Learning Boolean Networks from Tox Cast HighContent Imaging

Learning Boolean Networks from Tox. Cast High-Content Imaging Data Todor Antonijevic ORCID ID 0000 -0002 -0248 -8412 The views expressed in this presentation are those of the author[s] and do not necessarily reflect the views or policies of the U. S. Environmental Protection Agency.

Outline I. Introduction II. Methods: 1. 2. 3. 4. 5. 6. Dataset Data standardization, and Noise Threshold (z 0). Data Discretization. Learning Boolean Functions and Construction of Boolean Networks (BNs). Needleman-Wunsch (NW) optimal global alignment, and Error Estimation. Coverage. III. Results: 1. Discretized Trajectories and Total Perturbation. 2. Clustering of discretized trajectories, Error Estimation, and Coverage (first 10 BNs). 3. Learned BNs in case of Butachlor. IV. Summary 2

Outline I. Introduction II. Methods: 1. 2. 3. 4. 5. 6. Dataset Data standardization, and Noise Threshold (z 0). Data Discretization. Learning Boolean Functions and Construction of Boolean Networks (BNs). Needleman-Wunsch (NW) optimal global alignment, and Error Estimation. Coverage. III. Results: 1. Discretized Trajectories and Total Perturbation. 2. Clustering of discretized trajectories, Error Estimation, and Coverage (first 10 BNs). 3. Learned BNs in case of Butachlor. IV. Summary 3
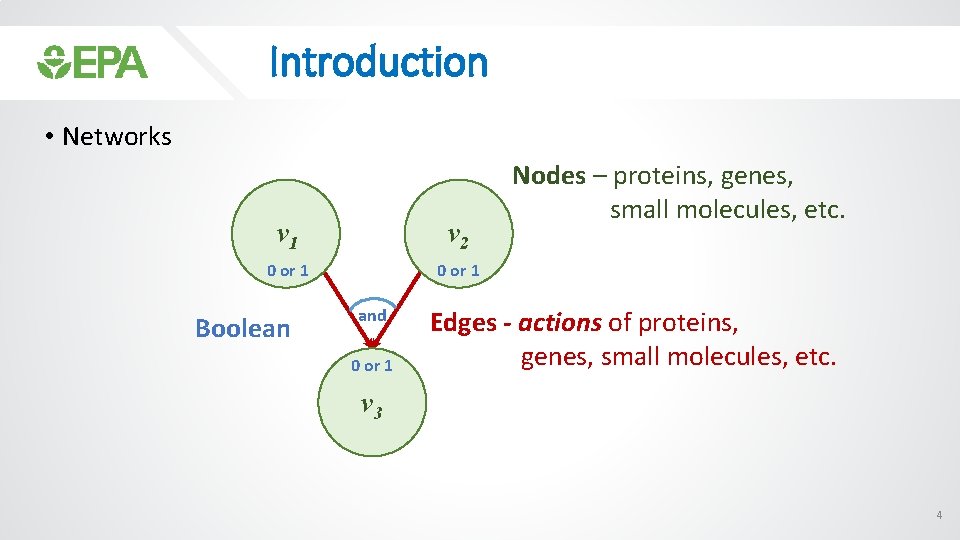
Introduction • Networks v 1 v 2 0 or 1 Boolean and 0 or 1 Nodes – proteins, genes, small molecules, etc. Edges - actions of proteins, genes, small molecules, etc. v 3 4
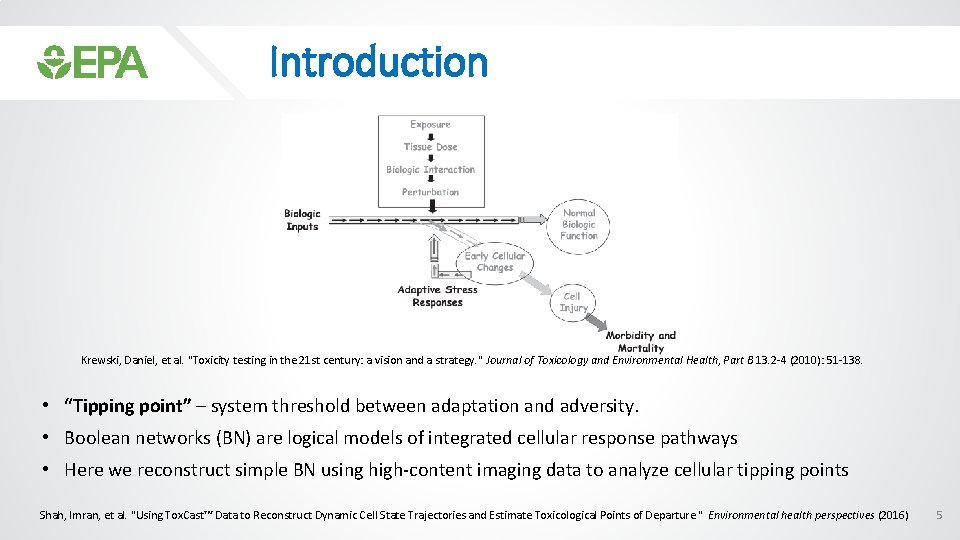
Introduction Krewski, Daniel, et al. "Toxicity testing in the 21 st century: a vision and a strategy. " Journal of Toxicology and Environmental Health, Part B 13. 2 -4 (2010): 51 -138. • “Tipping point” – system threshold between adaptation and adversity. • Boolean networks (BN) are logical models of integrated cellular response pathways • Here we reconstruct simple BN using high-content imaging data to analyze cellular tipping points Shah, Imran, et al. "Using Tox. Cast™ Data to Reconstruct Dynamic Cell State Trajectories and Estimate Toxicological Points of Departure. " Environmental health perspectives (2016) 5

Outline I. Introduction II. Methods: 1. 2. 3. 4. 5. 6. Dataset Data standardization, and Noise Threshold (z 0). Data Discretization. Learning Boolean Functions and Construction of Boolean Networks (BNs). Needleman-Wunsch (NW) optimal global alignment, and Error Estimation. Coverage. III. Results: 1. Discretized Trajectories and Total Perturbation. 2. Clustering of discretized trajectories, Error Estimation, and Coverage (first 10 BNs). 3. Learned BNs in case of Butachlor. IV. Summary 6
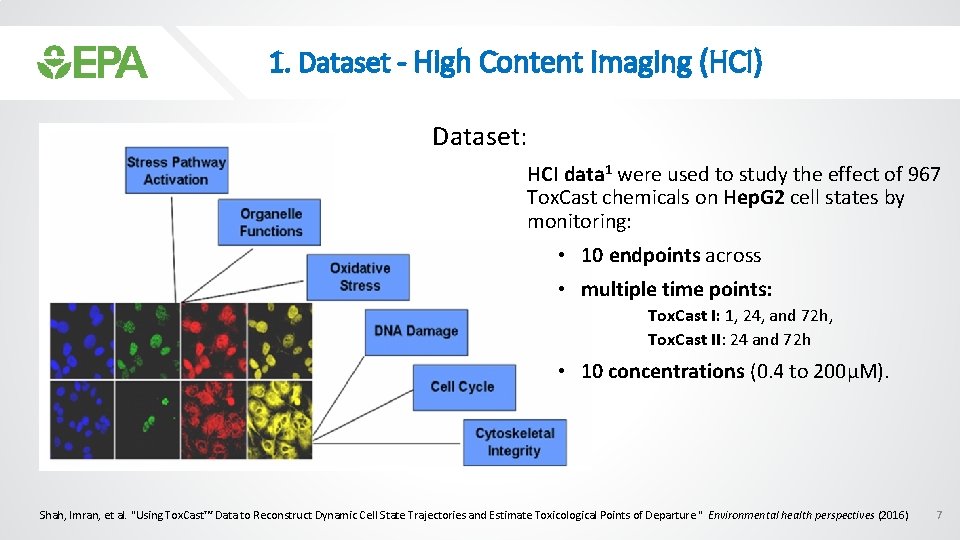
1. Dataset - High Content Imaging (HCI) Dataset: HCI data 1 were used to study the effect of 967 Tox. Cast chemicals on Hep. G 2 cell states by monitoring: • 10 endpoints across • multiple time points: Tox. Cast I: 1, 24, and 72 h, Tox. Cast II: 24 and 72 h • 10 concentrations (0. 4 to 200µM). Shah, Imran, et al. "Using Tox. Cast™ Data to Reconstruct Dynamic Cell State Trajectories and Estimate Toxicological Points of Departure. " Environmental health perspectives (2016) 7
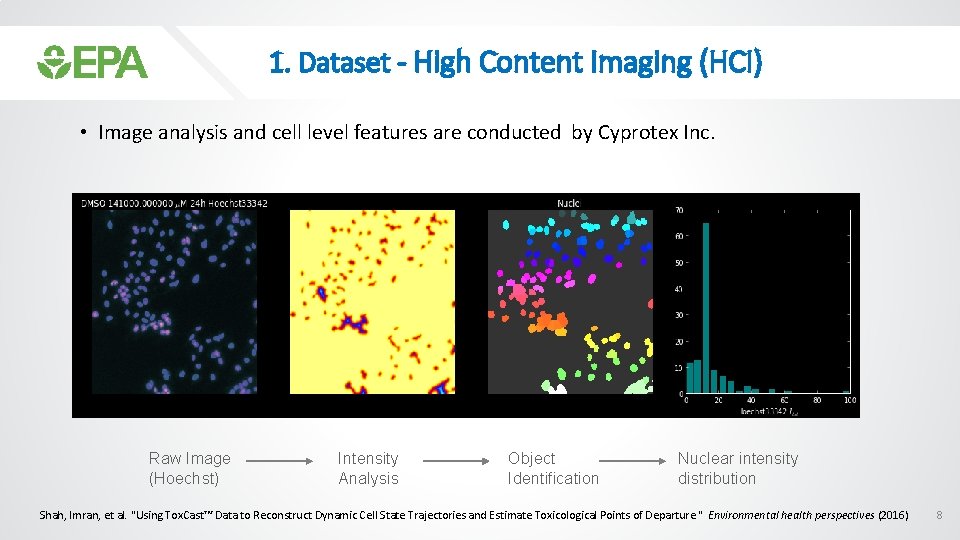
1. Dataset - High Content Imaging (HCI) • Image analysis and cell level features are conducted by Cyprotex Inc. Raw Image (Hoechst) Intensity Analysis Object Identification Nuclear intensity distribution Shah, Imran, et al. "Using Tox. Cast™ Data to Reconstruct Dynamic Cell State Trajectories and Estimate Toxicological Points of Departure. " Environmental health perspectives (2016) 8

1. Dataset - High Content Imaging (HCI) • The following cellular endpoints were quantified: 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. phosphorylated p 53 / p 53 activation (p 53), phosphorylated c-Jun/c-Jun activation (SK), phospho-Histone H 2 A. x (OS), phospho-Histone H 3 / mitotic arrest (MA), phosphorylated α-tubulin / microtubules (Mt), mitochondrial membrane potential (MMP), mitochondrial mass (MM), cell cycle arrest (CCA), nuclear size (NS), and cell number (CN). Shah, Imran, et al. "Using Tox. Cast™ Data to Reconstruct Dynamic Cell State Trajectories and Estimate Toxicological Points of Departure. " Environmental health perspectives (2016) 9
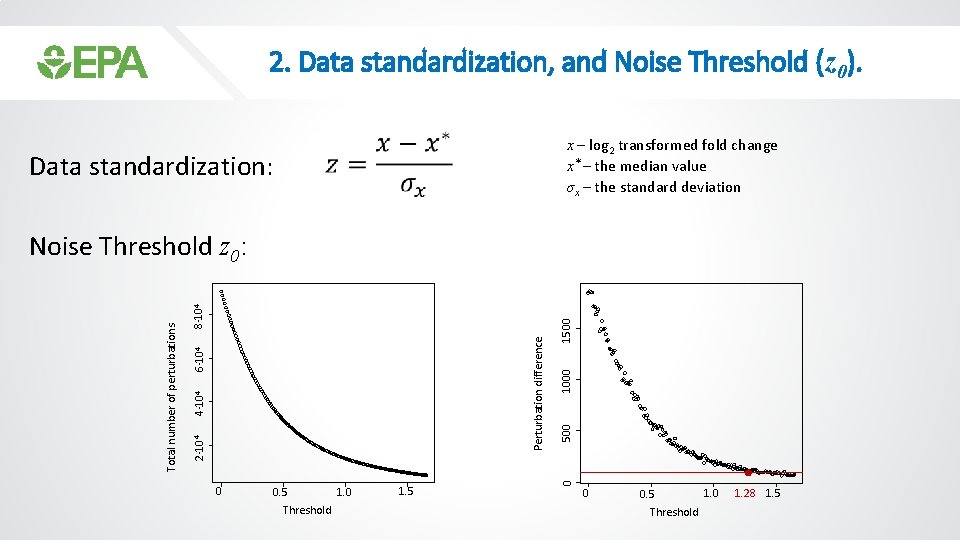
2. Data standardization, and Noise Threshold (z 0). Data standardization: • x – log 2 transformed fold change x* – the median value σx – the standard deviation 0 1. 0 0. 5 Threshold 1. 5 1500 1000 500 0 Perturbation difference 8· 104 6· 104 4· 104 2· 104 Total number of perturbations Noise Threshold z 0: 0 1. 0 0. 5 Threshold 1. 28 1. 5

3. Data Discretization. z score Motivation: Increase of p 53 causes decrease in OS, CCA or CN 11
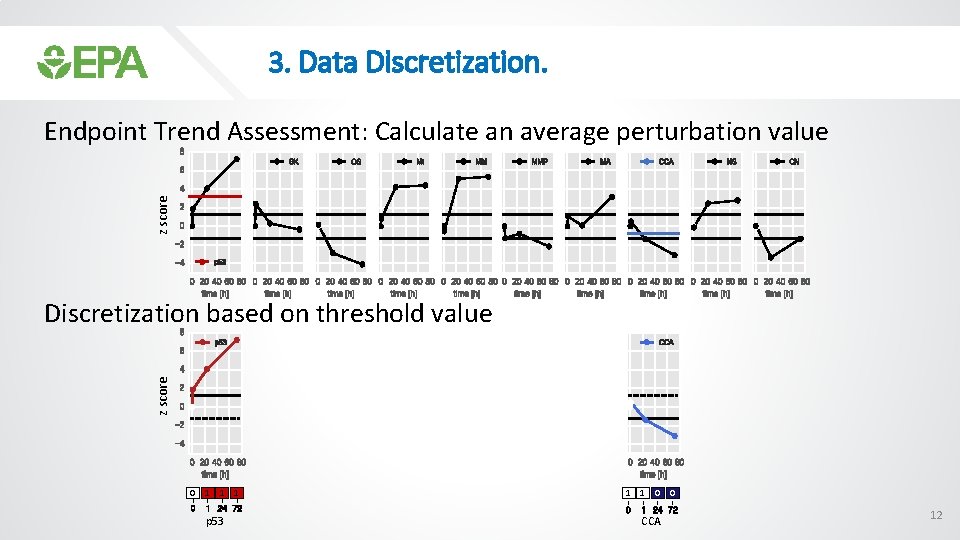
3. Data Discretization. z score Endpoint Trend Assessment: Calculate an average perturbation value z score Discretization based on threshold value 0 1 1 1 0 0 p 53 CCA 12
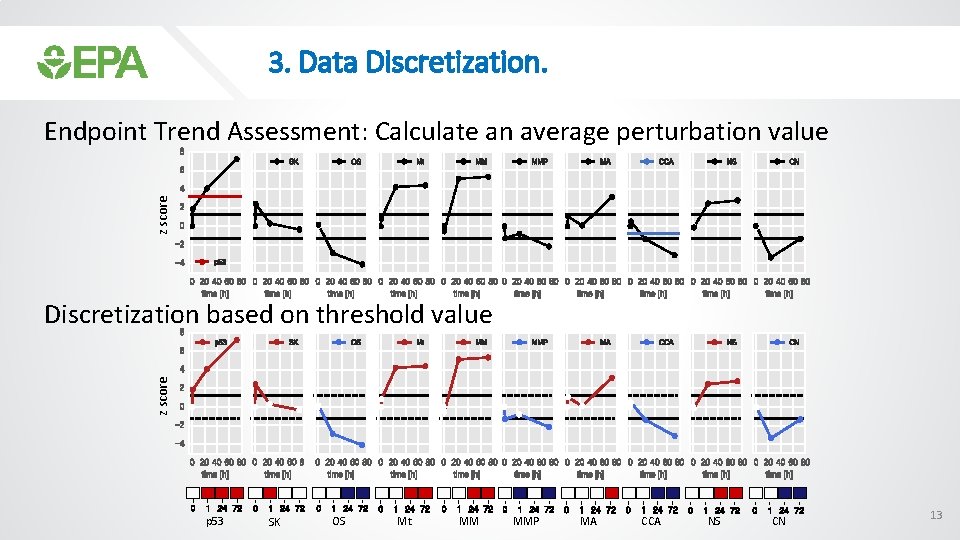
3. Data Discretization. z score Endpoint Trend Assessment: Calculate an average perturbation value z score Discretization based on threshold value p 53 SK OS Mt MM MMP MA CCA NS CN 13

4. Learning Boolean Functions and Construction of Boolean Networks (BNs). CN NS CCA MA MMP MM Mt SK OS p 53 time [h] 14
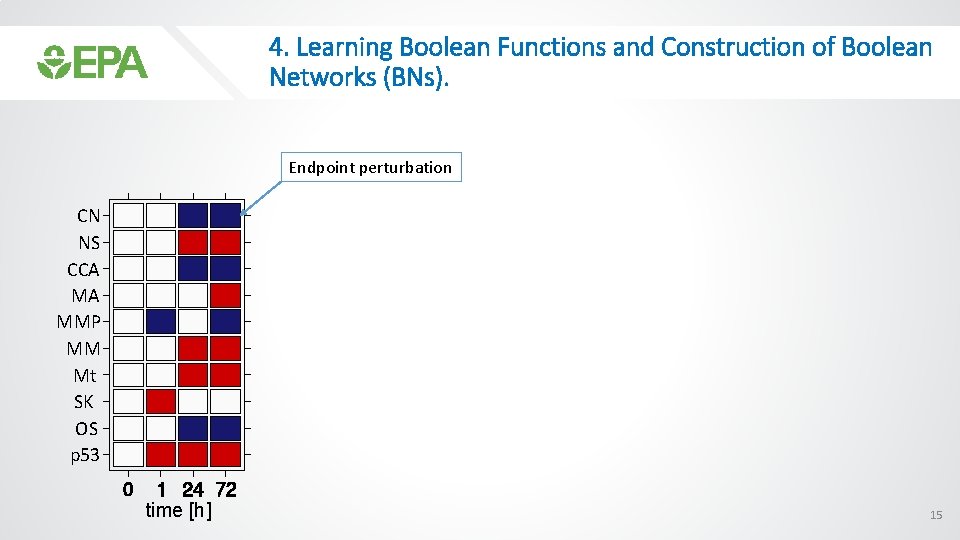
4. Learning Boolean Functions and Construction of Boolean Networks (BNs). Endpoint perturbation CN NS CCA MA MMP MM Mt SK OS p 53 time [h] 15

4. Learning Boolean Functions and Construction of Boolean Networks (BNs). State at 1 h CN NS CCA MA MMP MM Mt SK OS p 53 time [h] 16

4. Learning Boolean Functions and Construction of Boolean Networks (BNs). Discretized Trajectory of Hep. G 2 after application of Butachlor at 200 u. M State at 1 h CN NS CCA MA MMP MM Mt SK OS p 53 time [h] 17

4. Learning Boolean Functions and Construction of Boolean Networks (BNs). Discretized Trajectory of Hep. G 2 after application of Butachlor at 200 u. M State at 1 h CN NS CCA MA MMP MM Mt SK OS p 53 Find Boolean Functions “Best-Fit Extension” time [h] FCN ϵ [f 1 CN, …, f i. CN ] FNS ϵ [f 1 NS, …, f j. NS] FCCA ϵ [f 1 CCA, …, f k. CCA ] FMA ϵ [f 1 MA, …, f l. MA ] FMMP ϵ [f 1 MMP, …, f m. MMP ] FMM ϵ [f 1 MM, …, f n. MM ] FMt ϵ [f 1 Mt, …, f o. Mt] FSK ϵ [f 1 SK, …, f p. SK ] FOS ϵ [f 1 OS, …, f q OS] Fp 53 ϵ [f 1 p 53, …, f rp 53 ] 18
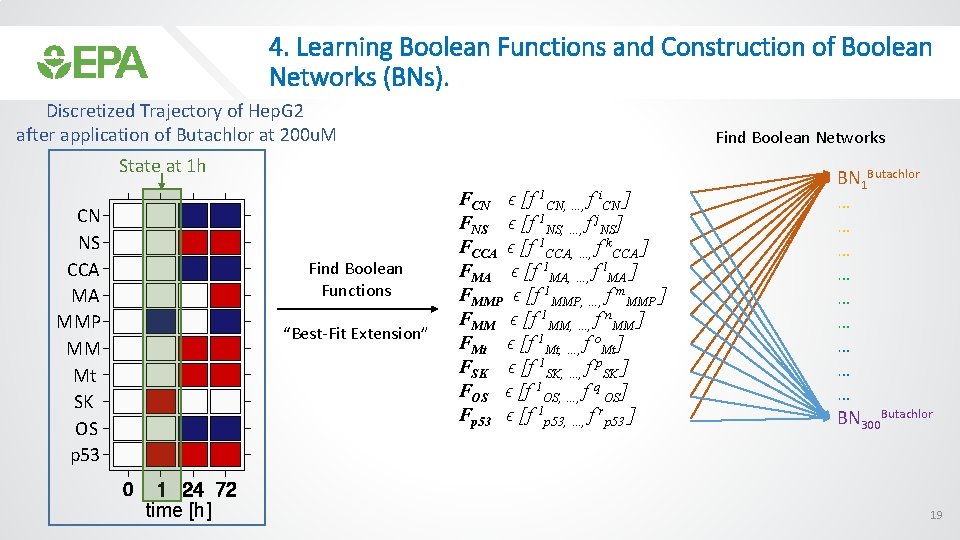
4. Learning Boolean Functions and Construction of Boolean Networks (BNs). Discretized Trajectory of Hep. G 2 after application of Butachlor at 200 u. M Find Boolean Networks State at 1 h CN NS CCA MA MMP MM Mt SK OS p 53 Find Boolean Functions “Best-Fit Extension” time [h] FCN ϵ [f 1 CN, …, f i. CN ] FNS ϵ [f 1 NS, …, f j. NS] FCCA ϵ [f 1 CCA, …, f k. CCA ] FMA ϵ [f 1 MA, …, f l. MA ] FMMP ϵ [f 1 MMP, …, f m. MMP ] FMM ϵ [f 1 MM, …, f n. MM ] FMt ϵ [f 1 Mt, …, f o. Mt] FSK ϵ [f 1 SK, …, f p. SK ] FOS ϵ [f 1 OS, …, f q OS] Fp 53 ϵ [f 1 p 53, …, f rp 53 ] BN 1 Butachlor … … … … … BN 300 Butachlor 19

5. Needleman–Wunsch optimal global alignment, and Error Estimation Error in BN prediction was estimated as the sum of the Hamming distances* between observed and predicted discretized trajectories. * The Hamming distance between two states is the number of positions at which the states are different 20

6. Coverage I. Error Estimation is performed: 1. For each trajectory – During this step we split BNs with the lowest error (“the baseline error”) from BNs with higher error. 2. Across all trajectories – During this analysis we estimated the number of trajectories predicted by each BN with an accuracy ≤ to the baseline error (“coverage”). 1 – BN covers traj. 0 – BN does not cover traj. BN 1 Coverage = 4 traj. BN 2 Coverage = 2 traj. BN 3 Coverage = 2 traj. II. The smallest set of BNs that covers all trajectories was inferred by selecting BNs with the largest coverage. 21

Outline I. Introduction II. Methods: 1. 2. 3. 4. 5. 6. Dataset Data standardization, and Noise Threshold (z 0). Data Discretization. Learning Boolean Functions and Construction of Boolean Networks (BNs). Needleman-Wunsch (NW) optimal global alignment, and Error Estimation. Coverage. III. Results: 1. Discretized Trajectories and Total Perturbation. 2. Clustering of discretized trajectories, Error Estimation, and Coverage (first 10 BNs). 3. Learned BNs in case of Butachlor. IV. Summary 22
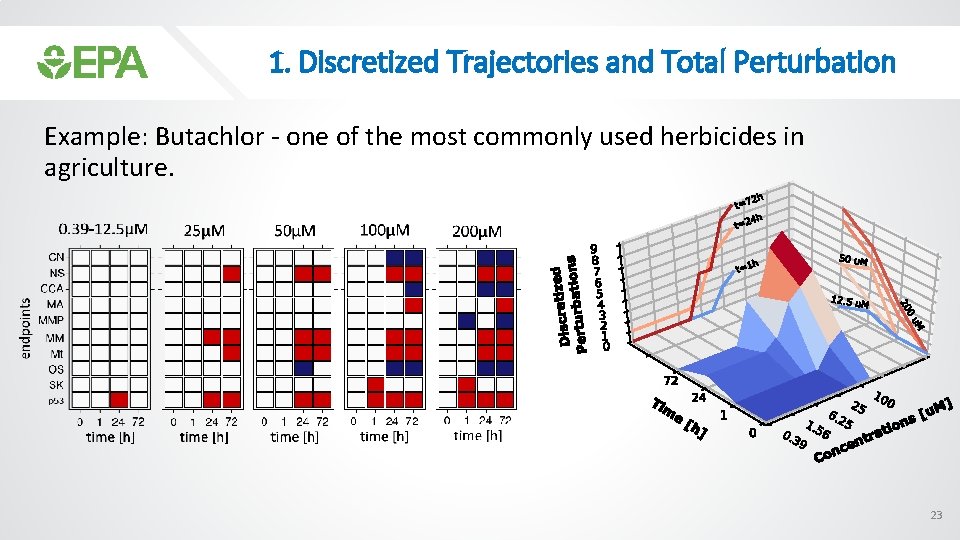
1. Discretized Trajectories and Total Perturbation Example: Butachlor - one of the most commonly used herbicides in agriculture. 23
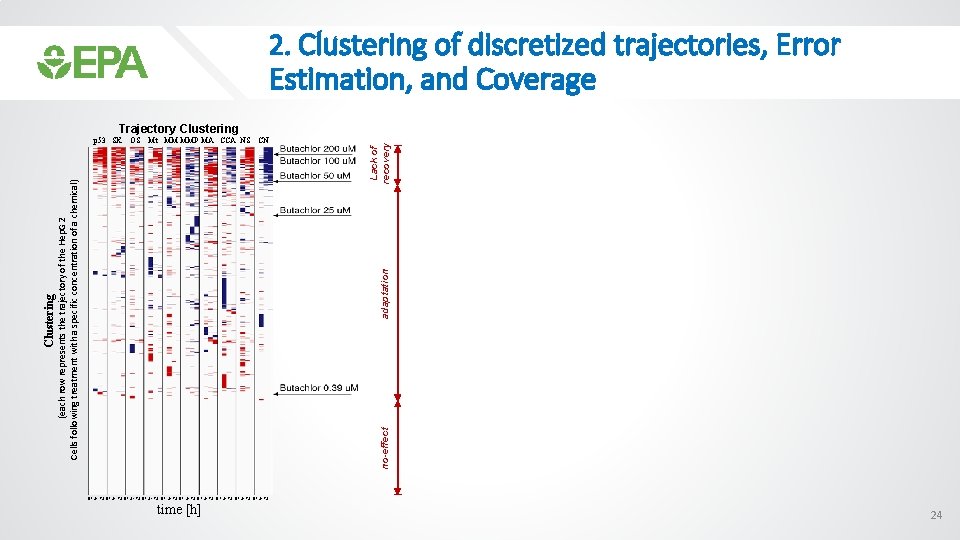
2. Clustering of discretized trajectories, Error Estimation, and Coverage adaptation no-effect Clustering (each row represents the trajectory of the Hep. G 2 Cells following treatment with a specific concentration of a chemical) OS Mt MM MMP MA CCA NS CN Lack of recovery Trajectory Clustering p 53 SK 0 1 24 72 0 1 24 72 0 1 24 72 time [h] 24
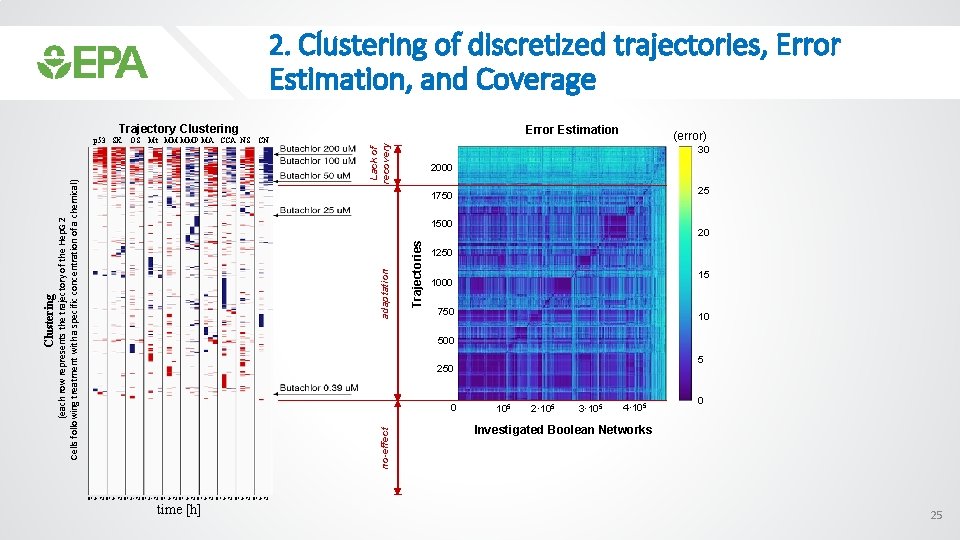
2. Clustering of discretized trajectories, Error Estimation, and Coverage Trajectory Clustering Lack of recovery Error Estimation (error) 30 2000 25 1750 Trajectories adaptation 1500 20 1250 15 1000 750 10 500 5 250 0 no-effect Clustering OS Mt MM MMP MA CCA NS CN (each row represents the trajectory of the Hep. G 2 Cells following treatment with a specific concentration of a chemical) p 53 SK 105 2· 105 3· 105 4· 105 0 Investigated Boolean Networks 0 1 24 72 0 1 24 72 0 1 24 72 time [h] 25
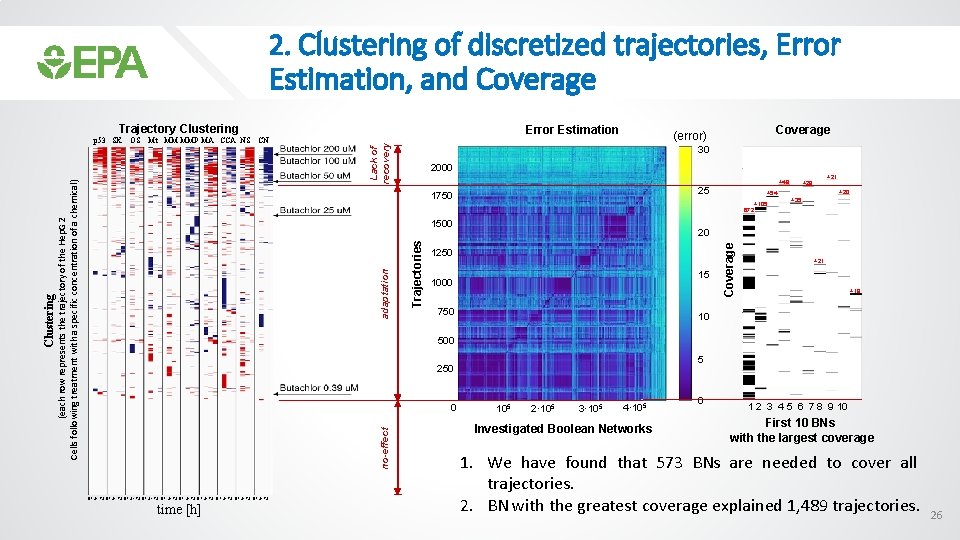
2. Clustering of discretized trajectories, Error Estimation, and Coverage Trajectory Clustering Lack of recovery Error Estimation Coverage (error) 30 2000 +48 25 1750 +105 +20 +35 20 1250 15 1000 750 Coverage Trajectories adaptation +21 +28 +54 672 1500 +21 +18 10 500 5 250 0 no-effect Clustering OS Mt MM MMP MA CCA NS CN (each row represents the trajectory of the Hep. G 2 Cells following treatment with a specific concentration of a chemical) p 53 SK 0 1 24 72 0 1 24 72 0 1 24 72 time [h] 105 2· 105 3· 105 4· 105 Investigated Boolean Networks 0 1 2 3 4 5 6 7 8 9 10 First 10 BNs with the largest coverage 1. We have found that 573 BNs are needed to cover all trajectories. 2. BN with the greatest coverage explained 1, 489 trajectories. 26

3. Learned BNs in case of Butachlor 200µM CN NS not and or CCA MA MMP MM Mt OS SK p 53 0 1 24 72 time [h] 27
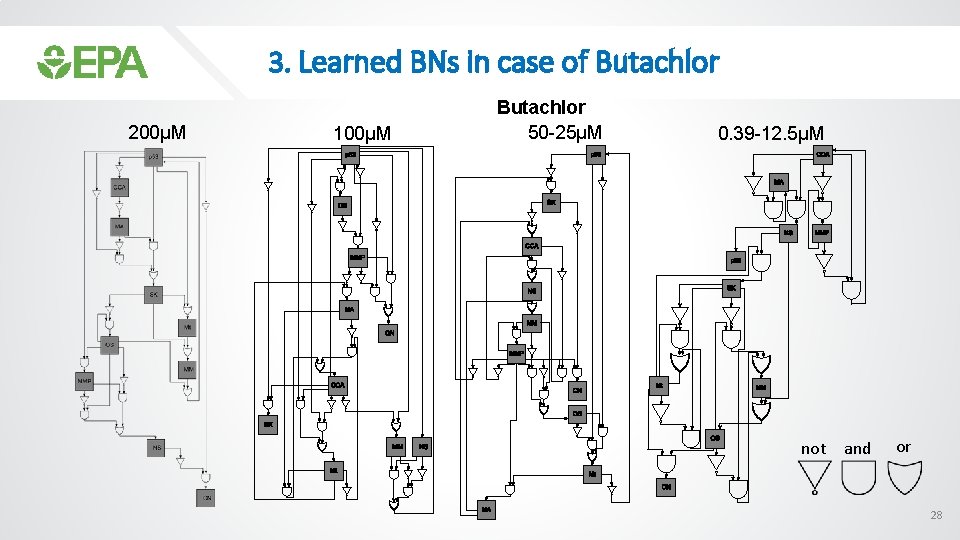
3. Learned BNs in case of Butachlor 200µM 100µM Butachlor 50 -25µM 0. 39 -12. 5µM not and or 28

Outline I. Introduction II. Methods: 1. 2. 3. 4. 5. 6. Dataset Data standardization, and Noise Threshold (z 0). Data Discretization. Learning Boolean Functions and Construction of Boolean Networks (BNs). Needleman-Wunsch (NW) optimal global alignment, and Error Estimation. Coverage. III. Results: 1. Discretized Trajectories and Total Perturbation. 2. Clustering of discretized trajectories, Error Estimation, and Coverage (first 10 BNs). 3. Learned BNs in case of Butachlor. IV. Summary 29

IV. Summary 1. Response of Hep. G 2 cells to concentration dependent chemical treatment shows three temporal trends: 1) no-effect, 2) adaptation, and 3) lack of recovery. 2. We have found that 573 BNs are needed to cover all trajectories. 3. BN with the greatest coverage explained 1, 489 trajectories. These trajectories were produced by low treatment concentrations and we believe they represent cellular recovery processes. 4. Trajectories produced by high concentration treatments, that resulted in cell death, were predicted by a different set of BNs. 5. Our findings illustrate the utility of BNs that differentiate cellular programs involved in adaptation versus injury. 30
Thank you 31
- Slides: 31