DNA Computing By Thierry Metais Email metaisenst fr

DNA Computing By Thierry Metais Email: metais@enst. fr
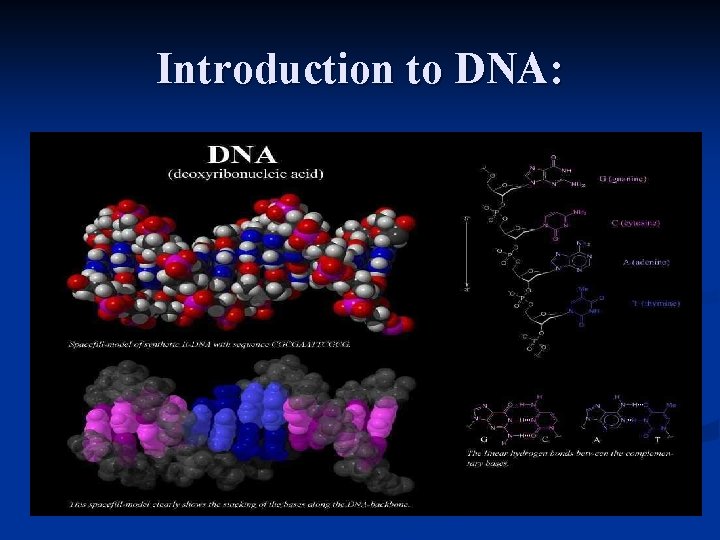
Introduction to DNA: n The life’s molecule:

Introduction: n What is DNA computing ? Around 1950 first idea (precursor Feynman) n First important experiment 1994: Leonard Adleman n Molecular level (just greater than 10 -9 meter) n Massive parallelism. n In a liter of water, with only 5 grams of DNA we get around 1021 bases ! n Each DNA strand represents a processor ! n

A bit of biology n n The DNA is a double stranded molecule. Each strand is based on 4 bases: n n n Adenine (A) Thymine (T) Cytosine (C) Guanine (G) Those bases are linked through a sugar (desoxyribose) IMPORTANT: n n The linkage between bases has a direction. There are complementarities between bases (Watson-Crick). (A) (T) (C) (G)

DNA manipulations: n n If we want to use DNA as an information bulk, we must be able to manipulate it. However we are talking of handling molecules… ENZYMES = Natural CATALYSERS. So instead of using physical processes, we would have to use natural ones, more effective: n n for lengthening: polymerases… for cutting: nucleases (exo/endo-nucleases)… for linking: ligases… Serialization: 1985: Kary Mullis PCR Thank this reaction we get millions of identical strands, and we are allowed to think of massive parallel computing.

And what now ? n Situation: Molecular level. n Lots of “agents”. (strands) n Tools provided by nature. (enzymes) n n How can we use all this? If there is a utility …

Coding the information: 1994: THE Adleman’s experiment. Given a directed graph can we find an hamiltonian path (more complex than the TSP). In this experiment there are 2 keywords: massive parallelism (all possibilities are generated) complementarity (to encode the information) n § This experiment proved that DNA computing wasn’t just a theoretical study but could be applied to real problems like cryptanalysis (breaking DES ).

Adleman experiment: n n n Each node is coded randomly with 20 bases. Let Si be a code, h be the complementarity mapping. h(ATCG) = TAGC. Each Si is decomposed into 2 sub strands of length 10: Si = Si’’ Edge(i, j) will be encode as h(Si’’Sj’) ( preserve edge orientation). Code: n n n Input(N) //All vertices and edges are mixed, Nature is working N B(N, S 0) //S 0 was chosen as input vertice. N E(N, S 4) //S 4 was chosen as output vertice. N E(N, <=140) // due to the size of the coding. For i=1 to 5 do N +N(N, Si) //Testing if hamiltonian path Detect(N) //conclusion …
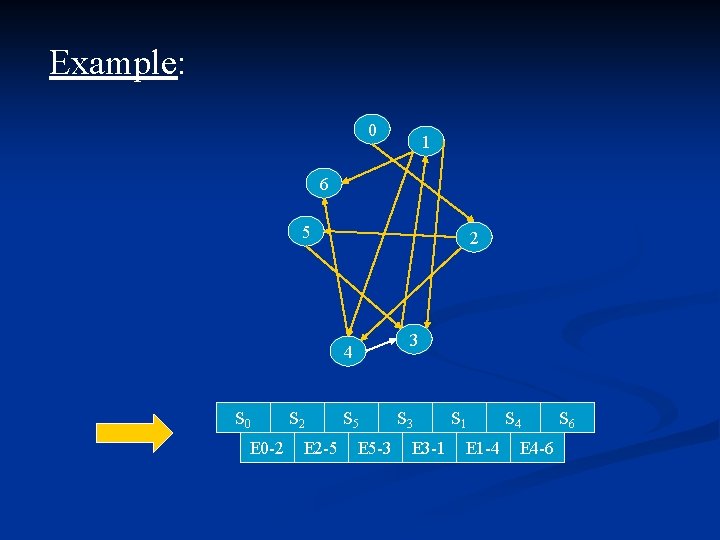
Example: 0 1 6 5 2 3 4 S 0 E 0 -2 S 2 E 2 -5 S 5 E 5 -3 S 3 E 3 -1 S 1 E 1 -4 S 4 E 4 -6 S 6
![New generation of computers? In the second part of [1], it is proven through New generation of computers? In the second part of [1], it is proven through](http://slidetodoc.com/presentation_image/3c0212668c87f52c3d41fbe2b2198b95/image-10.jpg)
New generation of computers? In the second part of [1], it is proven through language theory that DNA computing “guarantees universal computations”. n Many architectures have been invented for DNA computations. n The Adleman experiment is not the single application case of DNA computing… n

Stickers model: Memory complex = Strand of DNA (single or semi-double). n Stickers are segments of DNA, that are composed of a certain number of DNA bases. n To use correctly the stickers model, each sticker must be able to anneal only at a specific place in the memory complex. n
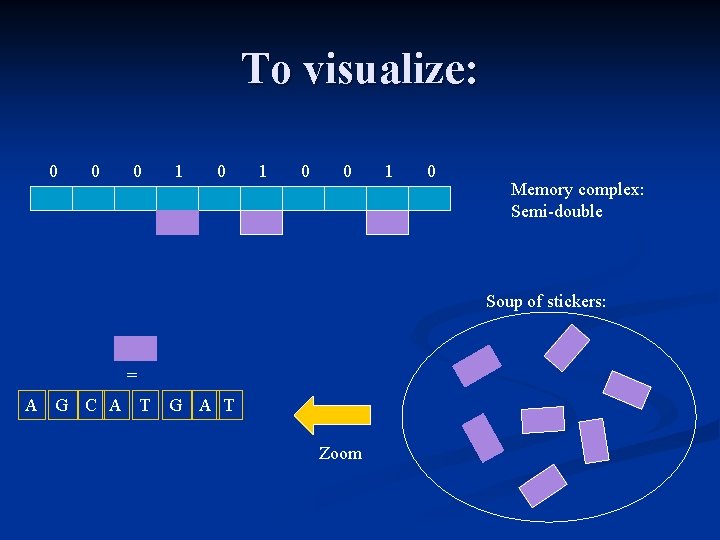
To visualize: 0 0 0 1 0 Memory complex: Semi-double Soup of stickers: = A G C A T G A T Zoom

About a stickers machine? Simple operations: merge, select, detect, clean. n Tubes are considered (cylinders with two entries) n However for a mere computation (DES): n Great number of tubes is needed (1000). n Huge amount of DNA needed as well. n n Practically no such machine has been created…. Too much engineering issues.

Why don’t we see DNA computers everywhere? n DNA computing has wonderful possibilities: Reducing the time of computations* (parallelism) n Dynamic programming ! n However one important issue is to find “the killer application”. n Great hurdles to overcome… n

Some hurdles: n Operations done manually in the lab. n Natural tools are what they are… Formation of a library (statistic way) Operations problems

Conclusion: The paradigm of DNA computing has lead to a very important theoretical research. n However DNA computers won’t flourish soon in our daily environment due to the technologic issues. n Adleman renouncement toward electronic computing. n Is all this work lost ? n NO ! “Wet computing” n

Bibliography: n DNA Computing, New Computing Paradigms. Gheorghe Paun, Grzegorz Rozenberg, Arto Salomaa n DIMACS: DNA based computers n Reducing Errors in DNA Computing by Appropriate Word Design. wdesign. pdf

Links: http: //www. cs. wayne. edu/~kjz/KPZ/Natural. Co mputing. html n http: //dna 2 z. com/dnacpu/dna. html n http: //www. intermonde. net/adn/liens. html n
- Slides: 18