Reproducible Data Analysis Workflows OPENBIS UGM 2019 Michal

Reproducible Data Analysis Workflows OPENBIS UGM 2019 Michal Okoniewski, Andrei Plamadă ETH Zürich – Scientific IT Services ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 1

Outline § Reproducibility and Scientific Computing § Best Practices § Workflow Management Systems § Introduction § Snakemake and Hands-On § Snakemake with Genomics Example § Reproducible Environment § Introduction § Open. BIS Integration § Containers and Conda Hands-On ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 2

Getting to know each other 1. 2. 3. 4. Which OS do you use: Windows 7, Windows 10, Linux, mac. OS, other? How often do you program: weekly, monthly, yearly? Do you use Python / R? What is your background: formal (Math+CS), physical, social, life sciences; engineering, medicine? 5. Did you have difficulties in reproducing your own work? 6. Did you hear about / use git? 7. Did you hear about / use workflow management systems? 8. Did you hear about / use containers? 9. Did you hear about / use conda? 10. Did you hear about / use MPI? ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 3

Outline § Reproducibility and Scientific Computing § Best Practices § Workflow Management Systems § Introduction § Snakemake and Hands-On § Snakemake with Genomics Example § Reproducible Environment § Introduction § Open. BIS Integration § Containers and Conda Hands-On ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 4

What is Reproducibility in Scientific Computing ? ? ? ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 5

What is Reproducibility in Scientific Computing Data Code Environment ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 6
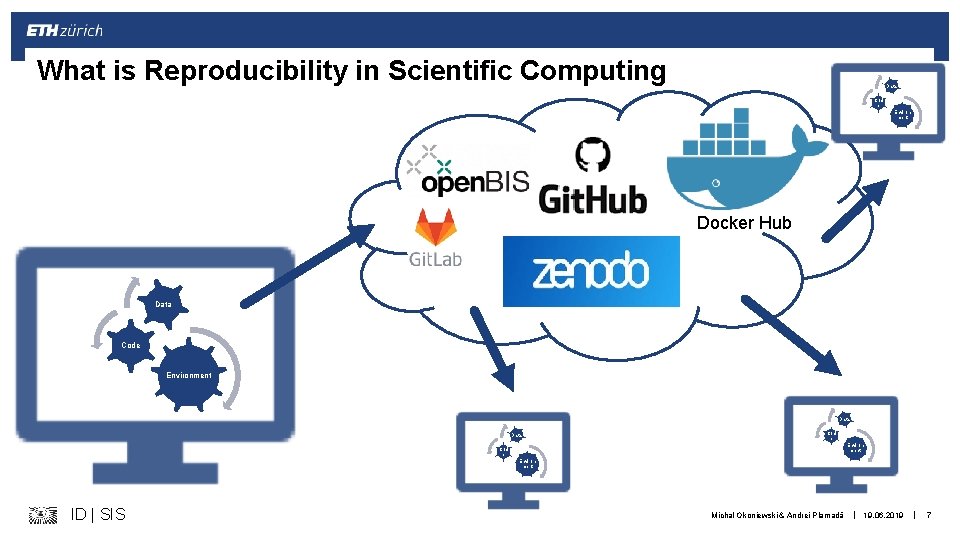
What is Reproducibility in Scientific Computing Data Cod e Environ ment Docker Hub Data Code Environment Data Cod e Environ ment ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 7

Reproducibility PI Manifesto "Reproducibility PI Manifesto", L. A. Barba. (13 December 2012). 10. 6084/m 9. figshare. 104539 1. I will teach my graduate students about reproducibility: a. lab notebook, b. version control, c. workflow, d. publication-quality plots at group meeting. 2. All our research code (and writing) is under version control. 3. We will always carry out verification and validation (V&V reports are posted to figshare). 4. For main results in a paper, we will share data, plotting script & figure under CC-BY. 5. We will upload the preprint to ar. Xiv at the time of submission of a paper. 6. We will release code at the time of submission of a paper. 7. We will add a "Reproducibility" declaration at the end of each paper. 8. I will keep an up-to-date web presence. ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 8

Best Practices for Reproducibility in Scientific Computing Kitzes, J. , Turek, D. , & Deniz, F. (Eds. ). (2018). The Practice of Reproducible Research: Case Studies and Lessons from the Data-Intensive Sciences. Oakland, CA: University of California Press. Lessons Learned – Kathryn Huff https: //www. practicereproducibleresearch. org/core-chapters/5 -lessons. html § Very common: § § § Version control your code Open your data Automate everywhere possible Document your process Test everything Use free and open tools ID | SIS § Less common: § Avoid excessive dependencies § When dependencies can’t be avoid, package their installation § Host code on collaborative platforms (e. g. Git. Hub) § Get a Digital Object Identifier for your data and code § Avoid spreadsheets, plain text data is preferred § Explicitly set pseudorandom number generator seeds § Workflow and provenance framework may be too clunky for most scientist Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 9

Best Practices for Scientific Computing Wilson G, Aruliah DA, Brown CT, Chue Hong NP, Davis M, Guy RT, et al. (2014) Best Practices for Scientific Computing. PLo. S Biol 12(1): e 1001745. https: //doi. org/10. 1371/journal. pbio. 1001745 1. Write Programs for People, not Computers • Readability and Style 2. Let the Computer Do the Work • • Scripts -> Automated workflows Unique version for code, data, dependencies 3. Make Incremental Changes • Version control (git) 4. Don’t Repeat Yourself (or others) • Re-use the code 6. Optimize Software Only after It Works Correctly • 5. + Profiling 7. Document Design and Purpose, Not Mechanism • Documentation 8. Collaborate • Issue tracking and Code Review (e. g. github, gitlab) 5. Plan for Mistakes • Testing and Continuous Integrations ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 10

So many things to learn! Where to start? ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 11

Outline § Reproducibility and Scientific Computing § Best Practices § Workflow Management Systems § Introduction § Snakemake and Hands-On § Snakemake with Genomics Example § Reproducible Environment § Introduction § Open. BIS Integration § Containers and Conda Hands-On ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 12

A Zoo of Data Workflow Systems § An incomplete list of 254 Computational Data Analysis Workflow Systems § https: //github. com/common-workflow-language/wiki/Existing-Workflowsystems § A curated list of 90 Awesome Pipeline frameworks & libraries + 27 Workflow platforms § https: //github. com/pditommaso/awesome-pipeline ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 13
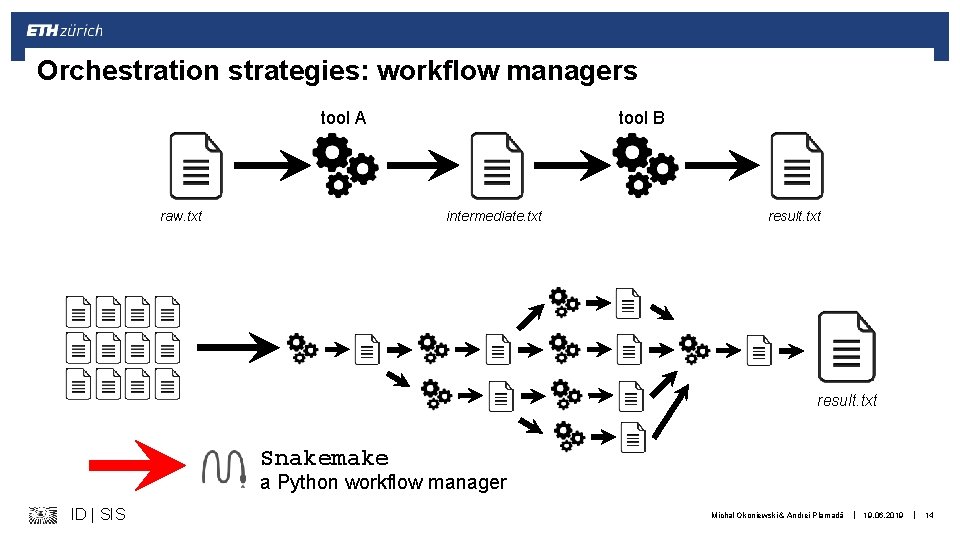
Orchestration strategies: workflow managers tool A raw. txt tool B intermediate. txt result. txt Snakemake a Python workflow manager ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 14

Snakemake § Workflow management system § Designed by Johannes Köster § Now PI at Uni. Essen § § § Python 3 – based cmake philosophy conda installation conda support http: //snakemake. readthedocs. io/ ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 15

Installation § Install miniconda § Download and run the installer (eg. Miniconda 3 -latest-Linux-x 86_64. sh) § Install snakemake with conda § conda install -c bioconda -c conda-forge snakemake-minimal § Test § snakemake --version ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 16

Parsing the workflow § rule_all defines the final product § Snakemake parses searches for files needed to do this final products § Then, recursively, searches for what needs to be done for the “substrates” § After successful parsing (in syntax and content): § Workflow is started from the “substrates” of lowest level § Proceeds as DAG (directed acyclic graph) towards the final product ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 17
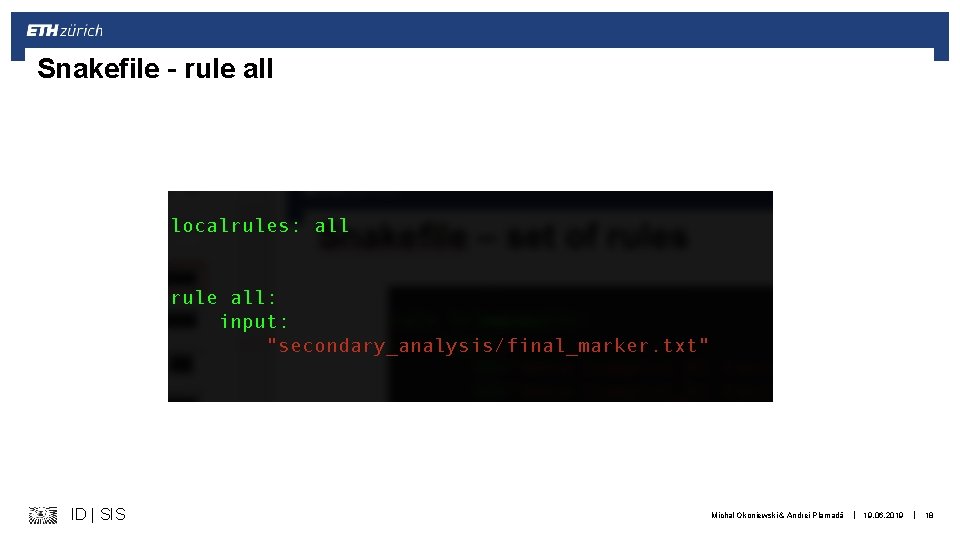
Snakefile – rule all ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 18
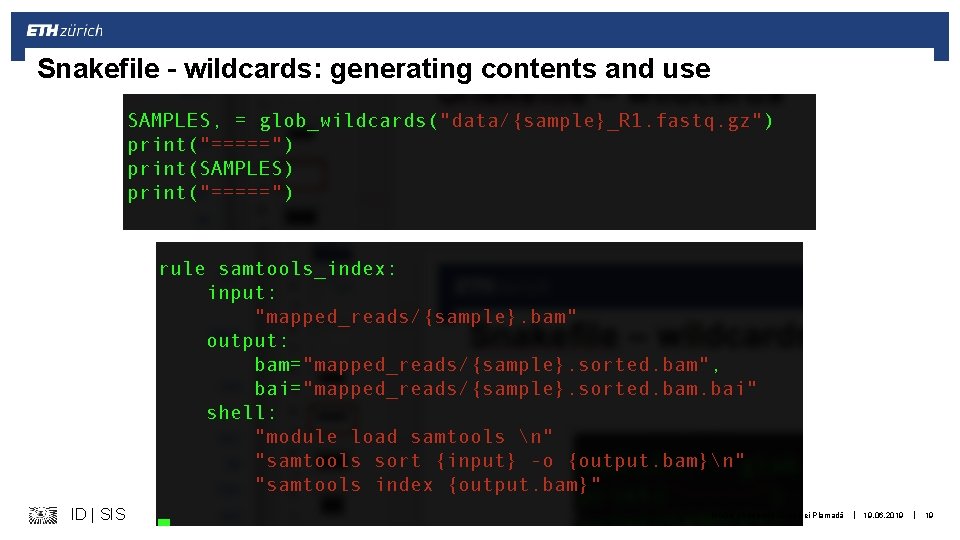
Snakefile – wildcards: generating contents and use ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 19
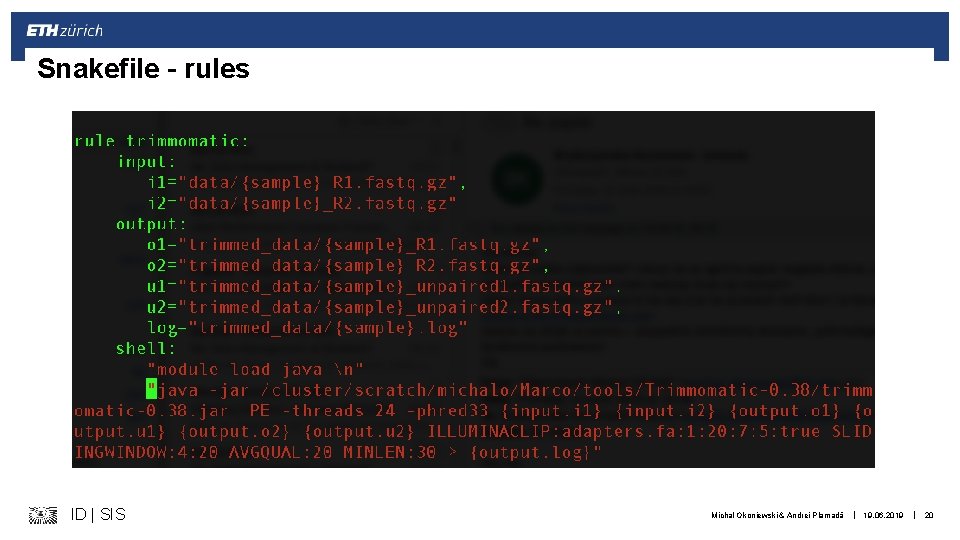
Snakefile – rules ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 20
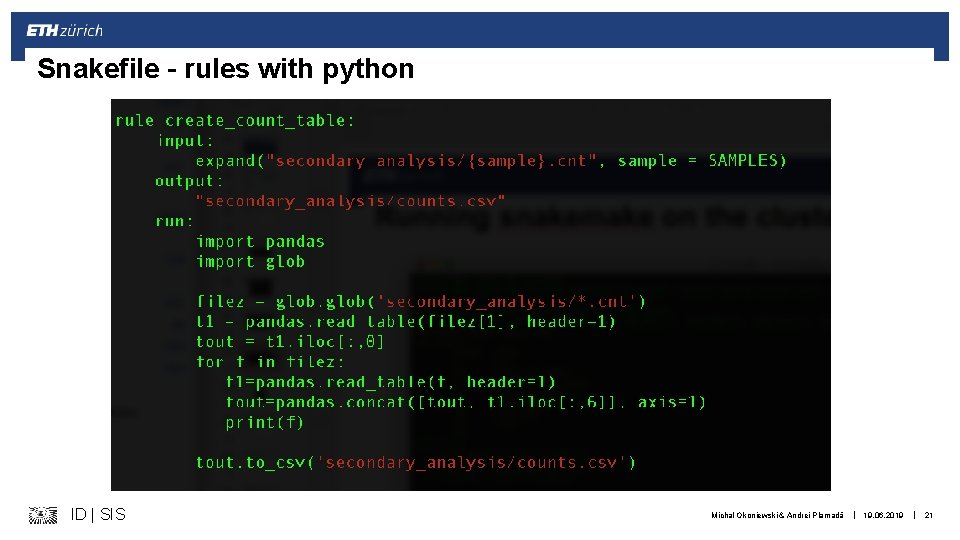
Snakefile – rules with python ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 21

Running snakemake on the cluster LSF snakemake -p -j 999 --cluster-config cluster. json --cluster "bsub -W {cluster. time} -n {cluster. n}” SLURM snakemake -j 999 --cluster-config cluster. json --cluster "sbatch -A {cluster. account} -p {cluster. partition} -n {cluster. n} -t {cluster. time}" Kubernetes snakemake --kubernetes --use-conda --default-remote-provider $REMOTE --default-remote-prefix $PREFIX ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 22
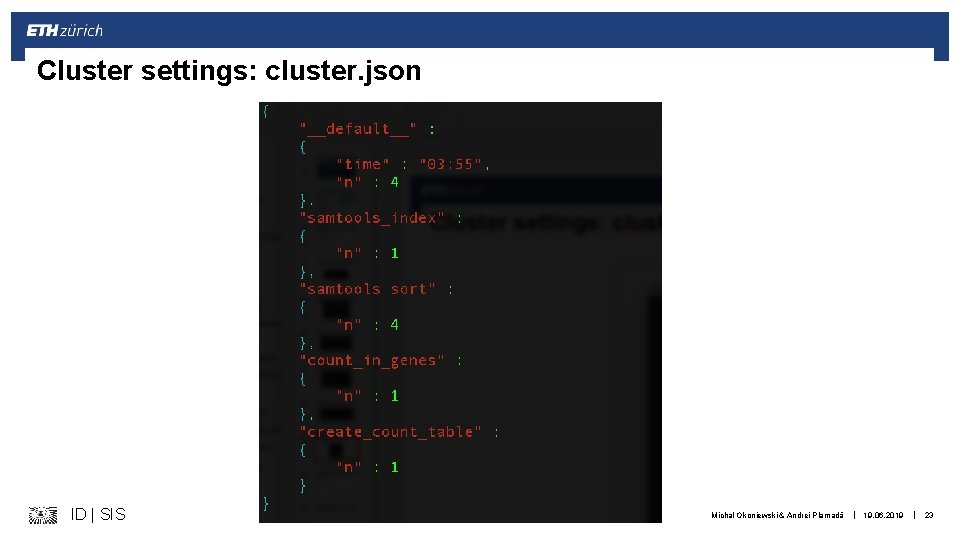
Cluster settings: cluster. json ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 23
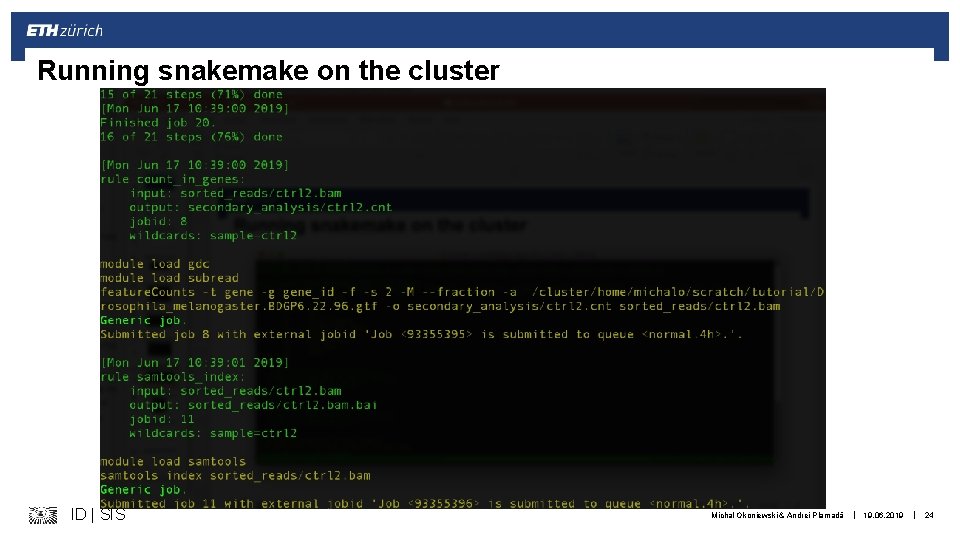
Running snakemake on the cluster ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 24

Demo on the computing cluster § § Genomic example 6 BAM (genome alignment) files on the input Operations: sorting, indexing, counting of read in genes, count table production Cluster. json specific for LSF on Euler cluster ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 25

Visualizing of what we actually done by snakemake § Directed acycylic graph of jobs § Can be seen with snakemake --dag > graph. dag dot -Tpdf graph. dag > aaa. pdf § Visualizes dependencies of rules ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 26
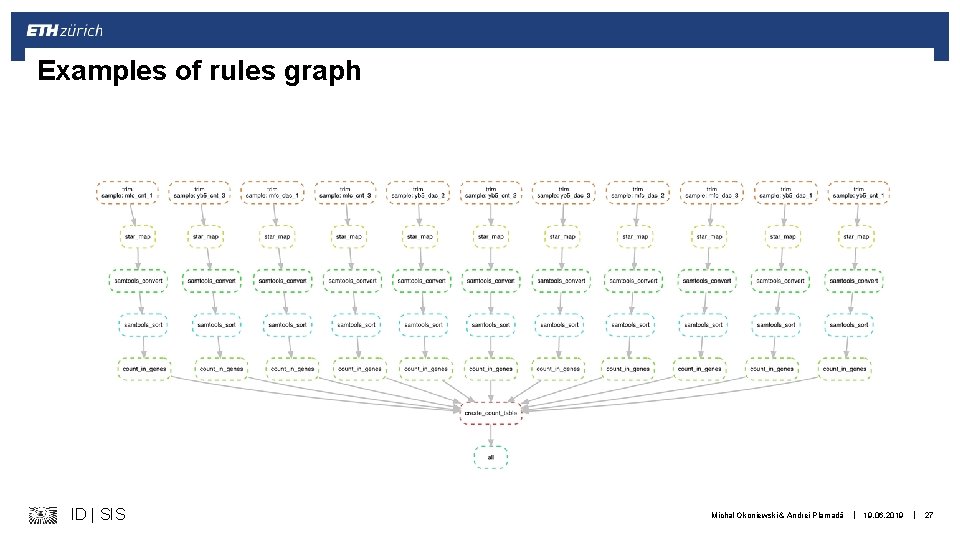
Examples of rules graph ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 27
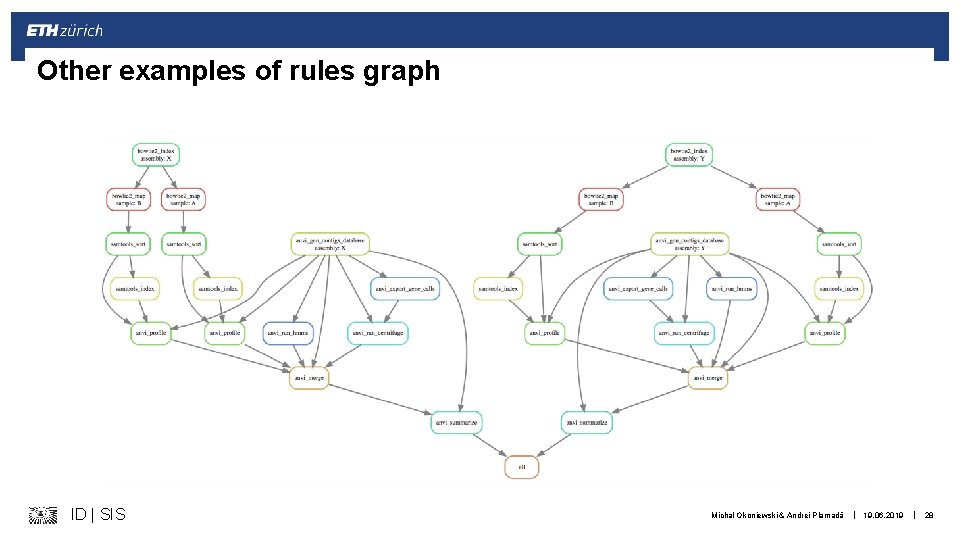
Other examples of rules graph ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 28
Snakemake happily finished ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 29

Advantages and difficulties of snakemake § Reproducibility § Control over workflow § Re-running § Encapsulation of typical tasks § “One-click” starting of a large process § You need to “speak python” § Learning curve steep at the beginning ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 30

Other reproducibility mechanisms that can be used by snakemake § § § Common Workflow Language Remote files Integrated package management with Conda Running jobs in containers Wrappers ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 31
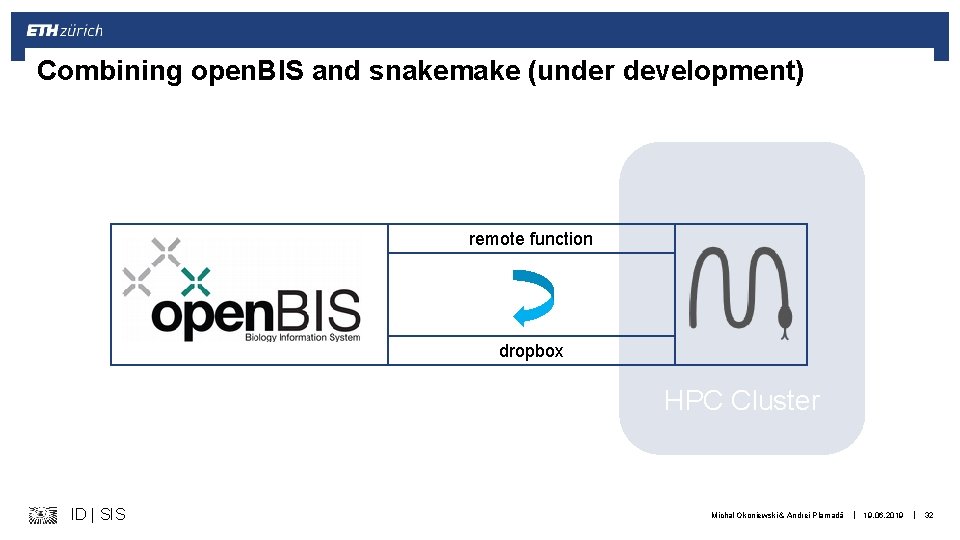
Combining open. BIS and snakemake (under development) remote function dropbox HPC Cluster ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 32

Practical advice § Test your workflow with a “dry run”: snakemake –np § Real run test – with small number of input files, eg 3 § On the cluster § § § run snakemake in a screen session on a login node run snakemake on personal scratch or other permanent storage use local rules whenever possible I/O rules – define as single core jobs in cluster. json check time, memory, cores settings for each job § Consider deleting intermediate files after use § Sometimes deleting. snakemake may be needed for re-run ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 33

Hands-on exercise on a single machine § https: //github. com/michalogit/snakemaketax ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 34

Outline § Reproducibility and Scientific Computing § Best Practices § Workflow Management Systems § Introduction § Snakemake and Hands-On § Snakemake with Genomics Example § Reproducible Environment § Introduction § Open. BIS Integration § Containers and Conda Hands-On ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 35

Reproducible Environment § Main idea: bundle your application and all dependencies § Virtual Machine (VM): Virtual. Box, VMware § Container - lightweight VM: Docker, Singularity § Isolated environment: § Python: Virtual Environment, Conda § R: Conda § As a side effect: No more version conflicts (Dependency hell) ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 36
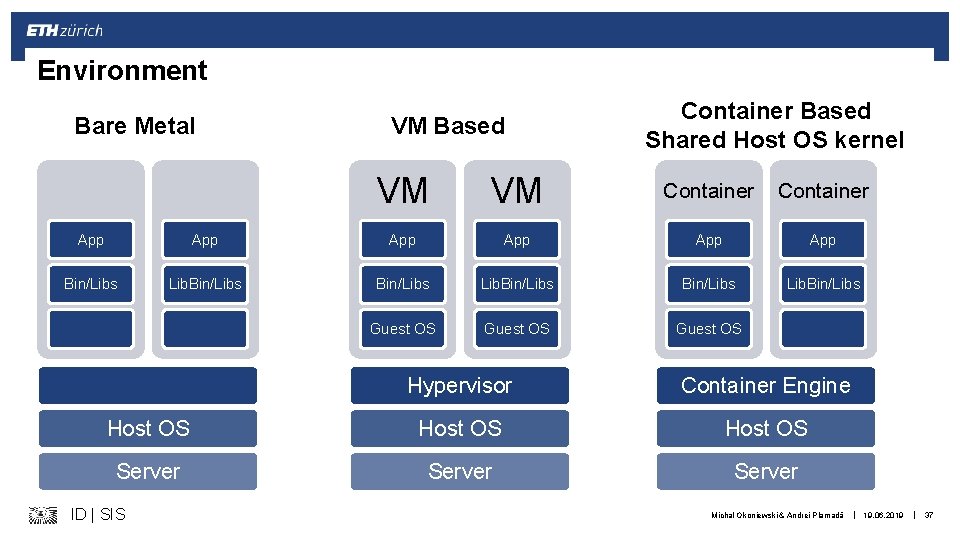
Environment Bare Metal VM Based Container Based Shared Host OS kernel VM VM Container App App App Bin/Libs Lib. Bin/Libs Guest OS Hypervisor Container Engine Host OS Server ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 37
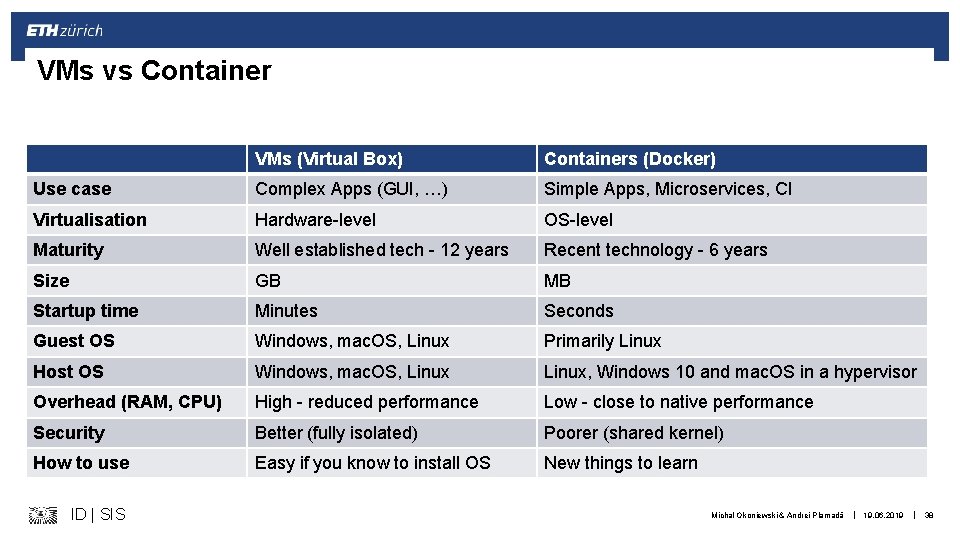
VMs vs Container VMs (Virtual Box) Containers (Docker) Use case Complex Apps (GUI, …) Simple Apps, Microservices, CI Virtualisation Hardware-level OS-level Maturity Well established tech - 12 years Recent technology - 6 years Size GB MB Startup time Minutes Seconds Guest OS Windows, mac. OS, Linux Primarily Linux Host OS Windows, mac. OS, Linux, Windows 10 and mac. OS in a hypervisor Overhead (RAM, CPU) High - reduced performance Low - close to native performance Security Better (fully isolated) Poorer (shared kernel) How to use Easy if you know to install OS New things to learn ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 38
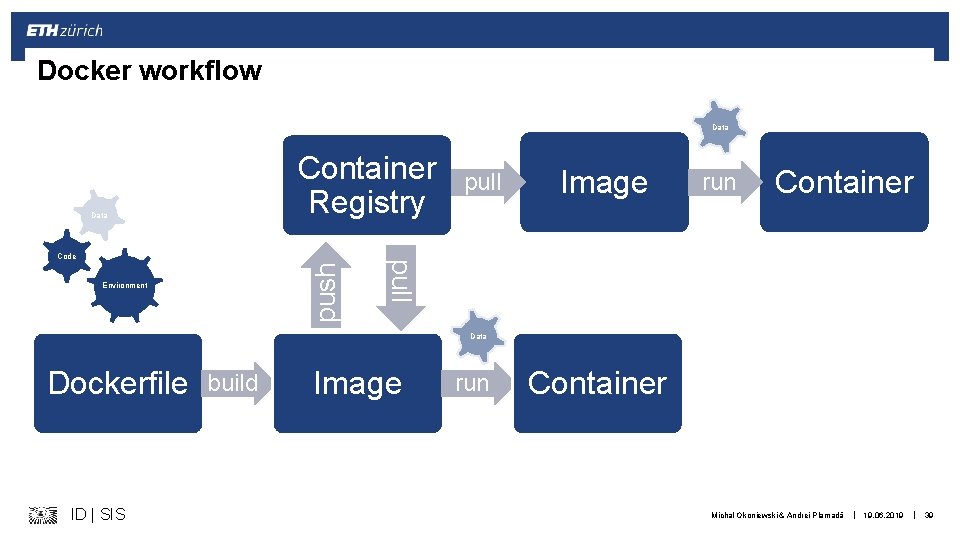
Docker workflow Data Container Registry Data push Environment Image run Container pull Code pull Data Dockerfile ID | SIS build Image run Container Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 39

Nice but Docker requires root access What about HPC systems? ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 40

Singularity as the container solution for HPC § Containers improve portability and can address the reproducibility issue in research (Enhance. R Survey - Science IT Consultants) § Singularity: § § § § Developed initially at LBL - Berkeley Lab - for HPC use case (multi-tenancy, single file) Open source with standard BSD 3 clause license https: //github. com/sylabs/singularity Under active development with 12 contributors with more than 100 commits Available also with commercial support: Singularity Pro Used world wide and recommended by vendors, e. g. NVIDIA, Azure Batch Big worldwide community (google groups, slack) Swiss community - Enhance. R 2 major versions: Singularity 2 and Singularity 3 ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 41

Singularity as the container solution for HPC § Containers improve portability and can address the reproducibility issue in research (Enhance. R Survey - Science IT Consultants) § Main idea Host OS+Drivers+Middleware (OSDM) MPI • mpirun • MPI Library ID | SIS SSH Server Host OS+Drivers+Middleware (OSDM) Container OSDM App MPI • Shared MPI Library • mpirun SSH Server • MPI • App • Shared MPI Library Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 42

User Experience for Containers – Docker + Singularity v 2. 6 Build Run • Docker • root access • on your PC • Singularity • on your PC or HPC infrastructure § Multi-node: MPICH ABI Compatibility initiative ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 43

Why to bother with Containers? I use only Python / R ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 44

Isolated Environment for R and Python - Conda § Conda https: //docs. conda. io/en/latest/ § Open source § Runs on Windows, mac. OS, Linux § Package management system https: //anaconda. org/search § Supported Programming languages: Python, R, … § Repository: https: //anaconda. org/ § Environment management system ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 45
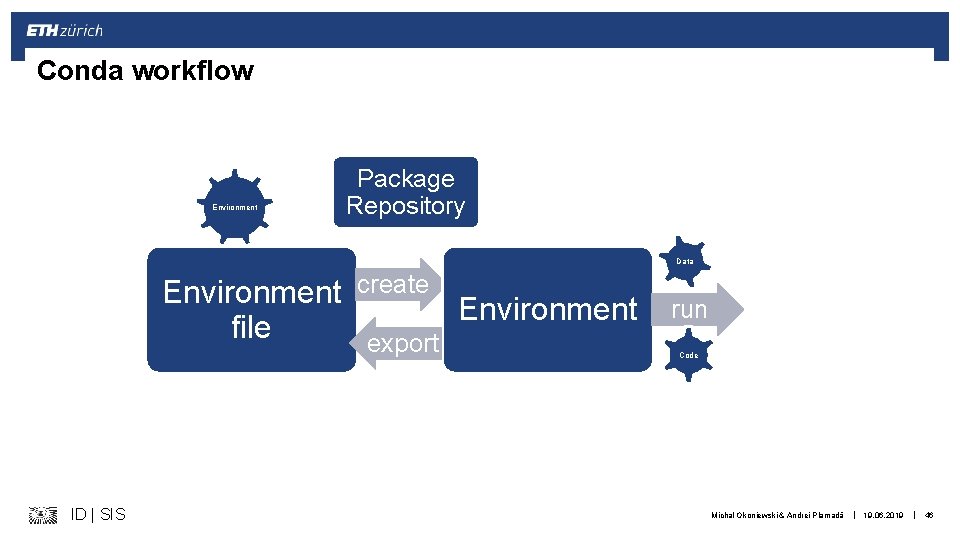
Conda workflow Data Code Environment Package Repository Data Environment create Environment file export ID | SIS run Code Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 46

What can go wrong? § Containers: § § The image is updated - same tag different content: e. g. centos: latest The image is deleted by the owner The old container does not work with the new Docker/Singularity (not very likely) The new container does not work with old Docker/Singularity § Conda § § The package metadata (dependency list) is updated (not very likely) The package is deleted by the owner Python: you mix pip and conda and do a conda update Conda packages are not platform independent ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 47

Things to consider § ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 48
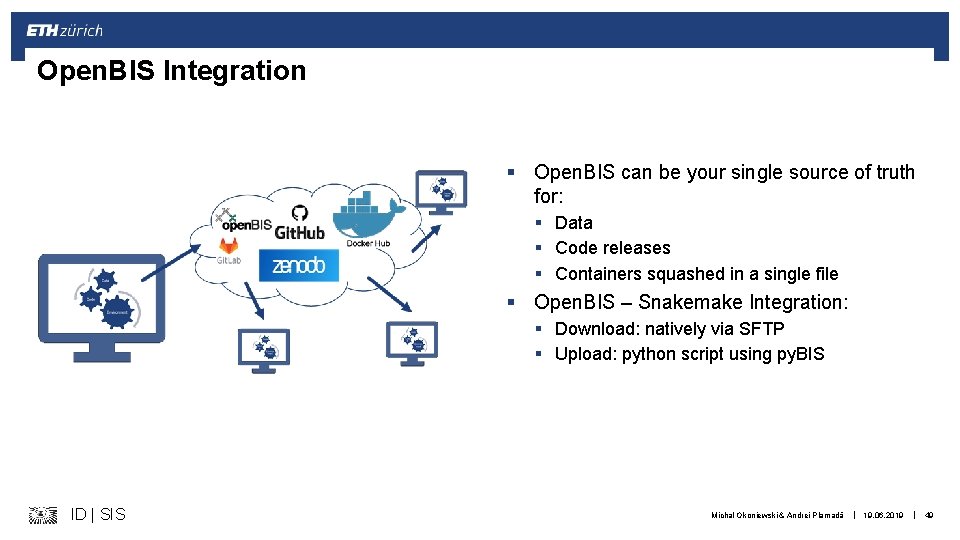
Open. BIS Integration § Open. BIS can be your single source of truth for: § Data § Code releases § Containers squashed in a single file § Open. BIS – Snakemake Integration: § Download: natively via SFTP § Upload: python script using py. BIS ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 49

Hands-on exercise on a single machine § https: //siscourses. ethz. ch/openbis_ugm_2019/Containers_and_Conda_Hands_ On. html ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 50

Summary § Reproducibility in Scientific Computing: Data, Code and Environment § We are studying the interplay between reproducible research techniques § For ETH Zurich we are responsible for providing solutions in this area § New developments in this area for open. BIS are coming ID | SIS Michal Okoniewski & Andrei Plamadă | 19. 06. 2019 | 51

Thanks!
- Slides: 52