Simulation Based Study of a Diffusion MRI Process

Simulation Based Study of a Diffusion MRI Process in Muscle Tissue Nadia Smith, Jessica Talbott, Chris Clark and Matt G Hall MEMPHYS Welcome to the National Physical Laboratory Metrology for Medical Physics

Me & Background § I have been working as a Research Scientist in Data Science at NPL for just over a year § MSc in Complexity Science § MSc in Mathematical Medicine and Biology § The slides have more words on them than normal as I am aware my voice is unclear, however please approach me with comments/questions, my friend Thea will interpret if necessary § Or, email jessica. talbott@npl. co. uk
Diffusion MRI § Diffusion Weighted Magnetic Resonance Imaging (MRI) is sensitive to the bulk motion of spins § Spins diffuse in tissue, and the tissue affects their diffusion § Measuring diffusion allows to infer details about the tissue § Diffusion happens on a very small scale: 1 -10 microns § Much smaller than a typical scan voxel: 1 -3 mm § Diffusion gives a window into microscopic structure non-invasively

Duchenne Muscular Dystrophy § Duchenne muscular dystrophy (DMD) is a genetic muscle wasting condition affecting approximately 1 in 3600 boys, it causes progressive muscle loss and is invariably fatal § DMD tissue is often imaged for fat-fraction, but replacement of muscle with fat is the end stage of the pathology § Diffusion MRI is a method that is sensitive to earlier stage microstructural changes § DMD is known to cause increased permeability of muscle fibres § Changes in permeability are not uniform across space however - permeable fibres are spatially randomly distributed and histological evidence for permeability changes reminiscent of a percolation cluster has been reported Images: N. Hornby

Overview of the work § The effect of spatial distribution of permeability on the diffusion signal has not been extensively studied § Previous simulation-based investigations typically considered permeability as a uniform change across tissue § Here we look at changing permeability in two scenarios: 1. uniformly across boundaries of tissue and interstitial space 2. when boundaries are permeable or impermeable with random probability In both cases the total expected flux across all barriers is the same for comparison sake

Methods § Synthetic pulsed gradient spin-echo (PGSE) diffusion-weighted signal curves are generated over a wide range of realistic MR scan parameters § A Finite Element Method (FEM) is used to numerically solve the Bloch. Torrey equation (fundamental to MR modalities) for spins in muscle tissue excited via a PGSE sequence § The solution is integrated over the sample volume to compute the synthetic signal

The diffusion equations Bloch-Torrey equation to describe the evolution of magnetisation undergoing diffusion, in the absence of flow: This needs to be solved subject to the effects of diffusion, boundary conditions and the time dependence of the magnetic gradients in the spin-echo pulse sequence. Taking the magnitude of the complex result allows determination of the signal attenuation.

Why COMSOL Multiphysics. TM? § Simulations for diffusion MRI typically adopt Monte Carlo approaches § Dealing with exchange between compartments with different diffusivities however leads to an equilibrium condition with higher concentrations of spins in low diffusivity regions § As a consequence, low diffusivity regions over contributes to the signal. COMSOL allows us to treat the spin population as a continuum and numerically solve fundamental equations that govern MRI
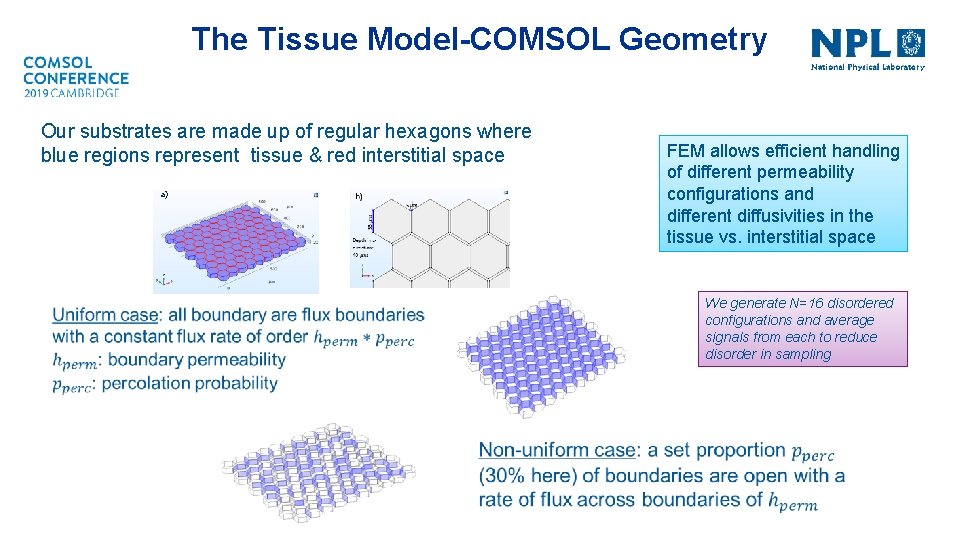
The Tissue Model-COMSOL Geometry Our substrates are made up of regular hexagons where blue regions represent tissue & red interstitial space a) FEM allows efficient handling of different permeability configurations and different diffusivities in the tissue vs. interstitial space We generate N=16 disordered configurations and average signals from each to reduce disorder in sampling

Geometry Construction Ø We use geometric expressions to form a lattice of regular hexagons that gives efficient packing Ø We make use of selects to impose conditions to set elements
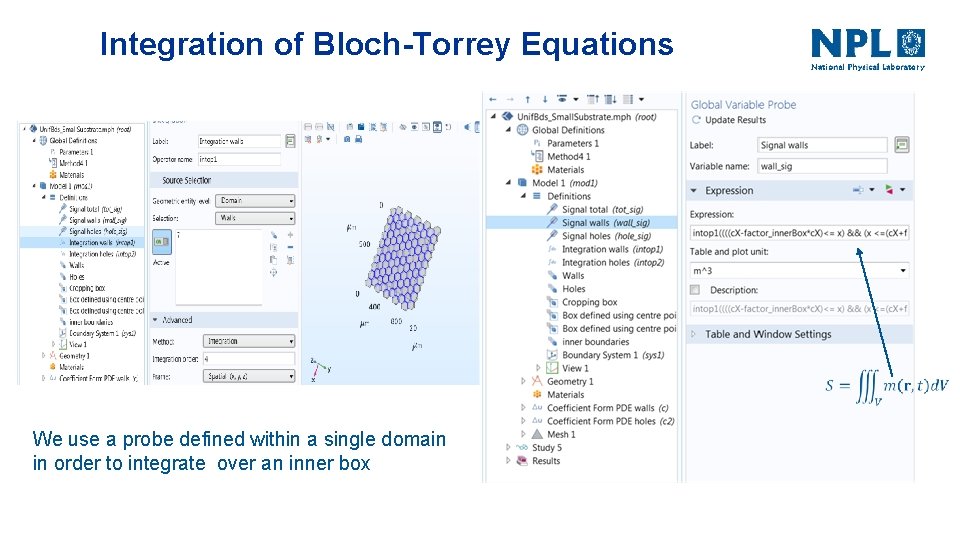
Integration of Bloch-Torrey Equations We use a probe defined within a single domain in order to integrate over an inner box
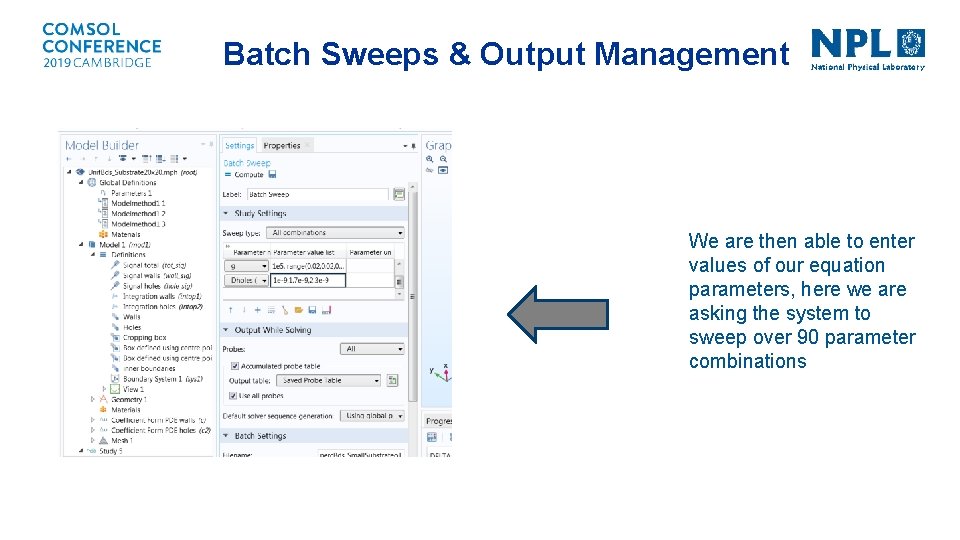
Batch Sweeps & Output Management We are then able to enter values of our equation parameters, here we are asking the system to sweep over 90 parameter combinations

Model Methods: Non-uniform Case double PERCOLATION_PROB = model. param(). evaluate ("perc_prob"); // Define random number between 0 and 1 - RAND_NUM for (int i = 0; i < flux_bnd_new. length; i++) { double rand_num = Math. random(); if (rand_num < PERCOLATION_PROB) { selected_bnds. add(new Integer(flux_bnd_new[i])); } } // set selection of flux boundaries to be selected_bnds in Comsol int[] ret. Val = new int[selected_bnds. size()]; for (int i = 0; i < ret. Val. length; i++) { ret. Val[i] = selected_bnds. get(i). int. Value(); } model. component("mod 1"). selection("sel 8"). set(ret. Val); For the percolation case we use a Model Method imposed on the inner boundaries of the lattice in order to randomly select a set percentage to be ‘open’

Running on a Cluster • This procedure rapidly becomes computationally expensive as we vary permeability and substrate size • FE models are built in COMSOL Multiphysics. TM 5. 3 a & passed through NPL’s HPC cluster, known as Minerva for run optimization § COMSOL files are integrated into a Minerva Slurm script to manage the 90 parameter sweep combination remotely. § The script forces simulations into parallel mode, running the sweep as a job array across nodes with one parameter combination per CPU and per node. § The parameter combinations are all defined in a single input parameter file, which is read one line at a time by the job submission script allowing to dispatch different parameter combinations across Minerva. § The script generates one COMSOL model per sub-job with one output file
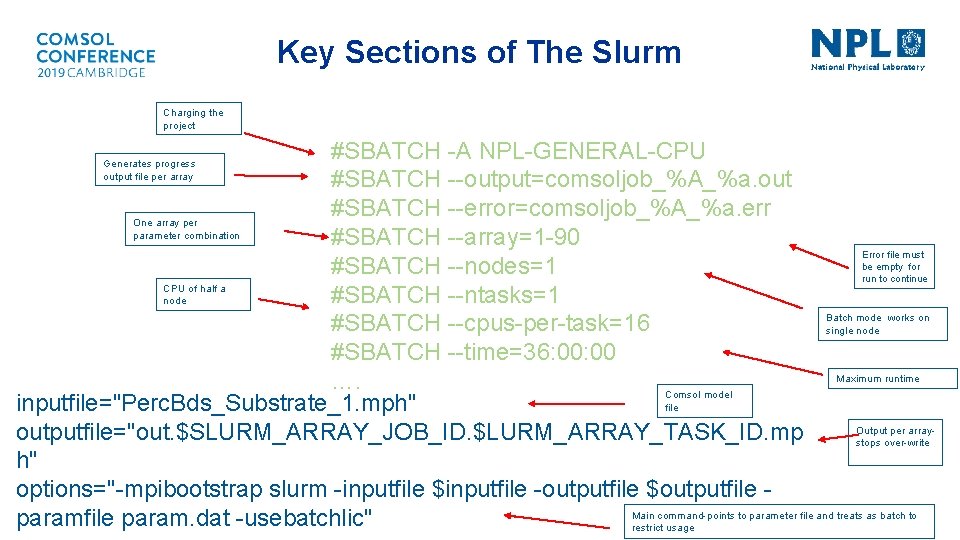
Key Sections of The Slurm Charging the project #SBATCH -A NPL-GENERAL-CPU #SBATCH --output=comsoljob_%A_%a. out #SBATCH --error=comsoljob_%A_%a. err One array per parameter combination #SBATCH --array=1 -90 Error file must be empty for #SBATCH --nodes=1 run to continue CPU of half a #SBATCH --ntasks=1 node Batch mode works on #SBATCH --cpus-per-task=16 single node #SBATCH --time=36: 00 Maximum runtime …. Comsol model inputfile="Perc. Bds_Substrate_1. mph" file Output per array- outputfile="out. $SLURM_ARRAY_JOB_ID. $LURM_ARRAY_TASK_ID. mp stops over-write h" options="-mpibootstrap slurm -inputfile $inputfile -outputfile $outputfile Main command-points to parameter file and treats as batch to paramfile param. dat -usebatchlic" restrict usage Generates progress output file per array
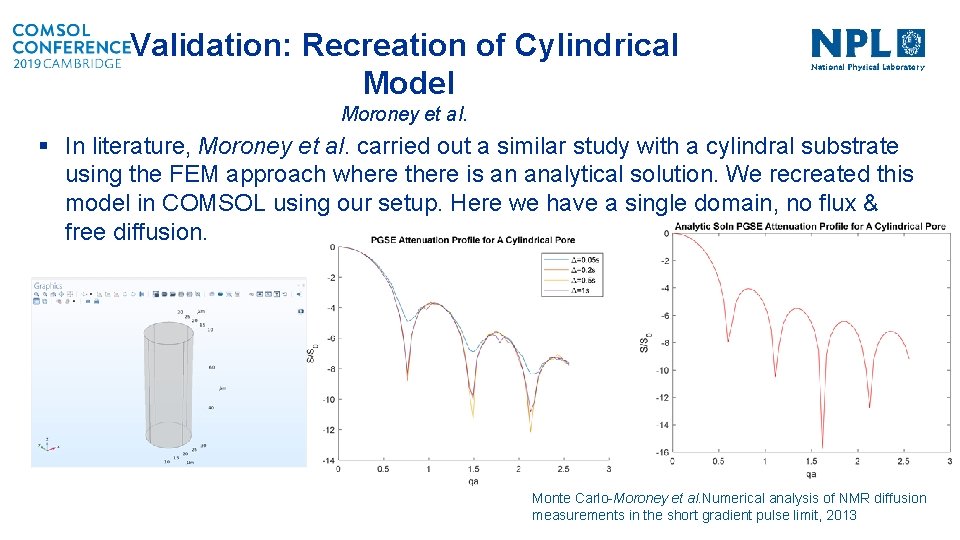
Validation: Recreation of Cylindrical Model Moroney et al. § In literature, Moroney et al. carried out a similar study with a cylindral substrate using the FEM approach where there is an analytical solution. We recreated this model in COMSOL using our setup. Here we have a single domain, no flux & free diffusion. Monte Carlo-Moroney et al. Numerical analysis of NMR diffusion measurements in the short gradient pulse limit, 2013
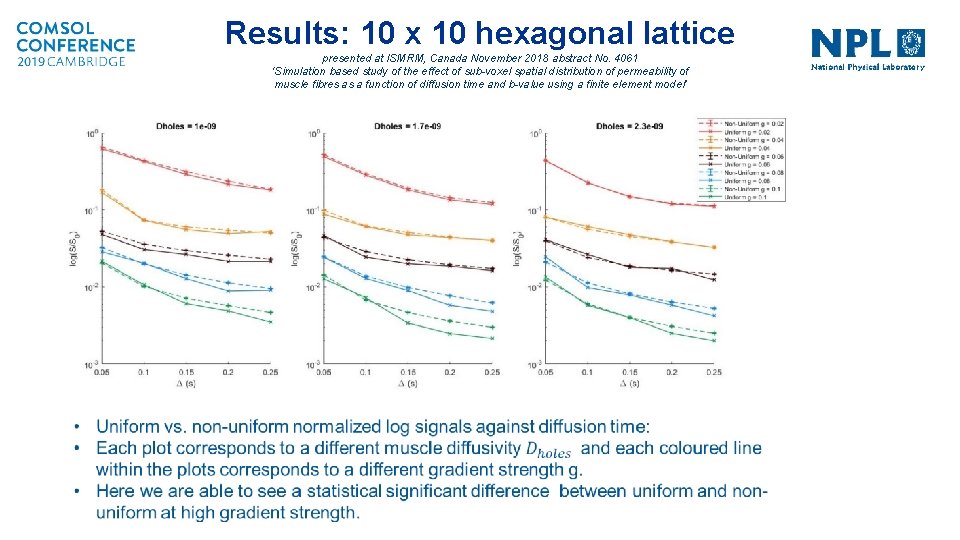
Results: 10 x 10 hexagonal lattice presented at ISMRM, Canada November 2018 abstract No. 4061 ‘Simulation based study of the effect of sub-voxel spatial distribution of permeability of muscle fibres as a function of diffusion time and b-value using a finite element model’

Discussion § COMSOL is ideal for our purposes, construction of the model is intuitive and numerical output is in line with hypotheses § The study has enabled us to present results so far in Canada (ISMRM) and Berlin (MYO-MRI)

THANK YOU Any Questions? jessica. talbott@npl. co. uk The National Physical Laboratory is operated by NPL Management Ltd, a wholly-owned company of the Department for Business, Energy and Industrial Strategy (BEIS).
- Slides: 19