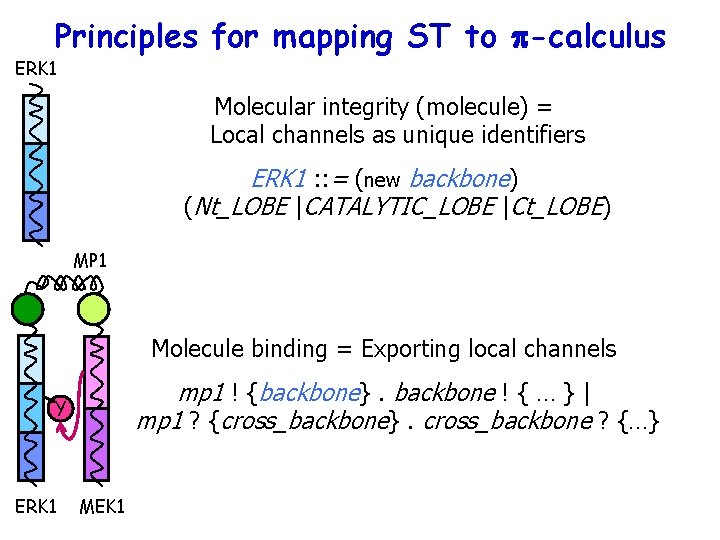
Principles for mapping ST to p-calculus ERK 1 Molecular integrity (molecule) = Local channels as unique identifiers ERK 1 : : = (new backbone) (Nt_LOBE |CATALYTIC_LOBE |Ct_LOBE) MP 1 Molecule binding = Exporting local channels mp 1 ! {backbone}. backbone ! { … } | mp 1 ? {cross_backbone}. cross_backbone ? {…} Y ERK 1 MEK 1
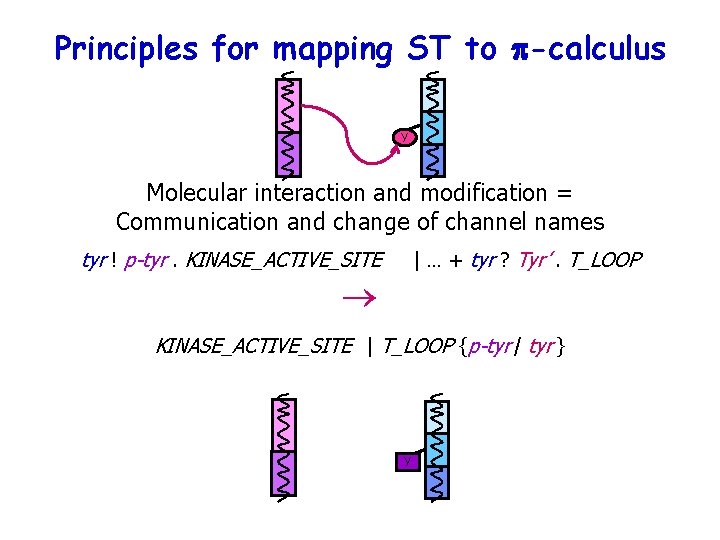
Principles for mapping ST to p-calculus Y Molecular interaction and modification = Communication and change of channel names tyr ! p-tyr. KINASE_ACTIVE_SITE | … + tyr ? Tyr’. T_LOOP ® KINASE_ACTIVE_SITE | T_LOOP {p-tyr / tyr } Y

Results: Unified view of structure and dynamics • Detailed molecular information (complexes, molecules, domains, residues) in visible form • Complex dynamic behavior (feedback, cross -talk, split and merge) without explicit modeling • Modular system
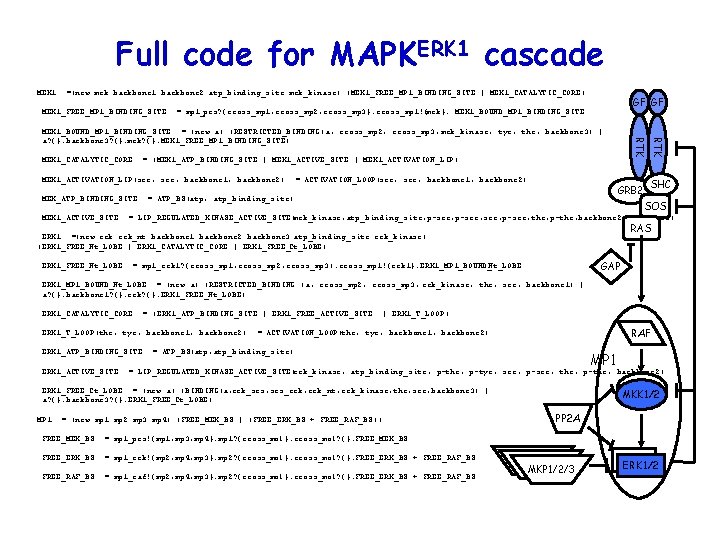
Full code for MAPKERK 1 cascade MEK 1: : =(new mek backbone 1 backbone 2 atp_binding_site mek_kinase) (MEK 1_FREE_MP 1_BINDING_SITE | MEK 1_CATALYTIC_CORE) GF GF MEK 1_FREE_MP 1_BINDING_SITE: : = mp 1_prs? {cross_mp 1, cross_mp 2, cross_mp 3}. cross_mp 1!{mek}. MEK 1_BOUND_MP 1_BINDING_SITE MEK 1_CATALYTIC_CORE: : = (MEK 1_ATP_BINDING_SITE | MEK 1_ACTIVE_SITE | MEK 1_ACTIVATION_LIP) MEK 1_ACTIVATION_LIP(ser, backbone 1, backbone 2): : = ACTIVATION_LOOP(ser, backbone 1, backbone 2) GRB 2 MEK_ATP_BINDING_SITE: : = ATP_BS(atp, atp_binding_site) RTK MEK 1_BOUND_MP 1_BINDING_SITE: : = (new a) (RESTRICTED_BINDING(a, cross_mp 2, cross_mp 3, mek_kinase, tyr, thr, backbone 3) | a? {}. backbone 3? {}. mek? {}. MEK 1_FREE_MP 1_BINDING_SITE) SHC SOS MEK 1_ACTIVE_SITE: : = LIP_REGULATED_KINASE_ACTIVE_SITE(mek_kinase, atp_binding_site, p-ser, p-ser, thr, p-thr, backbone 2, backbone 3) RAS ERK 1: : =(new erk_nt backbone 1 backbone 2 backbone 3 atp_binding_site erk_kinase) (ERK 1_FREE_Nt_LOBE | ERK 1_CATALYTIC_CORE | ERK 1_FREE_Ct_LOBE) GAP ERK 1_FREE_Nt_LOBE: : = mp 1_erk 1? {cross_mp 1, cross_mp 2, cross_mp 3). cross_mp 1!{erk 1}. ERK 1_MP 1_BOUND_ Nt_LOBE ERK 1_MP 1_BOUND_Nt_LOBE: : = (new a) (RESTRICTED_BINDING (a, cross_mp 2, cross_mp 3, erk_kinase, thr, ser, backbone 1) | a? {}. backbone 1? {}. erk? {}. ERK 1_FREE_Nt_LOBE) ERK 1_CATALYTIC_CORE: : = (ERK 1_ATP_BINDING_SITE | ERK 1_FREE_ACTIVE_SITE | ERK 1_T_LOOP) RAF ERK 1_T_LOOP(thr, tyr, backbone 1, backbone 2): : = ACTIVATION_LOOP(thr, tyr, backbone 1, backbone 2) ERK 1_ATP_BINDING_SITE: : = ATP_BS(atp, atp_binding_site) MP 1 ERK 1_ACTIVE_SITE: : = LIP_REGULATED_KINASE_ACTIVE_SITE(erk_kinase, atp_binding_site, p-thr, p-tyr, ser, p-ser, thr, p-thr, backbone 2) ERK 1_FREE_Ct_LOBE: : = (new a) (BINDING(a, erk_srs, srs_erk, erk_nt, erk_kinase, thr, ser, backbone 3) | a? {}. backbone 3? {}. ERK 1_FREE_Ct_LOBE) MP 1: : = (new mp 1 mp 2 mp 3 mp 4) (FREE_MEK_BS | (FREE_ERK_BS + FREE_RAF_BS)) MKK 1/2 PP 2 A FREE_MEK_BS: : = mp 1_prs!{mp 1, mp 3, mp 4}. mp 1? {cross_mol}. cross_mol? {}. FREE_MEK_BS FREE_ERK_BS: : = mp 1_erk!{mp 2, mp 4, mp 3}. mp 2? {cross_mol}. cross_mol? {}. FREE_ERK_BS + FREE_RAF_BS: : = mp 1_raf!{mp 2, mp 4, mp 3}. mp 2? {cross_mol}. cross_mol? {}. FREE_ERK_BS + FREE_RAF_BS MKP 1/2/3 ERK 1/2

p-calculus programs for ST pathways • • Unified coding of detailed and disparate data • Modular biology The Pi. FCP and SPi. FCP systems: semi- and fully quantitative (stochastic) computer simulation and tracing § p-calculus models for molecular and functional levels § Homology of processes
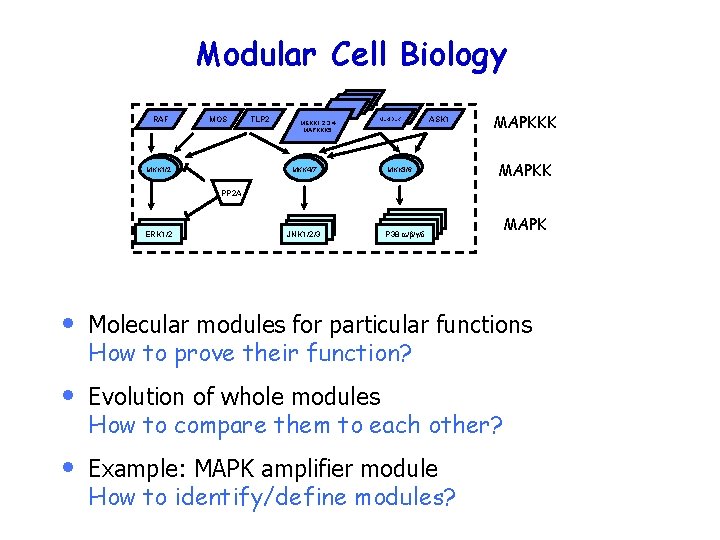
Modular Cell Biology RAF MOS MKK 1/2 TLP 2 MEKK 1, 2, 3, 4 MAPKKK 5 MKK 4/7 MLK/DLK ASK 1 MKK 3/6 MAPKKK MAPKK PP 2 A ERK 1/2 JNK 1/2/3 P 38 a/b/g/d MAPK • Molecular modules for particular functions How to prove their function? • Evolution of whole modules How to compare them to each other? • Example: MAPK amplifier module How to identify/define modules?

Establishing module function by a computational approach • Build two representations in the p-calculus § molecular level (implementation) § functional module level (specification) • Show the equivalence of both representations § by computer simulation § by formal verification (bisimulation)

Conclusions A comprehensive theory for § Unified formal representation of pathways and modules § Simulation and analysis § Comparative studies of process homologies We have developed § The theory of molecular processes as concurrent computation § A method for representing ST in the p-calculus § The Pi. FCP and SPi. FCP simulation systems

Future work • • Study various systems with simulation tools Improve representation § Dual face of interaction § Module and complex integrity • Comparative measures § Pathway and function § Process homology

Acknowledgements TAU WIS • • • Eva Jablonka Yehuda Ben-Shaul Udi Shapiro Bill Silverman Naama Barkai
- Slides: 35