Bioinformatics for comparative quantitative LCMS2E proteomics data analysis

Bioinformatics for comparative quantitative LC-MS(2/E) proteomics data analysis Joost de Groot, Twan America, Roeland van Ham

Introduction n Joost de Groot n Scientific Software Developer n Wageningen University and Research (WUR)
Introduction

Introduction n WUR = WU + R R=Research n n DLO (research institutes) Plant Research International Bio. Science (bu) Applied Bio. Informatics (clust)

Introduction The Bio. Science: High-throughput analyses of DNA, RNA, proteins and metabolites Genome analyses and bioinformatics Research on bioactive and health promoting compounds Investigate the plant as factory, e. g. for the production of pharmaceutical proteins Perform research on stress biology Explore quality traits of plants, such as taste, flavour, insect resistance plant architecture

Introduction n Bio. Science is (among others) involved in

Introduction n I am involved in Bioinformatics for Proteomics n Bioinformatics for label-free comparative quantitative LC-MS(2/E) proteomics data analysis

Introduction n n Data from Waters Q-TOF, Synapt MS systems PLGS software data acquisition/processing + other software (e. g. Mascot, Progenesis) We focus on post alignment data quality control and data quality improvement Several Proteomics experiments l l Differential protein expression in fungi infected plants Allergens in mother’s milk Apoplast protein identification etc
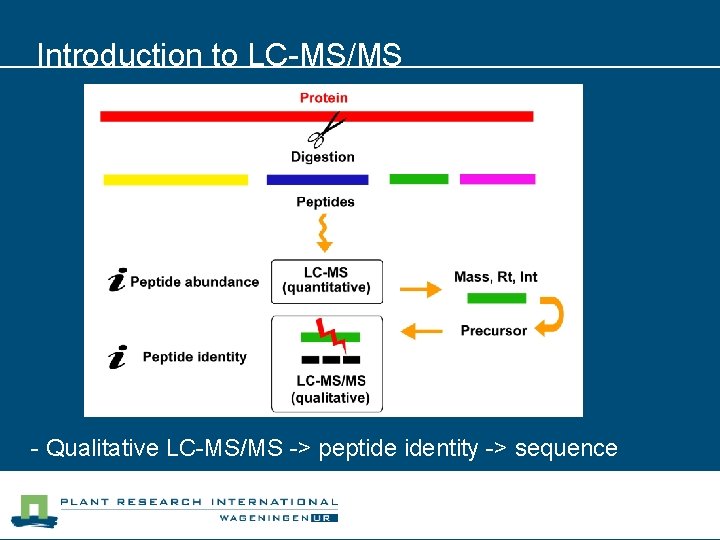
Introduction to LC-MS/MS - Qualitative LC-MS/MS -> peptide identity -> sequence
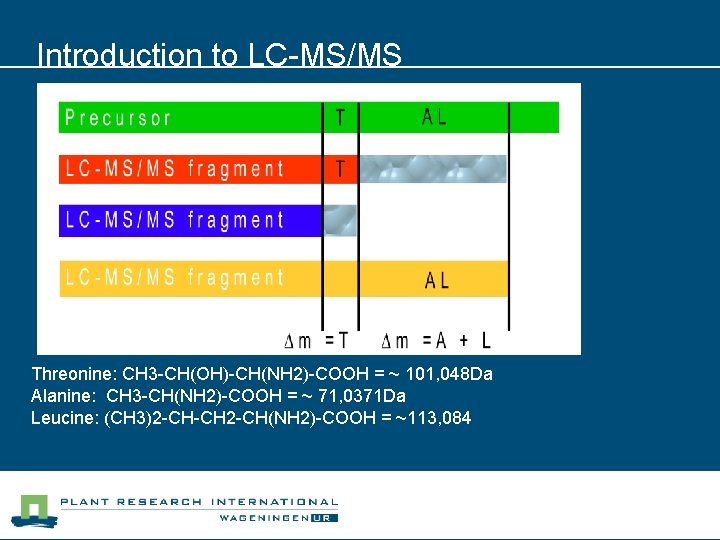
Introduction to LC-MS/MS Threonine: CH 3 -CH(OH)-CH(NH 2)-COOH = ~ 101, 048 Da Alanine: CH 3 -CH(NH 2)-COOH = ~ 71, 0371 Da Leucine: (CH 3)2 -CH-CH 2 -CH(NH 2)-COOH = ~113, 084
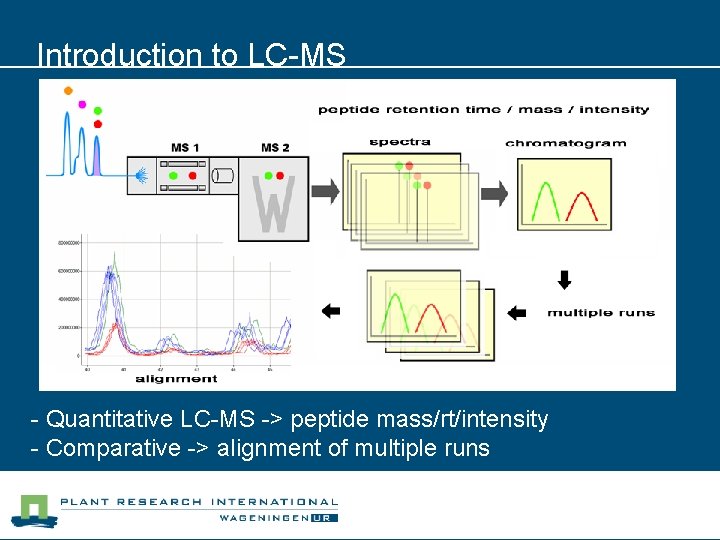
Introduction to LC-MS - Quantitative LC-MS -> peptide mass/rt/intensity - Comparative -> alignment of multiple runs

Introduction to LC-MS (what is (was: ) the problem? ) l l This simplified example shows one peak in three runs (replicates) of a single sample. Chromatogram of a single peptide (present in every replicate). Problem: data processing software can make ‘mistakes’ at peak detection. Result: split peaks. Peaks of high abundant peptides or tailing peaks are prone to fragmentation.

History (how I’ve got involved) n 2006/2007 CBSG Ind 3 bottleneck project l l Bioinformatics solutions for urgent issues in comparative quantitative proteomics data analysis (Twan America). Highest priority: • Solve LC-MS peak detection fragmentation over multiple chromatograms (which needs some explanation I guess )
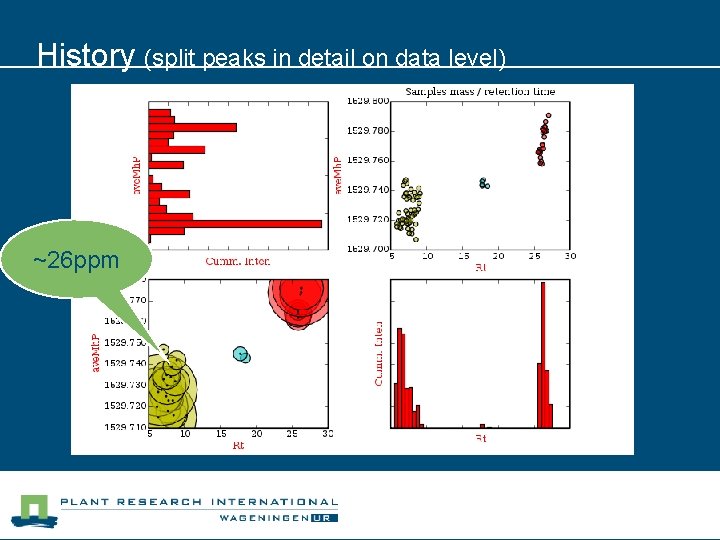
History (split peaks in detail on data level) ~26 ppm
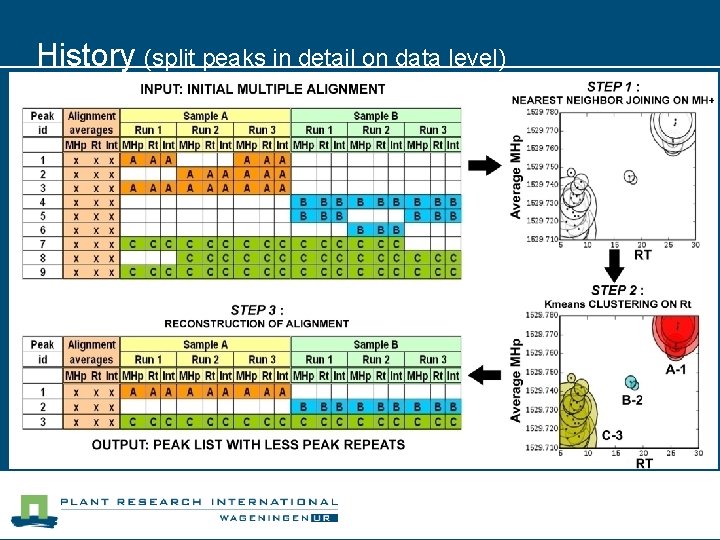
History (split peaks in detail on data level) - Quantitative -> peptide mass/rt/intensity - Comparative -> multiple samples = runs
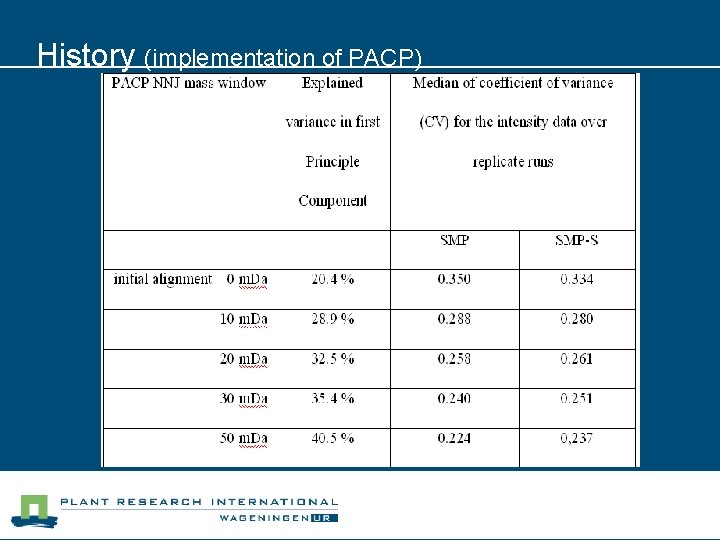
History (implementation of PACP)

History (implementation of PACP)

History (PACP) n Procedure published in Proteomics n Post alignment clustering procedure for comparative quantitative proteomics LC-MS Data. De Groot, JC et. al. Proteomics 2008 V 8#1. p 32 -36

Future n n We applied for additional Bioinformatics for Proteomics funding (Twan America (supervisor) and Joost de Groot (bioinformatics developer)). Granted: l CBSG 2 BB 6 project: • Scientific programmer 2 year (~0, 5 fte = ~0, 25 fte/y) l NBIC/NPC/BIOASSIST/NGI = NBPP (Netherlands Bioinformatics for Proteomics Platform) • Scientific programmer 2 year (~1 fte = ~ 0, 5 fte/y)
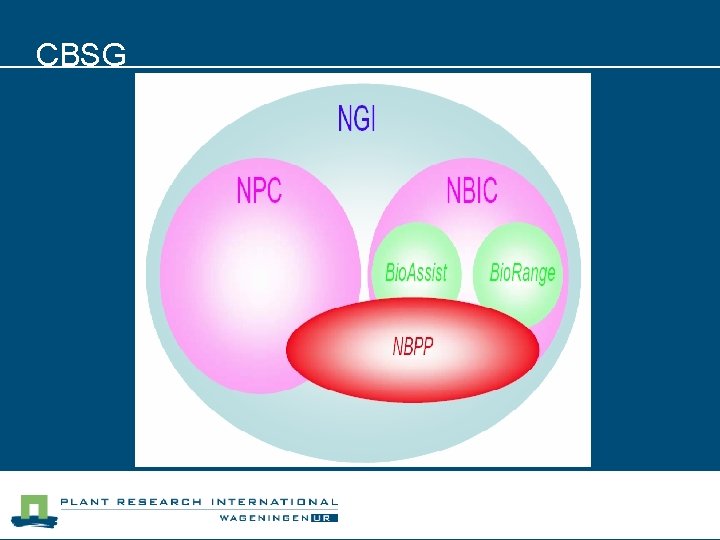
CBSG
NBPP
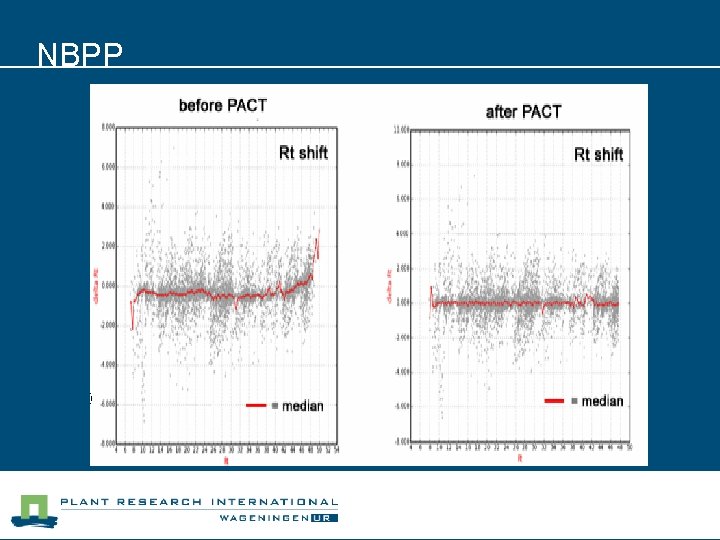
NBPP

Issues to address n CBSG BB 6 l Retention time correction of LC-MS results. • Several effects can cause (small) drifts in retention time which can result in less accurate alignments. • PACP and SEDMAT results expect to be improved by Rt correction methods. l Solution: retention time correction algorithm.

Issues to address (CBSG BB 6)
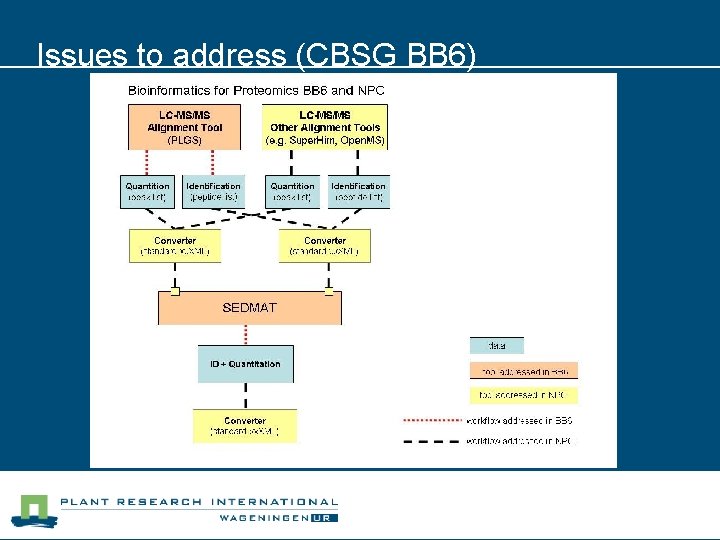
Issues to address (CBSG BB 6)

Issues to address n NBPP (Bio. Assist) l Make tools available (via webservices) • Wrap tools in web services • Enables workflow management systems (like Taverna) • Re-engineer PACP (Python ->Java WS) l Solution: build web service providers/consumers
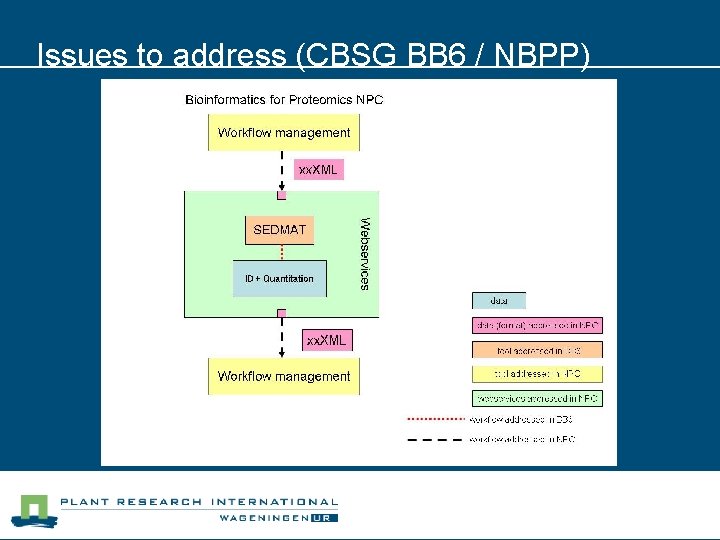
Issues to address (CBSG BB 6 / NBPP)
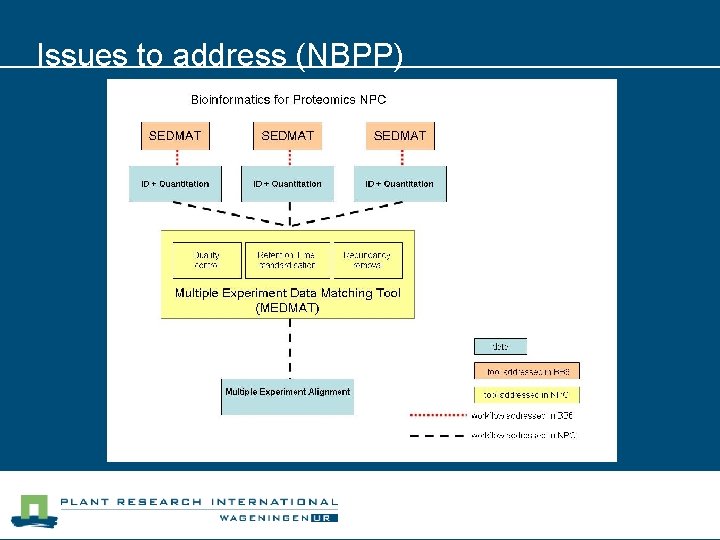
Issues to address (NBPP)
Issues (NBIC/NPC)
Netbeans / Java n SEDMAT
Glass. Fish Application Server

Thanks for your attention. Feel free to give comments, remarks or suggestions. © Wageningen UR
- Slides: 32