Bioinformatics for Proteomics studies Tamanna Sultana Bioinformatics Analysis

Bioinformatics for Proteomics studies Tamanna Sultana Bioinformatics Analysis Core (BAC) Genomics & Proteomics Core Laboratories (GPCL) University of Pittsburgh

Proteomics • The term proteomics refers but not limited to the analysis of proteins in terms of separation, identification, quantification, expression and function.

Bioinformatics for proteomics • Informatics is a field of study that focuses on the use of technology for improving access to and utilization of information. . library. ahima. org/xpedio/groups/public/documents/ahima /bok 1_025042. hcsp • Information science: – the sciences concerned with gathering, manipulating, storing, retrieving, and classifying recorded information wordnetweb. princeton. edu/perl/webwn – its broad meaning is the science of processing data. Within health and social care, it is used to refer to the processing of data on patients and clients, normally – but by no means exclusively – through IT systems. www. smarthealthcare. com/glossary
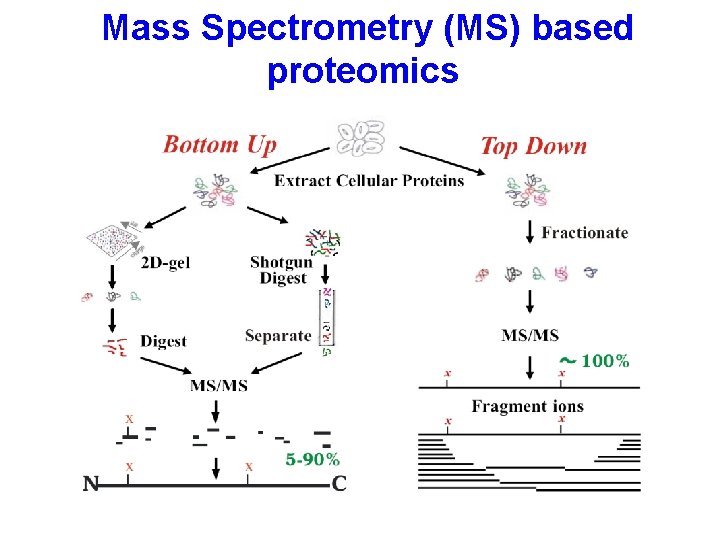
Mass Spectrometry (MS) based proteomics
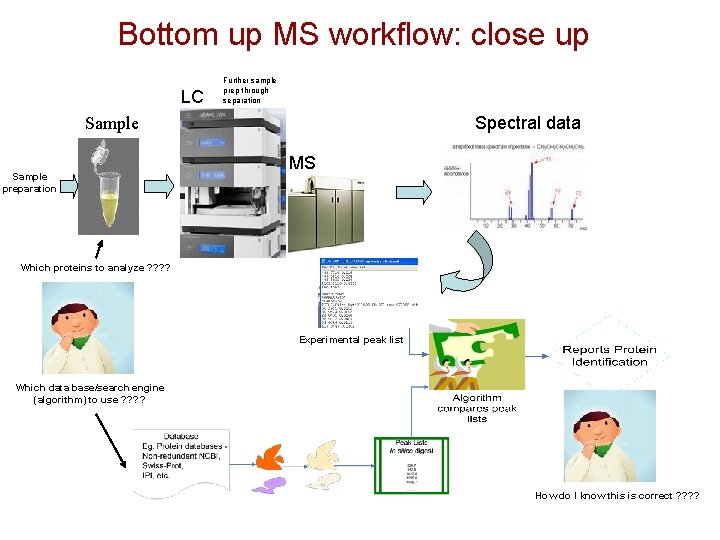
Bottom up MS workflow: close up LC Further sample prep through separation Spectral data Sample preparation MS Which proteins to analyze ? ? Experimental peak list Which data base/search engine (algorithm) to use ? ? How do I know this is correct ? ?
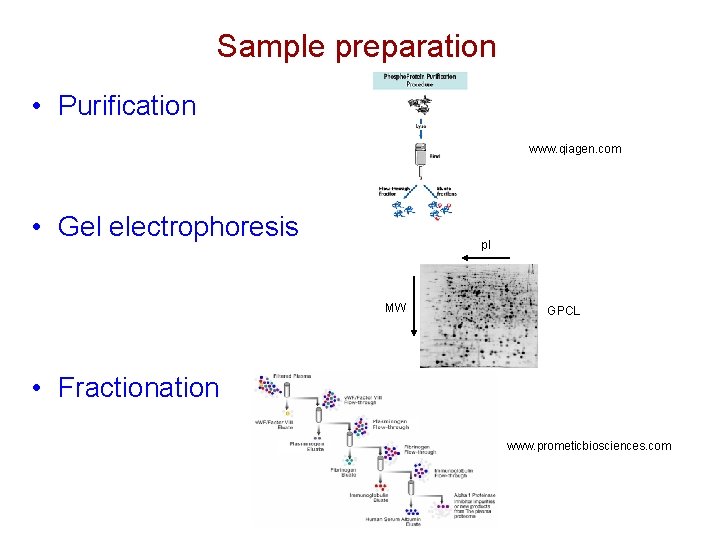
Sample preparation • Purification www. qiagen. com • Gel electrophoresis p. I MW GPCL • Fractionation www. prometicbiosciences. com

Sample analysis by Bottom up MS • Digestion (cleavage) of proteins by an enzyme MGLSDGEWQQ VLNVWGKVEA DIAGHGQEVL IRLFTGHPET LEKFDKFKHL KTEAEMKASE DLKKHGTVVL TALGGILKKK GHHEAELKPL AQSHATKHKI PIKYLEFISD AIIHVLHSKH PGDFGADAQG AMTKALELFR NDIAAKYKEL GFQG • Fractionation of peptides – Off line – On line • Analysis by MS and tandem MS (MS/MS)
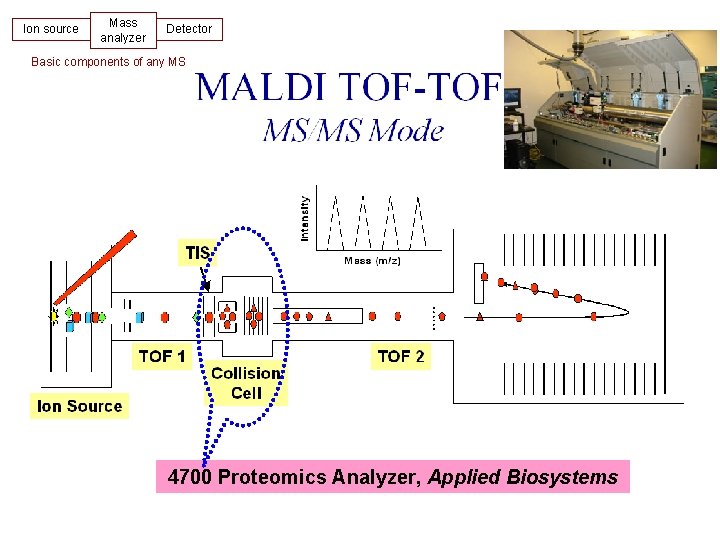
Ion source Mass analyzer Detector Basic components of any MS Tandem MS 4700 Proteomics Analyzer, Applied Biosystems
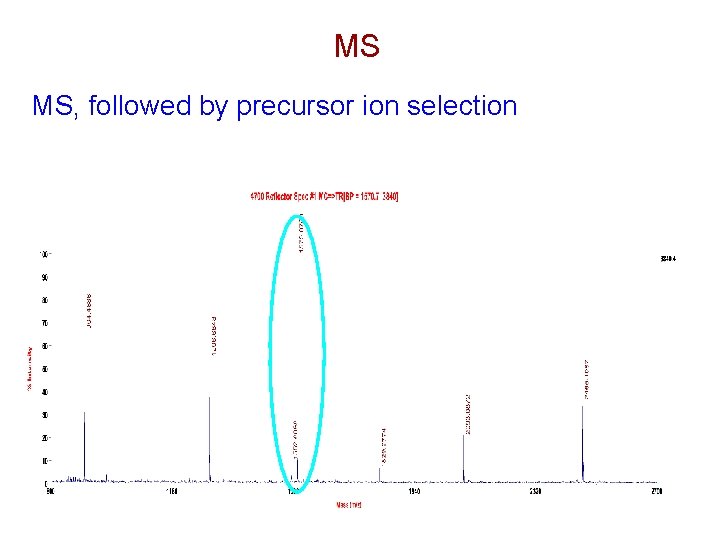
MS MS, followed by precursor ion selection
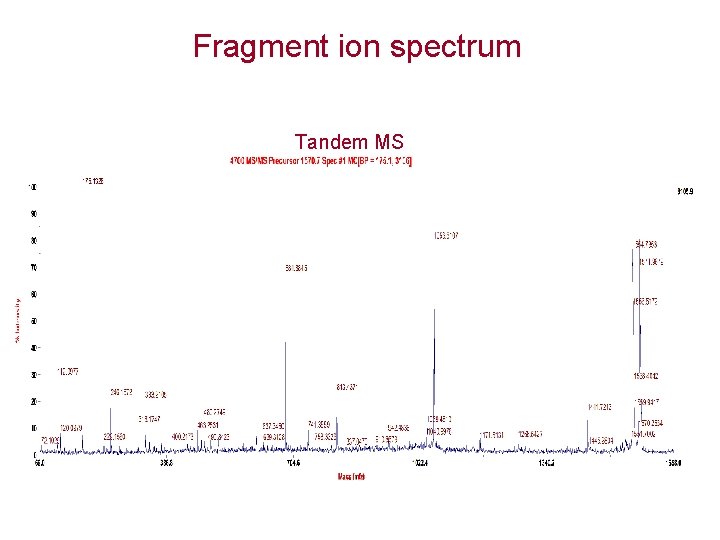
Fragment ion spectrum Tandem MS
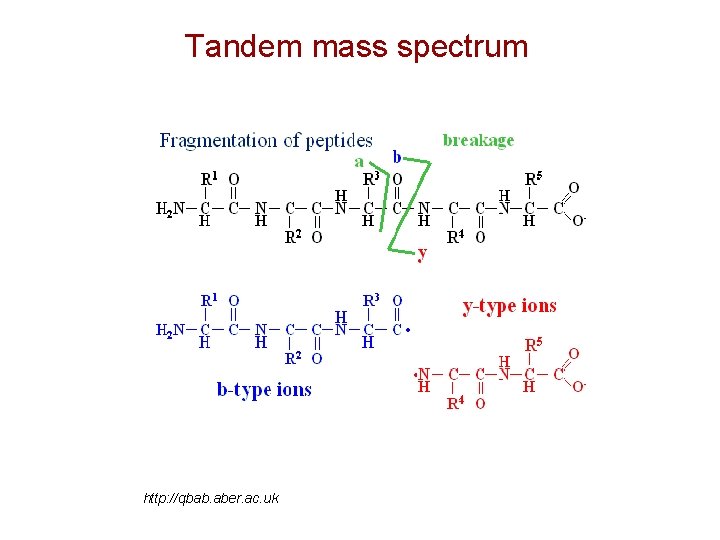
Tandem mass spectrum http: //qbab. aber. ac. uk
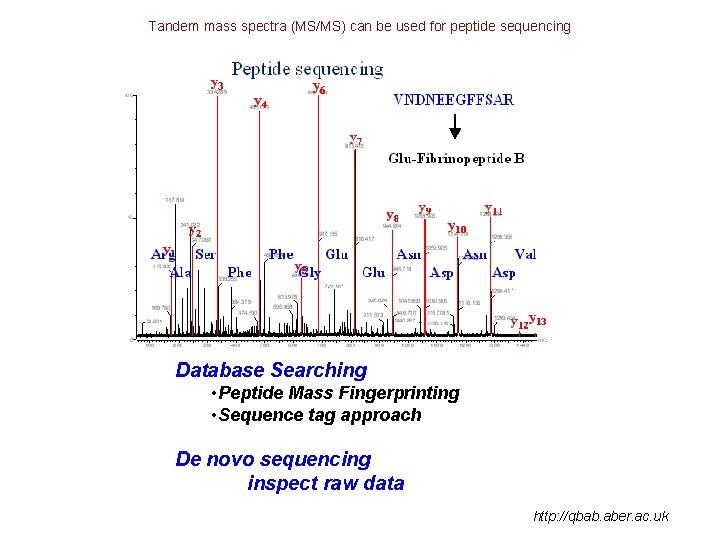
Tandem mass spectra (MS/MS) can be used for peptide sequencing Database Searching • Peptide Mass Fingerprinting • Sequence tag approach De novo sequencing inspect raw data http: //qbab. aber. ac. uk

Mascot Search Results Search title : Sample. Set. ID: 362, Analysis. ID: 567, Maldi. Well. ID: 15790, Spectrum. ID: 17225, Path=Mani102004New Analysis 1 Database : NCBInr 20040606 (1846720 sequences; 611532004 residues) Timestamp : 20 Oct 2004 at 14: 52: 50 GMT Top Score : 681 for gi|180570, creatine kinase [Homo sapiens] Probability Based Mowse Score is -10*Log(P), where P is the probability that the observed match is a random event. Protein scores greater than 75 are significant (p<0. 05).

Top hits from Mascot Search – there are multiple accession numbers for the same protein

Creatine kinase B is the highest scoring protein Match to: gi|21536286 ; Score: 681 Creatine kinase - B [Homo sapiens] Nominal mass (Mr): 42591; Calculated p. I value: 5. 34 Observed Mass & p. I: 43 kd, 6. 2 -6. 27 Sequence Coverage: 46% 1 MPFSNSHNAL KLRFPAEDEF PDLSAHNNHM AKVLTPELYA ELRAKSTPSG 51 FTLDDVIQTG VDNPGHPYIM TVGCVAGDEE SYEVFKDLFD PIIEDRHGGY 101 KPSDEHKTDL NPDNLQGGDD LDPNYVLSSR VRTGRSIRGF CLPPHCSRGE 151 RRAIEKLAVE ALSSLDGDLA GRYYALKSMT EAEQQQLIDD HFLFDKPVSP 201 LLSASGMARD WPDARGIWHN DNKTFLVWVN EEDHLRVISM QKGGNMKEVF 251 TRFCTGLTQI ETLFKSKDYE FMWNPHLGYI LTCPSNLGTG LRAGVHIKLP 301 NLGKHEKFSE VLKRLRLQKR GTGGVDTAAV GGVFDVSNAD RLGFSEVELV 351 QMVVDGVKLL IEMEQRLEQG QAIDDLMPAQ K

Problems frequently faced in proteomics • How to choose which proteins to analyze – Depends on the goal and the availability of support • Which methods to choose to quantify the proteins – Again it depends on the goal • How do I know I have all the correct proteins – Complicated – Depends on the sample, methods, instruments, software’s used

BAC’s fee for service methods for supporting proteomic studies • Differential protein expression analysis – Tradition 2 D – Di. GE • Consensus peptide identification – Protein database searches with Mascot, Sequest, X!Tandem, Phenyx etc. – Consensus searches among Mascot, Sequest and X!Tandem – Pathway analysis of identified proteins • Intelligent Systems and Bioinformatics Laboratory – Pathway Express
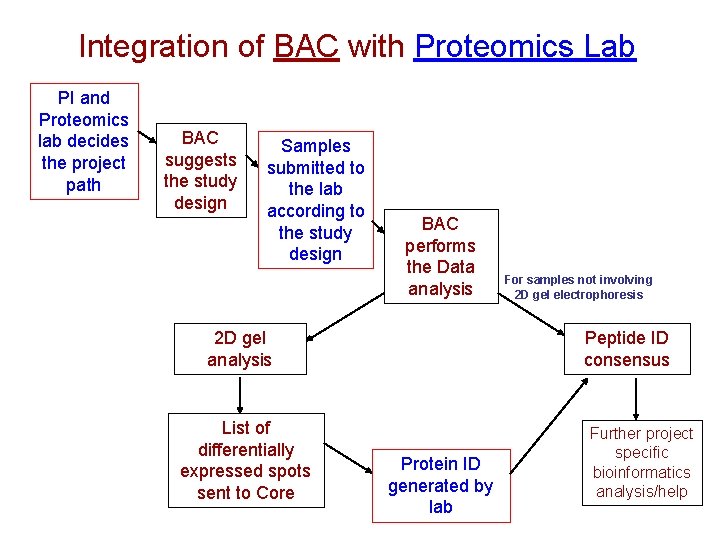
Integration of BAC with Proteomics Lab PI and Proteomics lab decides the project path BAC suggests the study design Samples submitted to the lab according to the study design BAC performs the Data analysis 2 D gel analysis List of differentially expressed spots sent to Core For samples not involving 2 D gel electrophoresis Peptide ID consensus Protein ID generated by lab Further project specific bioinformatics analysis/help
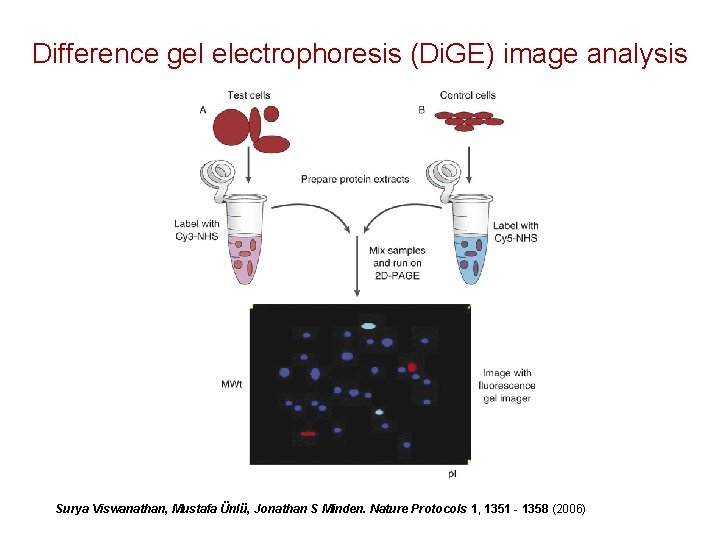
Difference gel electrophoresis (Di. GE) image analysis Surya Viswanathan, Mustafa Ünlü, Jonathan S Minden. Nature Protocols 1, 1351 - 1358 (2006)
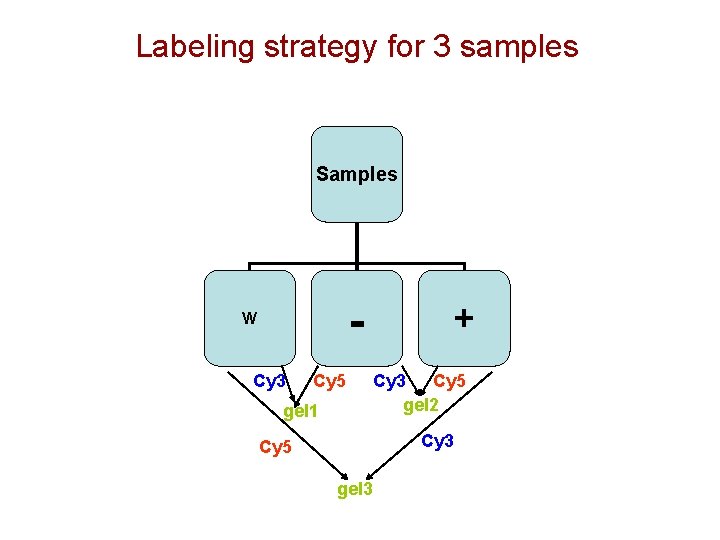
Labeling strategy for 3 samples Samples - W Cy 3 Cy 5 gel 1 + Cy 3 Cy 5 gel 2 Cy 3 Cy 5 gel 3

Di. GE analysis of protein isoform expression in STAT 3 constitutively activated versus STAT 3 loss variant multiple myeloma cell line U 266 Date: May 22, 2008 PI: ___ BAC Analyst: Tamanna Sultana Project Location: www. genetics. pitt. edu/mygpcl/ Example of Di. GE analysis and Reporting to PI

Specific aims/objectives To assess proteomic differences between U 266 with constitutively activated STAT 3 and the U 266 STAT 3 loss variant using 2 D-DIGE, with particular emphasis on mapping the CENPM 58 AA isoform

Sample Details • Samples – U 266 with constitutively activated STAT 3: 266#1 – U 266 STAT 3 loss variant: 266#2 • Sample condition – Protein extracted and cleaned by PI lab • Sample amount – 100 µg each sample was used • Sample buffer – Lysis buffer, 20 µL

Sample processing (Traditional 2 D/Di. GE) • Sample prep – labeling: 266#1 cy 3, 266#2 cy 5 and reciprocal • 1 st dimension: Protein IEF cell, Bio. Rad – Gel-strip: 3 -10 NL, 17 cm – Sample volume load: 300 µL – Running conditions: 250 V for 15 min. , ramp to 10000 V in 3 hrs. , reach 60000 V/hr, hold at 500 V • 2 nd dimension: Protein II xi cell, Bio. Rad – – Gel: Jule, 8 -16% Running buffer: TGS (Bio. Rad), 2 X on top chamber and 1 X on bottom chamber Running conditions: 16 m. A for 45 min. followed by 30 m. A for 5 hrs. # gels generated: 2

Gel processing • Gel fixing – – • Gel staining – • • None Gel imaging – – – • Buffer: 40% Methanol, 5% acidic acid Time: overnight Di. GE scanner: custom made with Prometrix® CCD camera Image generated per gel: 2 (total 4 images) Image label: • • Brickner. Gel_A_30 sec_Cy 3 -266#1 Brickner. Gel_A_30 sec_Cy 5 -266#2 Brickner. Gel_B_30 sec_Cy 5 -266#1 Brickner. Gel_B_30 sec_Cy 3 -266#2 Image Storage location: http: //www. genetics. pitt. edu/mygpcl/050208 Scanned gels and left over sample storage location – Gels: in Rm. 9035, BST 3 at 4°C, samples: -80°C

Previous analysis summary • None
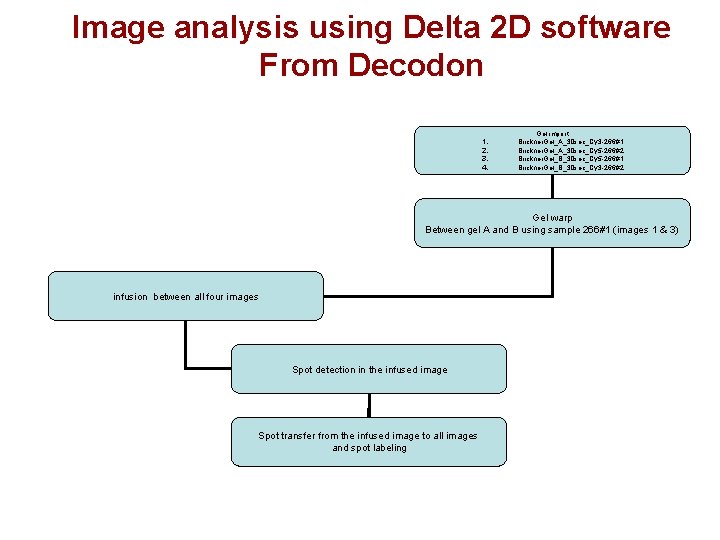
Image analysis using Delta 2 D software From Decodon 1. 2. 3. 4. Gel import Brickner. Gel_A_30 sec_Cy 3 -266#1 Brickner. Gel_A_30 sec_Cy 5 -266#2 Brickner. Gel_B_30 sec_Cy 5 -266#1 Brickner. Gel_B_30 sec_Cy 3 -266#2 Gel warp Between gel A and B using sample 266#1 (images 1 & 3) infusion between all four images Spot detection in the infused image Spot transfer from the infused image to all images and spot labeling

Dual-view image of 266#1 and 266#2 Possible knockdown Over expression in 266#2 labeled 266#2: blue spots 266#1: orange spots Overlap: black Under expression in 266#2 labeled
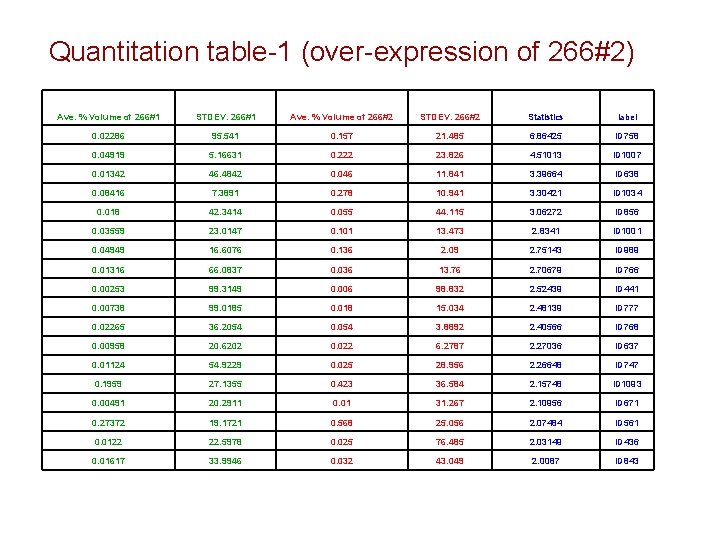
Quantitation table-1 (over-expression of 266#2) Ave. % Volume of 266#1 STDEV. 266#1 Ave. % Volume of 266#2 STDEV. 266#2 Statistics label 0. 02286 95. 541 0. 157 21. 485 6. 86425 ID 758 0. 04919 5. 16631 0. 222 23. 826 4. 51013 ID 1007 0. 01342 46. 4842 0. 046 11. 841 3. 39664 ID 638 0. 08416 7. 3891 0. 278 10. 941 3. 30421 ID 1034 0. 018 42. 3414 0. 055 44. 115 3. 06272 ID 856 0. 03559 23. 0147 0. 101 13. 473 2. 8341 ID 1001 0. 04949 16. 6076 0. 136 2. 09 2. 75143 ID 989 0. 01316 66. 0837 0. 036 13. 76 2. 70679 ID 766 0. 00253 99. 3149 0. 006 98. 832 2. 52439 ID 441 0. 00738 99. 0185 0. 018 15. 034 2. 48139 ID 777 0. 02265 36. 2054 0. 054 3. 8892 2. 40566 ID 768 0. 00958 20. 6202 0. 022 6. 2787 2. 27036 ID 637 0. 01124 54. 9229 0. 025 28. 956 2. 26648 ID 747 0. 1959 27. 1355 0. 423 36. 584 2. 15748 ID 1093 0. 00491 20. 2911 0. 01 31. 267 2. 10956 ID 671 0. 27372 19. 1721 0. 568 25. 056 2. 07484 ID 561 0. 0122 22. 5978 0. 025 76. 485 2. 03149 ID 436 0. 01617 33. 9946 0. 032 43. 049 2. 0087 ID 843
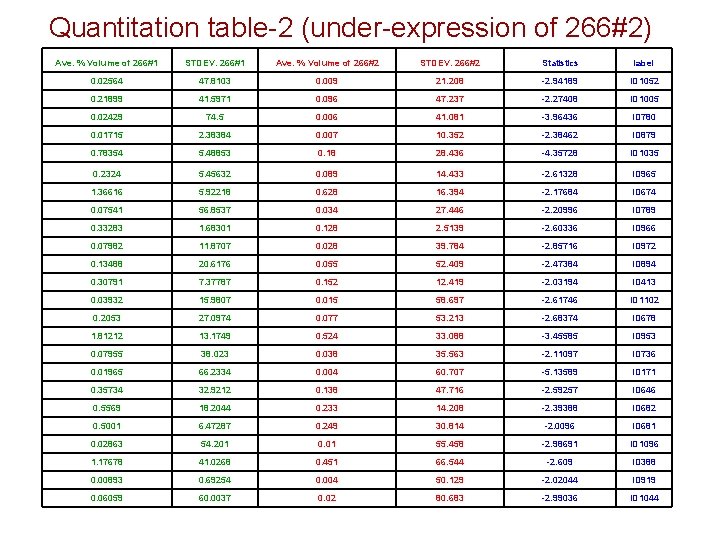
Quantitation table-2 (under-expression of 266#2) Ave. % Volume of 266#1 STDEV. 266#1 Ave. % Volume of 266#2 STDEV. 266#2 Statistics label 0. 02564 47. 8103 0. 009 21. 208 -2. 94189 ID 1052 0. 21899 41. 5971 0. 096 47. 237 -2. 27408 ID 1005 0. 02429 74. 5 0. 006 41. 081 -3. 96436 ID 780 0. 01715 2. 38384 0. 007 10. 352 -2. 38462 ID 879 0. 78354 5. 48853 0. 18 28. 436 -4. 35728 ID 1035 0. 2324 5. 45632 0. 089 14. 433 -2. 61328 ID 965 1. 36616 5. 92218 0. 628 16. 394 -2. 17684 ID 674 0. 07541 56. 8537 0. 034 27. 446 -2. 20996 ID 789 0. 33283 1. 68301 0. 128 2. 5139 -2. 60336 ID 966 0. 07982 11. 8707 0. 028 39. 784 -2. 85716 ID 972 0. 13488 20. 6176 0. 055 52. 409 -2. 47384 ID 894 0. 30791 7. 37787 0. 152 12. 419 -2. 03194 ID 413 0. 03932 15. 9807 0. 015 58. 697 -2. 61746 ID 1102 0. 2053 27. 0974 0. 077 53. 213 -2. 68374 ID 678 1. 81212 13. 1749 0. 524 33. 088 -3. 45585 ID 953 0. 07955 38. 023 0. 038 35. 563 -2. 11097 ID 736 0. 01965 66. 2334 0. 004 60. 707 -5. 13589 ID 171 0. 35734 32. 9212 0. 138 47. 716 -2. 59257 ID 646 0. 5569 18. 2044 0. 233 14. 208 -2. 39388 ID 682 0. 5001 6. 47287 0. 249 30. 814 -2. 0096 ID 681 0. 02863 54. 201 0. 01 55. 458 -2. 98691 ID 1096 1. 17678 41. 0268 0. 451 66. 544 -2. 609 ID 388 0. 00893 0. 69254 0. 004 50. 129 -2. 02044 ID 919 0. 06059 60. 0037 0. 02 80. 683 -2. 99036 ID 1044
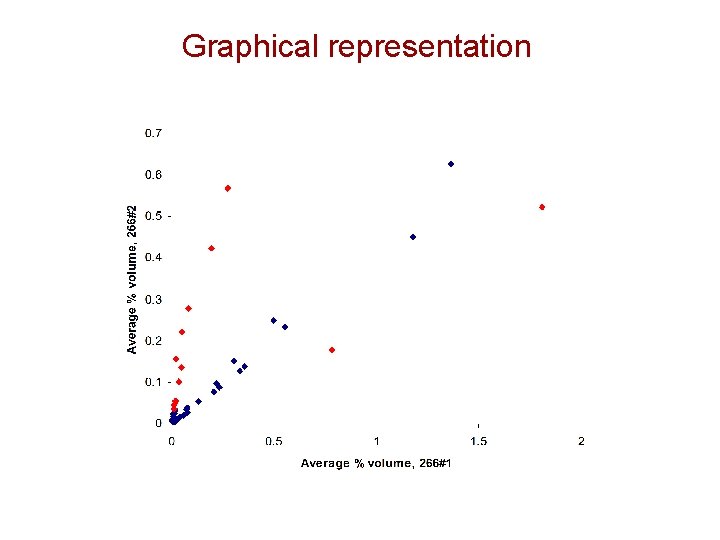
Graphical representation

Preliminary Conclusions • 266#1 versus 266#2 – Number of over-expressed spots: 18 – Number of under-expression: 24 • Report storage location – Folder: 052708_Delta 2 D_image analysis – Contents: Power point presentation, pick list, snapshot images with labeling and Delta 2 D report

Mass spectrometry (MS) based peptide identification • Gel bands of interests are excised for in-gel digestion • Alternatively, a protein sample of interest can be digested in solution • The digest is then subjected to MS or MS/MS for peptide identification – You can chose to run LC before MS • Some instruments like FT-MS allows MS and MS/MS of undigested protein samples

List of useful proteomics websites • • • http: //www. fixingproteomics. org/ http: //www. ionsource. com/ www. ebi. ac. uk www. proteomecommons. org www. peptideatlas. org http: //www. biochem. mpg. de/mann http: //ncrr. pnl. gov/software/ http: //tools. proteomecenter. org http: //www. pil. sdu. dk/ www. proteomesoftware. com www. hprd. org www. expasy. org Click on each link to get familiar

Protein database search engines (Algorithms) • Commercial – – – Mascot (Matrix sciences) Sequest (Thermo Scientific) Phenyx (Gene. Bio) Spectrum Mills (Agilent Technologies) Paragon etc. (Applied Biosystems) • Open source – Mascot (for small dataset) • http: //www. matrixscience. com/search_form_select. html – X!Tandem • http: //www. thegpm. org/tandem/ • Requires command-line use – OMSSA etc. • http: //pubchem. ncbi. nlm. nih. gov/omssa/

Source of ambiguity in proteomics data analysis • • Quality of the MS/MS spectra Redundant databases Search strategy used Nature of the model used by database search engines (scoring algorithms) – The most common source of ambiguity and incorrect assignment
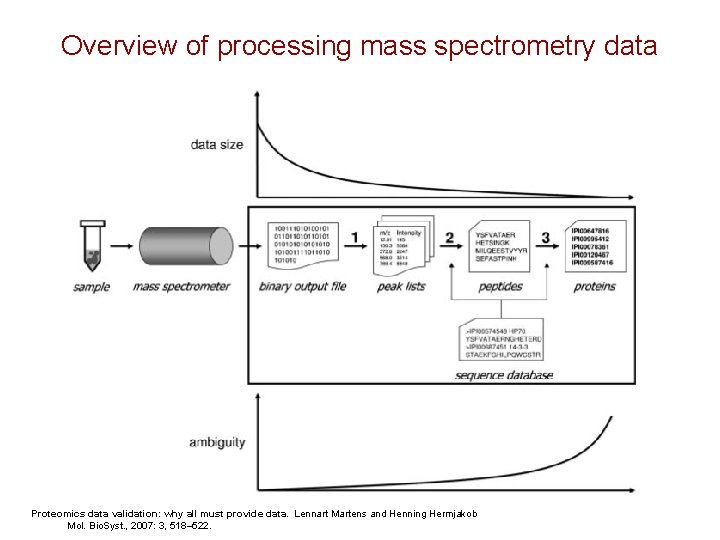
Overview of processing mass spectrometry data Proteomics data validation: why all must provide data. Lennart Martens and Henning Hermjakob Mol. Bio. Syst. , 2007: 3, 518– 522.

Why different search engines generate different peptide lists from the same dataset? ? • Mascot – Probability base MOWSE scoring • Sequest – Cross-correlation (Xcorr) among experimental and theoretical spectra is used – Reports delta. Cn • X-Tandem – Considers only B/Y-type ions – Creates a database of proteins identified and performs an extensive search on only identified proteins

Protein databases • NCBI – NCBI is Entrez Protein database from National Center for Biotechnology information and contains redundant protein sequences with poor annotation. • • Ref. Seq is NCBI’s Reference Sequence database with a comprehensive, integrated, non-redundant, wellannotated set of sequences. Uniprot/Swiss-Port – The Uni. Prot Knowledgebase (Uni. Prot. KB) consists of two sections: • • manually annotated and reviewed Uni. Prot. KB/Swiss-Prot and automatically annotated Uni. Prot. KB/Tr. EMBL. – Uni. Prot. KB/Swiss-Prot is well-curated, well annotated, non redundant and considerably smaller than NCBI, therefore widely used. • • IPI – IPI, International Protein Index databases, is used for species specific searches and is maintained by European Bioinformatics Institute (EBI). The decision as which databases to use solely depends on aim of the project and type of the experiment in concern – If the goal is to receive highest sensitivity, NCBI is more desirable as a first step. • However, it is time consuming to search against a large database and it requires manual validation as a second step and/or further distillation of the protein list based on other specific databases, but for identifying sequence variant, NCBI is a better starting point. – Uni. Prot. KB/Swiss-Prot, on the other hand, is a better option for investigators seeking faster and reliable search results. – If species information is known, IPI database is a good candidate containing protein sequences with cross-references to all its source data e. g. Ensembl, Uni. Prot, Ref. Seq. .

#1 problem Proliferation of new search algorithms, with a variety of settings; which one(s)?

Importance of database search algorithms in peptide identification Each search engine identifies about the same number of spectra, SEQUEST But the overlap is surprisingly small. 9% 22% 4% Different search engines match different spectra. 34% X!Tandem 19% 7% 5% Courtesy: Proteome Software Inc. Mascot

Sequest, mascot and X-tandem scores – SEQUEST: XCorr>2. 5, Delta. Cn>0. 1 – Mascot: Ion Score-Identity Score>0 – X! Tandem: E-Value<0. 01 How do we compare these? ? That’s when Scaffold comes in Courtesy: Proteome Software Inc.
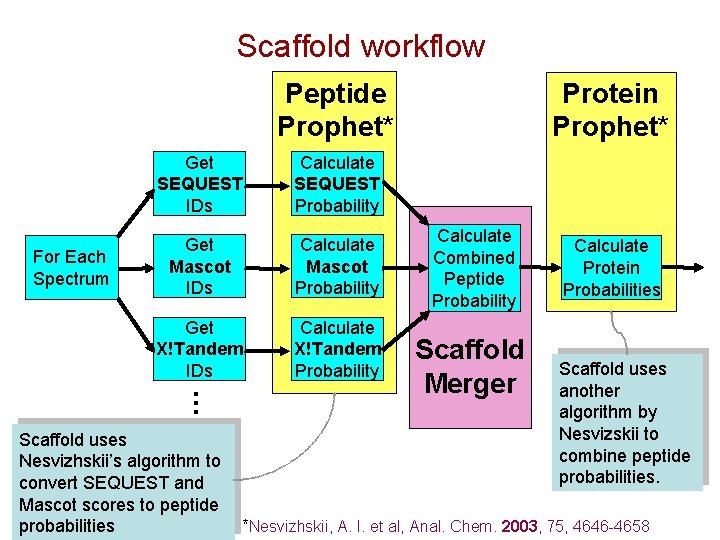
Scaffold workflow Peptide Prophet* Calculate SEQUEST Probability Get Mascot IDs Calculate Mascot Probability Get X!Tandem IDs Calculate X!Tandem Probability … For Each Spectrum Get SEQUEST IDs Protein Prophet* Calculate Combined Peptide Probability Scaffold Merger Calculate Protein Probabilities Scaffold uses another algorithm by Nesvizskii to combine peptide probabilities. Scaffold uses Nesvizhskii’s algorithm to convert SEQUEST and Mascot scores to peptide probabilities *Nesvizhskii, A. I. et al, Anal. Chem. 2003, 75, 4646 -4658
Scaffold View Click on Scaffold and import file MSX 50. sfd We will be able to import this file into protein center after exporting it as Prot. XML file format ( MSX 50. xml)

Consensus Study Conducted by BAC • • • Mascot only (MO) Sequest only (SO) S X!Tandem only (XO) ? ? ? Union of M & S (MSU) M ? MXU ? ? SXU ? X MSXU Intersection of M & S (MSI) Evaluate the performance of each methods using a SXI standard protein dataset MXI Performance measures are sensitivity and specificity MSXI of each methods Sultana T, Jordan R, Lyons-Weiler J. 2009. Optimization of the use of consensus methods for the detection and putative identification of peptides via mass spectrometry using protein standard mixtures. J Proteomics Bioinform 2: 263 -273.

Scaffold confidence filter settings • Minimum Protein – defines the probability that a protein’s identification is correct (20, 50, 80, 95, 99. 9) • Minimum # Peptides – filters results by the number of unique peptides on which the identification is based (1, 2, 3, 4, 5) • Minimum Peptide – requires a minimum probability from at least one spectrum (0, 20, 50, 80, 95)

GPCL-BAC recommendations (Sultana et al. , 2009 publication) • ‘Most Accurate’ peptide identification – Union of Sequest, Mascot & X!Tandem (MSXU) – Scaffold filter: 95% protein probability, 2 minimum unique peptides & 50% peptide probability. • ‘Most Sensitive’ peptide identification – Union of Sequest & Mascot (MSU) or union of Sequest & X!Tandem (MSXU) or Sequest only (SO) – Scaffold filter: 80% protein probability, 1 minimum unique peptides & 50% peptide probability. • ‘Most Specific’ peptide identification – Union of Mascot & X!Tandem (MXU) or Mascot only (MO) – Scaffold filter: 99% protein probability, 3 minimum unique peptides & 50% peptide probability. Sultana T, Jordan R, Lyons-Weiler J. 2009. Optimization of the use of consensus methods for the detection and putative identification of peptides via mass spectrometry using protein standard mixtures. J Proteomics Bioinform 2: 263 -273. Sensitivity and specificity trade off

Biology of MICA Protein in Human Sarcoma Cells Final Research Report March 5, 2010 PI: _____ GPCL-Bioinformatics Analysis Core Analysts: Tamanna Sultana and James Lyons Weiler Project Location: http: //mygpcl/ Example of consensus peptide ID data analysis and reporting

Specific Aims (obtained from PI) • ID proteins in sample – Identification of other closely interacting proteins with MICA in human sarcoma cells

Study Details • # Samples: protein extract of osteo-sarcoma cell lines – SCH 2473 A 8 MA 3_sample 1 – SCH 2473 A 9 MA 3_sample 2 • Sample preparation: – Stable over expressed MICA: immuno-precipitated with MICA antibody and then the complex were pull down – ~5 ug/10 u. L protein was reduced with TCEP, alkylated with iodoacetamide and trypsin digested. – LCQ-Deca-XL (LC-ESI-MS) was used for MS and MS/MS data generation • # Data sets generated – SCH 2473 A 8 MA 3 -Spectra. File. RAW – SCH 2473 A 9 MA 3 -Spectra. File. RAW

Previous Analysis Summary • Sequest search results provided by Manny Schreiber from Proteomics Lab

GPCL-BAC Profile Data Analysis • Start with the LC-MS Raw data file – Convert to peak list (MGF file format) – Run Mascot and X!Tandem search • Using Scaffold, combine the search results of Sequest, Mascot and X!Tandem • Provide protein lists – (Most) Accurate (list 1) – (Most) Sensitive (list 2) – (Most) Specific (list 3)
SCH 2473 A 8 MA 3. RAW SCH 2473 A 9 MA 3. RAW

Database Search Parameters • Database – IPI human v. 3. 57 • Search algorithms – Mascot, Sequest & X!Tandem • Modifications (variable) – Carbamidomethyl (+57 @C) and oxidation (+16 @M) • Missed cleavages: 2 maximum • Error tolerance: 2 Da on both parent and fragment ions • Peak list conversion – Raw file were converted into Mascot generic format (MGF) peak list using extract_msn provided by Xcalibur software of LCQ instrument
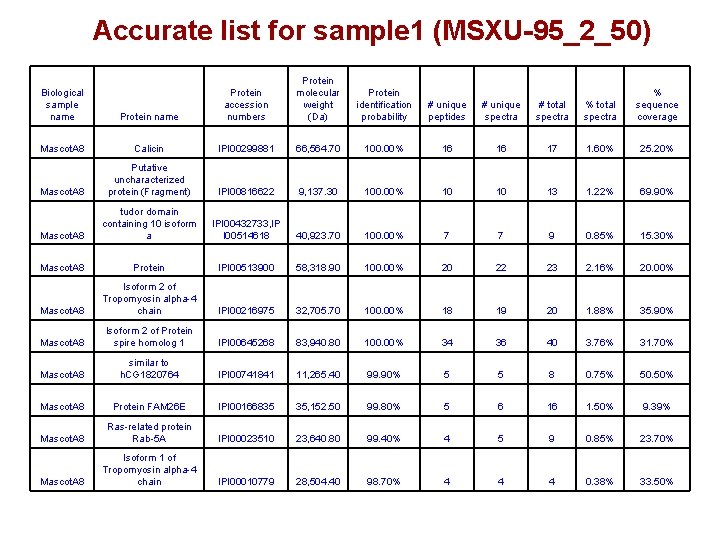
Accurate list for sample 1 (MSXU-95_2_50) Protein name Protein accession numbers Protein molecular weight (Da) Protein identification probability # unique peptides # unique spectra # total spectra % sequence coverage Mascot. A 8 Calicin IPI 00299881 66, 564. 70 100. 00% 16 16 17 1. 60% 25. 20% Mascot. A 8 Putative uncharacterized protein (Fragment) IPI 00816622 9, 137. 30 100. 00% 10 10 13 1. 22% 69. 90% Mascot. A 8 tudor domain containing 10 isoform a IPI 00432733, IP I 00514618 40, 923. 70 100. 00% 7 7 9 0. 85% 15. 30% Mascot. A 8 Protein IPI 00513900 58, 318. 90 100. 00% 20 22 23 2. 16% 20. 00% Mascot. A 8 Isoform 2 of Tropomyosin alpha-4 chain IPI 00216975 32, 705. 70 100. 00% 18 19 20 1. 88% 35. 90% Mascot. A 8 Isoform 2 of Protein spire homolog 1 IPI 00645268 83, 940. 80 100. 00% 34 36 40 3. 76% 31. 70% Mascot. A 8 similar to h. CG 1820764 IPI 00741841 11, 265. 40 99. 90% 5 5 8 0. 75% 50. 50% Mascot. A 8 Protein FAM 26 E IPI 00166835 35, 152. 50 99. 80% 5 6 16 1. 50% 9. 39% Mascot. A 8 Ras-related protein Rab-5 A IPI 00023510 23, 640. 80 99. 40% 4 5 9 0. 85% 23. 70% Mascot. A 8 Isoform 1 of Tropomyosin alpha-4 chain IPI 00010779 28, 504. 40 98. 70% 4 4 4 0. 38% 33. 50% Biological sample name
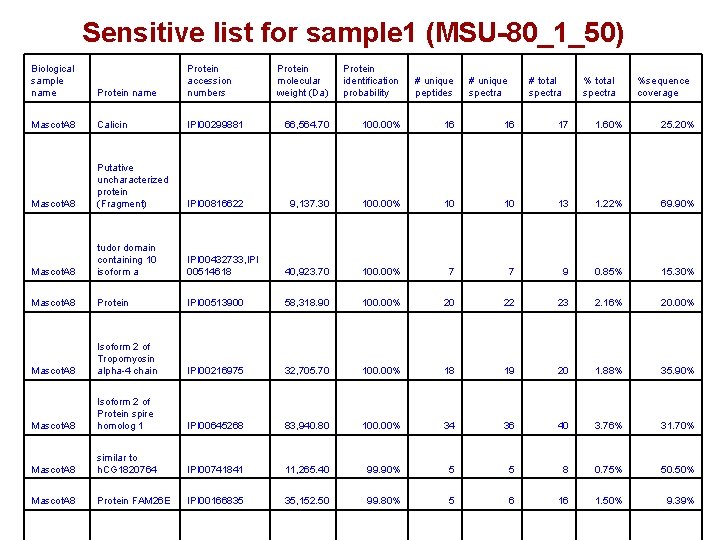
Sensitive list for sample 1 (MSU-80_1_50) Biological sample name Protein accession numbers Protein molecular weight (Da) Protein identification probability # unique peptides Mascot. A 8 Calicin IPI 00299881 66, 564. 70 100. 00% 16 16 17 1. 60% 25. 20% Mascot. A 8 Putative uncharacterized protein (Fragment) IPI 00816622 9, 137. 30 100. 00% 10 10 13 1. 22% 69. 90% Mascot. A 8 tudor domain containing 10 isoform a IPI 00432733, IPI 00514618 40, 923. 70 100. 00% 7 7 9 0. 85% 15. 30% Mascot. A 8 Protein IPI 00513900 58, 318. 90 100. 00% 20 22 23 2. 16% 20. 00% Mascot. A 8 Isoform 2 of Tropomyosin alpha-4 chain IPI 00216975 32, 705. 70 100. 00% 18 19 20 1. 88% 35. 90% Mascot. A 8 Isoform 2 of Protein spire homolog 1 IPI 00645268 83, 940. 80 100. 00% 34 36 40 3. 76% 31. 70% Mascot. A 8 similar to h. CG 1820764 IPI 00741841 11, 265. 40 99. 90% 5 5 8 0. 75% 50. 50% Mascot. A 8 Protein FAM 26 E IPI 00166835 35, 152. 50 99. 80% 5 6 16 1. 50% 9. 39% # unique spectra # total spectra %sequence coverage
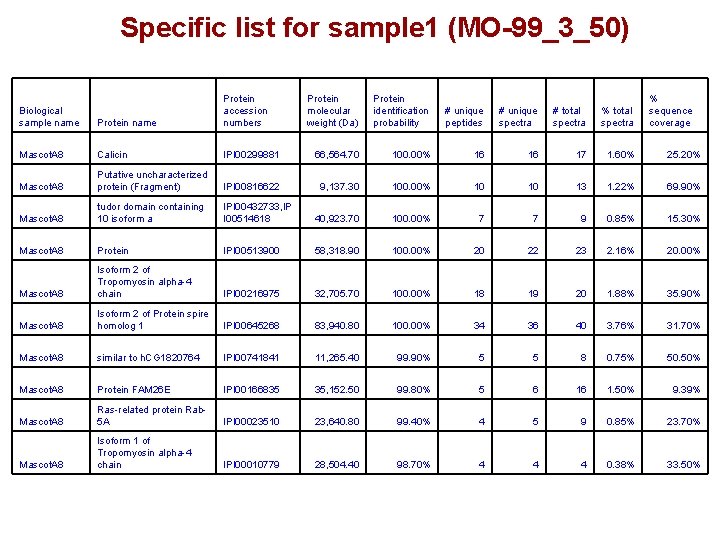
Specific list for sample 1 (MO-99_3_50) Biological sample name Protein accession numbers Protein molecular weight (Da) Protein identification probability # unique peptides # unique spectra # total spectra % total spectra Mascot. A 8 Calicin Mascot. A 8 % sequence coverage IPI 00299881 66, 564. 70 100. 00% 16 16 17 1. 60% 25. 20% Putative uncharacterized protein (Fragment) IPI 00816622 9, 137. 30 100. 00% 10 10 13 1. 22% 69. 90% Mascot. A 8 tudor domain containing 10 isoform a IPI 00432733, IP I 00514618 40, 923. 70 100. 00% 7 7 9 0. 85% 15. 30% Mascot. A 8 Protein IPI 00513900 58, 318. 90 100. 00% 20 22 23 2. 16% 20. 00% Mascot. A 8 Isoform 2 of Tropomyosin alpha-4 chain IPI 00216975 32, 705. 70 100. 00% 18 19 20 1. 88% 35. 90% Mascot. A 8 Isoform 2 of Protein spire homolog 1 IPI 00645268 83, 940. 80 100. 00% 34 36 40 3. 76% 31. 70% Mascot. A 8 similar to h. CG 1820764 IPI 00741841 11, 265. 40 99. 90% 5 5 8 0. 75% 50. 50% Mascot. A 8 Protein FAM 26 E IPI 00166835 35, 152. 50 99. 80% 5 6 16 1. 50% 9. 39% Mascot. A 8 Ras-related protein Rab 5 A IPI 00023510 23, 640. 80 99. 40% 4 5 9 0. 85% 23. 70% Mascot. A 8 Isoform 1 of Tropomyosin alpha-4 chain IPI 00010779 28, 504. 40 98. 70% 4 4 4 0. 38% 33. 50%

Venn diagram for sample 1 using sensitive list SEQUEST 29% 0% 3% 0% X!Tandem 19% 42% 7% Mascot
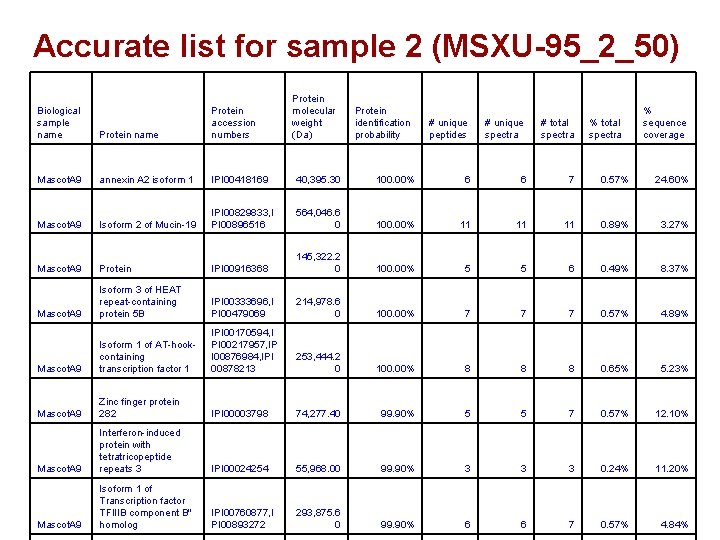
Accurate list for sample 2 (MSXU-95_2_50) Protein molecular weight (Da) Protein identification probability # unique peptides # unique spectra # total spectra IPI 00418169 40, 395. 30 100. 00% 6 6 7 0. 57% 24. 60% Isoform 2 of Mucin-19 IPI 00829833, I PI 00896516 564, 046. 6 0 100. 00% 11 11 11 0. 89% 3. 27% Mascot. A 9 Protein IPI 00916368 145, 322. 2 0 100. 00% 5 5 6 0. 49% 8. 37% Mascot. A 9 Isoform 3 of HEAT repeat-containing protein 5 B IPI 00333696, I PI 00479069 214, 978. 6 0 100. 00% 7 7 7 0. 57% 4. 89% Mascot. A 9 Isoform 1 of AT-hookcontaining transcription factor 1 IPI 00170594, I PI 00217957, IP I 00876984, IPI 00878213 253, 444. 2 0 100. 00% 8 8 8 0. 65% 5. 23% Mascot. A 9 Zinc finger protein 282 IPI 00003798 74, 277. 40 99. 90% 5 5 7 0. 57% 12. 10% Mascot. A 9 Interferon-induced protein with tetratricopeptide repeats 3 IPI 00024254 55, 968. 00 99. 90% 3 3 3 0. 24% 11. 20% Mascot. A 9 Isoform 1 of Transcription factor TFIIIB component B'' homolog IPI 00760877, I PI 00893272 293, 875. 6 0 99. 90% 6 6 7 0. 57% 4. 84% Biological sample name Protein accession numbers Mascot. A 9 annexin A 2 isoform 1 Mascot. A 9 % total spectra % sequence coverage
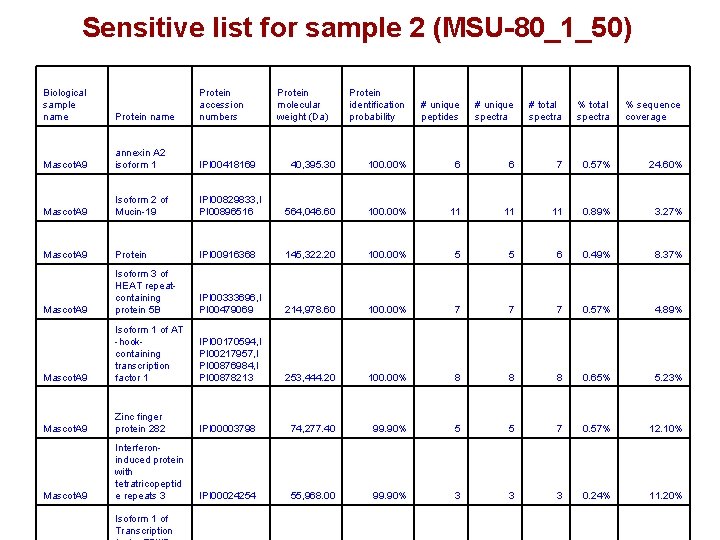
Sensitive list for sample 2 (MSU-80_1_50) Biological sample name Protein accession numbers Mascot. A 9 annexin A 2 isoform 1 IPI 00418169 Mascot. A 9 Isoform 2 of Mucin-19 Mascot. A 9 Protein identification probability # unique peptides # unique spectra # total spectra % sequence coverage 40, 395. 30 100. 00% 6 6 7 0. 57% 24. 60% IPI 00829833, I PI 00896516 564, 046. 60 100. 00% 11 11 11 0. 89% 3. 27% Protein IPI 00916368 145, 322. 20 100. 00% 5 5 6 0. 49% 8. 37% Mascot. A 9 Isoform 3 of HEAT repeatcontaining protein 5 B IPI 00333696, I PI 00479069 214, 978. 60 100. 00% 7 7 7 0. 57% 4. 89% Mascot. A 9 Isoform 1 of AT -hookcontaining transcription factor 1 IPI 00170594, I PI 00217957, I PI 00876984, I PI 00878213 253, 444. 20 100. 00% 8 8 8 0. 65% 5. 23% Mascot. A 9 Zinc finger protein 282 IPI 00003798 74, 277. 40 99. 90% 5 5 7 0. 57% 12. 10% Mascot. A 9 Interferoninduced protein with tetratricopeptid e repeats 3 IPI 00024254 55, 968. 00 99. 90% 3 3 3 0. 24% 11. 20% Isoform 1 of Transcription Protein molecular weight (Da)
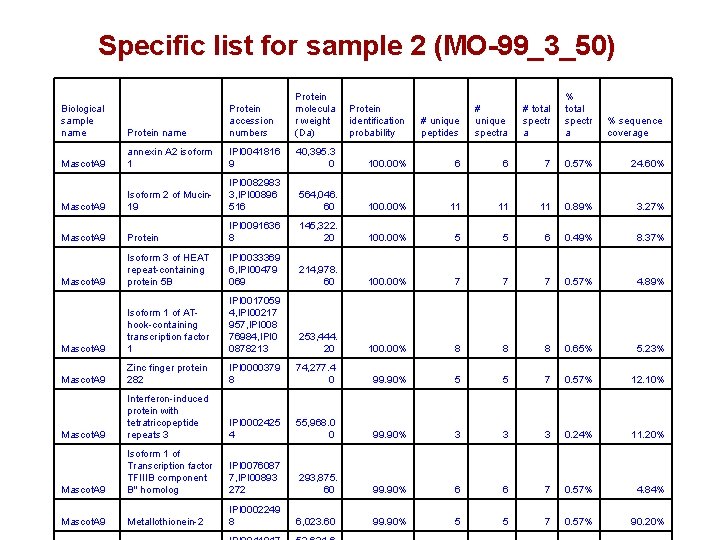
Specific list for sample 2 (MO-99_3_50) Protein name Protein accession numbers Protein molecula r weight (Da) Protein identification probability # unique peptides # unique spectra # total spectr a % sequence coverage Mascot. A 9 annexin A 2 isoform 1 IPI 0041816 9 40, 395. 3 0 100. 00% 6 6 7 0. 57% 24. 60% Mascot. A 9 Isoform 2 of Mucin 19 IPI 0082983 3, IPI 00896 516 564, 046. 60 100. 00% 11 11 11 0. 89% 3. 27% Mascot. A 9 Protein IPI 0091636 8 145, 322. 20 100. 00% 5 5 6 0. 49% 8. 37% Mascot. A 9 Isoform 3 of HEAT repeat-containing protein 5 B IPI 0033369 6, IPI 00479 069 214, 978. 60 100. 00% 7 7 7 0. 57% 4. 89% Mascot. A 9 Isoform 1 of AThook-containing transcription factor 1 IPI 0017059 4, IPI 00217 957, IPI 008 76984, IPI 0 0878213 253, 444. 20 100. 00% 8 8 8 0. 65% 5. 23% Mascot. A 9 Zinc finger protein 282 IPI 0000379 8 74, 277. 4 0 99. 90% 5 5 7 0. 57% 12. 10% Mascot. A 9 Interferon-induced protein with tetratricopeptide repeats 3 IPI 0002425 4 55, 968. 0 0 99. 90% 3 3 3 0. 24% 11. 20% Mascot. A 9 Isoform 1 of Transcription factor TFIIIB component B'' homolog IPI 0076087 7, IPI 00893 272 293, 875. 60 99. 90% 6 6 7 0. 57% 4. 84% Mascot. A 9 Metallothionein-2 IPI 0002249 8 6, 023. 60 99. 90% 5 5 7 0. 57% 90. 20% Biological sample name

Venn diagram for sample 2 using sensitive list SEQUEST 35% 0% 13% 3% X!Tandem 5% 2% 42% Mascot
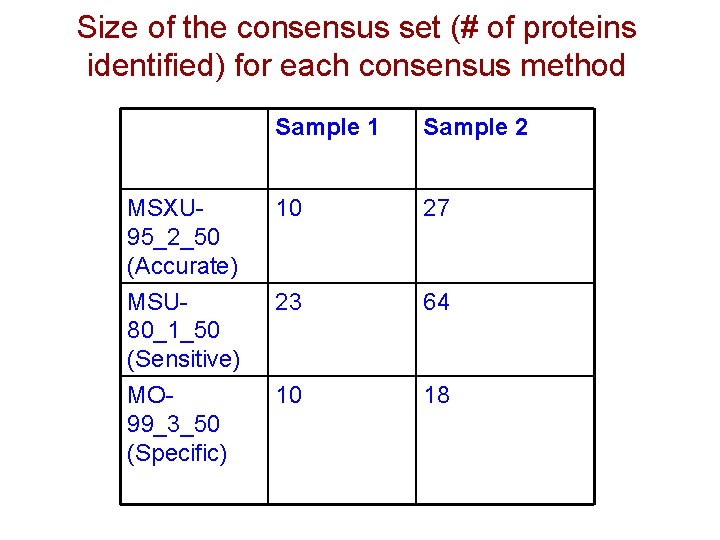
Size of the consensus set (# of proteins identified) for each consensus method Sample 1 Sample 2 MSXU 95_2_50 (Accurate) MSU 80_1_50 (Sensitive) 10 27 23 64 MO 99_3_50 (Specific) 10 18

Sample 1 consensus list for each consensus method Acurate protein list Specific protein list Sensitive protein list Calicin Putative uncharacterized protein (Fragment) tudor domain containing 10 isoform a Protein Isoform 2 of Tropomyosin alpha-4 chain Isoform 2 of Tropomyosin alpha-4 chain Isoform 2 of Protein spire homolog 1 similar to h. CG 1820764 Protein FAM 26 E Ras-related protein Rab-5 A Isoform 1 of Tropomyosin alpha-4 chain Isoform 1 of Tropomyosin alpha-4 chain Isoform 2 of Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 Corrupt Accession: PI: IPI 00890727. 1|SWISS-PROT: Q 8 NB 90 -2 Isoform 1 of Uncharacterized protein KIAA 1107 Isoform 2 of Protein FAM 184 A Isoform 2 of Metallothionein-1 G GTPase IMAP family member 8 Isoform 1 of WSC domain-containing protein 2 similar to GOLGA 8 A protein

Sample 2 consensus list for each consensus method Accurate protein list Specific protein list Sensitive protein list annexin A 2 isoform 1 Isoform 2 of Mucin-19 Protein IPI 00916368 Isoform 3 of HEAT repeatcontaining protein 5 B Isoform 3 of HEAT repeat-containing protein 5 B Isoform 1 of AT-hook-containing transcription factor 1 Zinc finger protein 282 Interferon-induced protein with tetratricopeptide repeats 3 Isoform 1 of Transcription factor TFIIIB component B'' homolog Metallothionein-2 Vimentin Isoform 1 of Disabled homolog 2 interacting protein Isoform 1 of Disabled homolog 2 -interacting protein Isoform 1 of Disabled homolog 2 -interacting protein Similar to Signal peptidase complex subunit 2 Vimentin Isoform 2 of Protein Dok-7 Vimentin Annexin A 5 Isoform GTBP-N of DNA mismatch repair protein Msh 6 Annexin A 5 Isoform 2 of Protein Dok-7 Isoform 1 of HEAT repeatcontaining protein 5 A Isoform 2 of Protein Dok-7 Isoform GTBP-N of DNA mismatch repair protein Msh 6 Isoform 1 of Cullin-4 A Isoform GTBP-N of DNA mismatch repair protein Msh 6

Methods Text • • Sample Preparation – Protein extracts of osteo-sarcoma cell lines, SCH 2473 A 8 MA 3_sample 1 and SCH 2473 A 9 MA 3_sample 2 were processed in the Genomics and Proteomics Core Laboratories (GPCL) at the University of Pittsburgh prior to performing tandem mass spectrometry. In brief, immuno precipitated samples provided by PI’s lab were reduced with tris-2 -carboxyethyl-phosphine (TCEP), alkylated with iodoacetamide (IAC), and digested with trypsin (Promega). The ESI-MS and information dependent (IDA) MS/MS spectra were acquired at GPCL with an LCQ-Deca-XL coupled with a nano-LC system (Thermo Scientific, Waltham, MA). The IDA was set so that MS/MS was done on the top three intense peaks per cycle. Database Search – Experimental Raw spectra files were searched against the human database, IPI Human v 3. 57, for identifying peptides using following three search algorithms: Sequest, Mascot, and X!Tandem. The search parameters for searching candidate peptides were: precursor ion tolerance: 2 Da; fragment ion tolerance: 2 Da; variable modifications: Carbamidomethyl on cysteine, and oxidation on methionine; maximum missed cleavages: 2. For Mascot and X!Tandem search, the raw files were converted to MGF peak list. Merging the Data – Files containing database search results derived from Mascot, Sequest, and X!Tandem were imported into Scaffold. The software then merged the peptide lists identified by all the three search algorithms, re-scored, and re-ranked. Scaffold uses Peptide. Prophet and Protein. Prophet, that employ Bayesian statistics to combine the probability of identifying spectra with the probability that all search methods agree with each other. Protein List Generation – Accurate list: unions of Mascot, Sequest and X!Tandem (MSXU) at 95% minimum protein probability, 2 minimum unique peptides and 50% minimum peptide probability – Sensitive list: unions of Mascot, and Sequest (MSU) at 80% minimum protein probability, 1 minimum unique peptides and 50% minimum peptide probability – Specific list: mascot only at 99% minimum protein probability, 3 minimum unique peptides and 50% minimum peptide probability

Pathway express • Pathway express analysis* – Convert the ‘Protein accession numbers’ to ‘Genbank accession’ IDs using DAVID • That is your input file for pathway express *
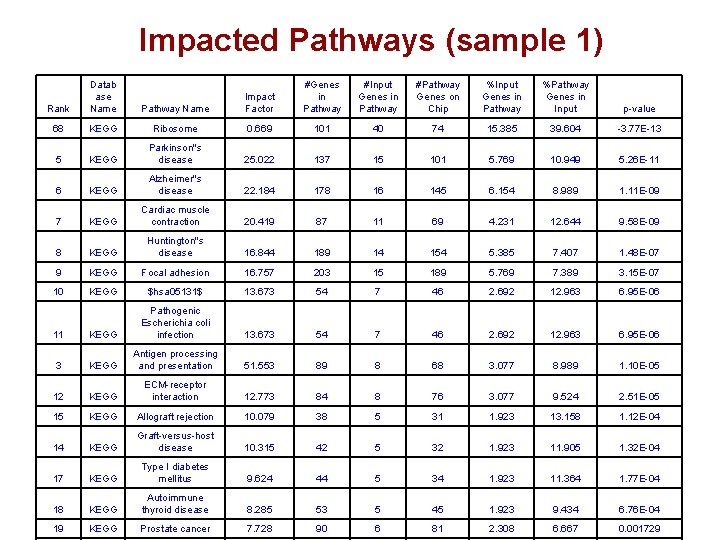
Impacted Pathways (sample 1) Rank Datab ase Name Pathway Name Impact Factor #Genes in Pathway #Input Genes in Pathway #Pathway Genes on Chip %Input Genes in Pathway %Pathway Genes in Input p-value 68 KEGG Ribosome 0. 669 101 40 74 15. 385 39. 604 -3. 77 E-13 5 KEGG Parkinson''s disease 25. 022 137 15 101 5. 769 10. 949 5. 26 E-11 6 KEGG Alzheimer''s disease 22. 184 178 16 145 6. 154 8. 989 1. 11 E-09 7 KEGG Cardiac muscle contraction 20. 419 87 11 69 4. 231 12. 644 9. 58 E-09 8 KEGG Huntington''s disease 16. 844 189 14 154 5. 385 7. 407 1. 48 E-07 9 KEGG Focal adhesion 16. 757 203 15 189 5. 769 7. 389 3. 15 E-07 10 KEGG $hsa 05131$ 13. 673 54 7 46 2. 692 12. 963 6. 95 E-06 11 KEGG Pathogenic Escherichia coli infection 13. 673 54 7 46 2. 692 12. 963 6. 95 E-06 3 KEGG Antigen processing and presentation 51. 553 89 8 68 3. 077 8. 989 1. 10 E-05 12 KEGG ECM-receptor interaction 12. 773 84 8 76 3. 077 9. 524 2. 51 E-05 15 KEGG Allograft rejection 10. 079 38 5 31 1. 923 13. 158 1. 12 E-04 14 KEGG Graft-versus-host disease 10. 315 42 5 32 1. 923 11. 905 1. 32 E-04 17 KEGG Type I diabetes mellitus 9. 624 44 5 34 1. 923 11. 364 1. 77 E-04 18 KEGG Autoimmune thyroid disease 8. 285 53 5 45 1. 923 9. 434 6. 76 E-04 19 KEGG Prostate cancer 7. 728 90 6 81 2. 308 6. 667 0. 001729
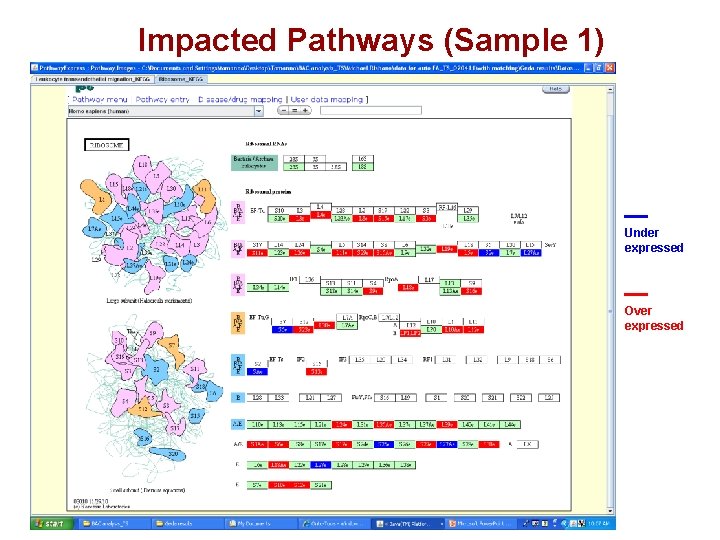
Impacted Pathways (Sample 1) Under expressed Over expressed

Protein Center v 3. 0. 4 A Proteomics bioinformatics tool available through HSLS

Import and view a protein dataset, export desired info • Go to http: //hsls. proteincenter. proxeon. com/Pro. Xweb/ • Log in • Click on the file MSX 50. xml • Play with peptide, protein and cluster view • Learn how to select information of interest, export files or create a report • Learn how to compare two datasets (A 8. xml and A 9. xml)

Bioinformatics and statistical analysis • Single protein – Look up using Accession Key 6807647 • Dataset: Human Red blood cell (h. RBC) dataset – Datasets • Proxeon – Tutorials » Data set comparison and click on h. RBC_proteome For further help email Protein Center support at proteincenter-support@proxeon. com

Acknowledgement • Genomics and Proteomics Core Laboratories • James Lyons-Weiler, Director of BAC • Rick Jordan, programmer of BAC
- Slides: 73