Quantitative Proteomics II Targeted Quantitation Traditional Affinitybased proteomics

Quantitative Proteomics II: Targeted Quantitation
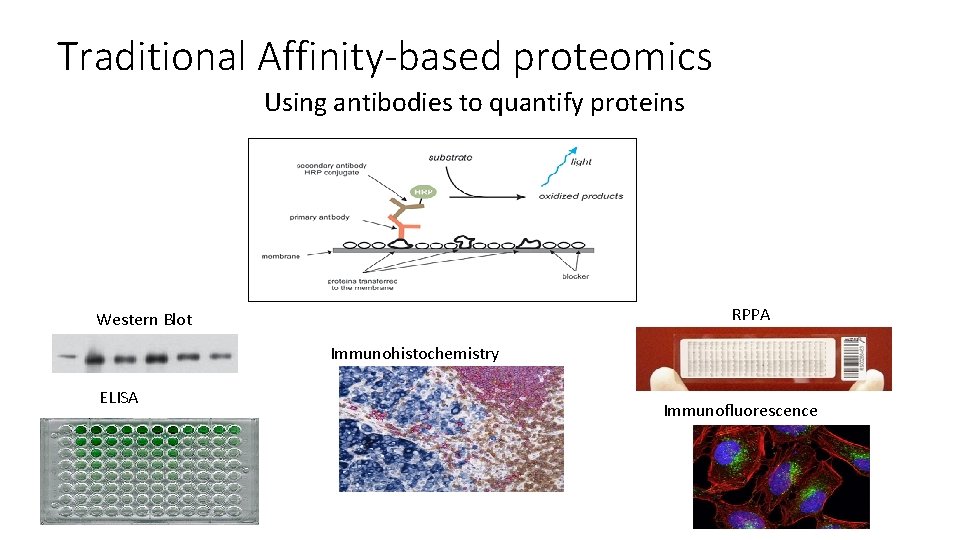
Traditional Affinity-based proteomics Using antibodies to quantify proteins RPPA Western Blot Immunohistochemistry ELISA Immunofluorescence

Using MS/MS for quantitative protein assays • Most clinical and biology studies in the literature rely on measurement of proteins which already have antibodies available • Goal of targeted proteomics is to create reliable, high quality assays to measure proteins that do not require antibodies and instead rely on MS/MS

Targeted Proteomics • Focuses on a subset of proteins of interest • • • Disease related changes in proteins Signaling processes Highly multiplexed alternative method to western blots/antibodies Can focus on unique and informative peptides for protein of interest Hypothesis driven questions! Nature Method of the Year 2012

Targeted Proteomics Overview
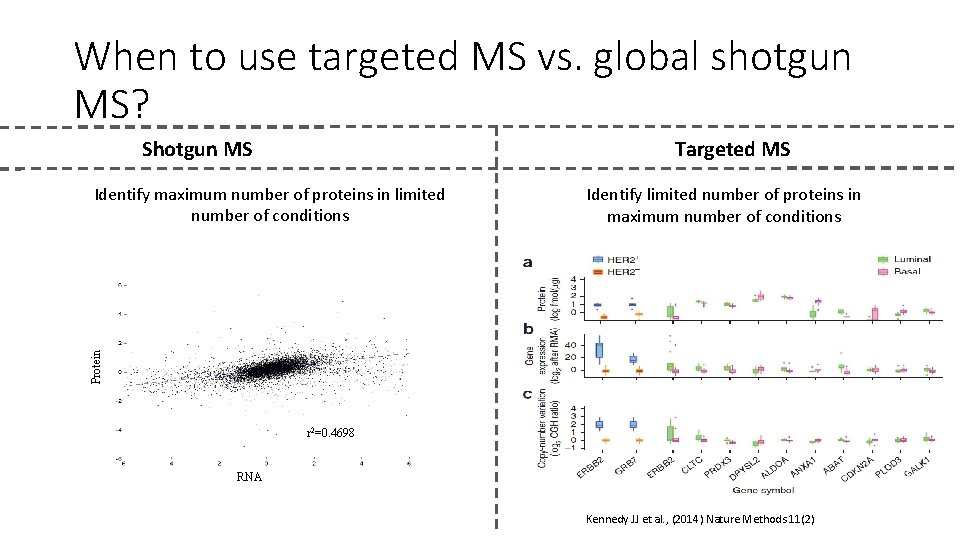
When to use targeted MS vs. global shotgun MS? Shotgun MS Targeted MS Identify limited number of proteins in maximum number of conditions Protein Identify maximum number of proteins in limited number of conditions r 2=0. 4698 RNA Kennedy JJ et al. , (2014) Nature Methods 11(2)
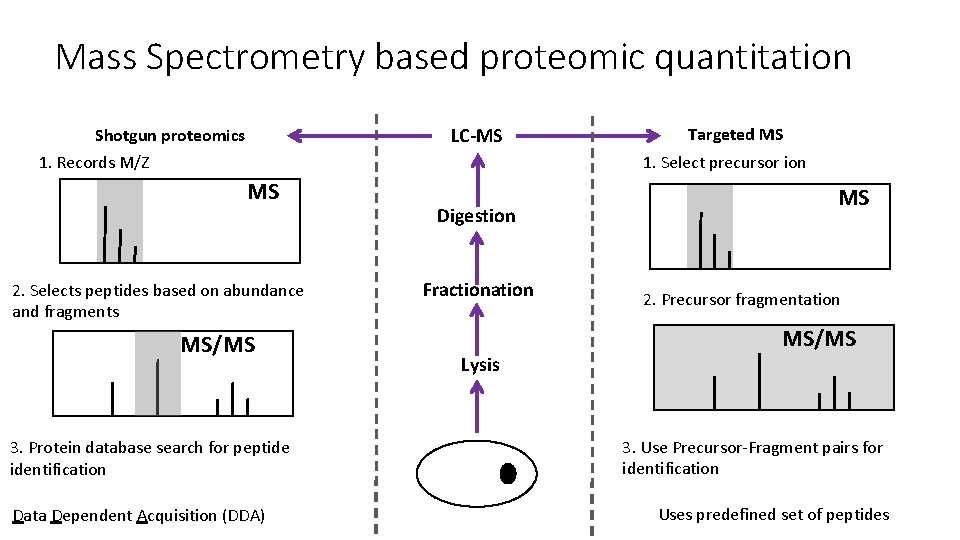
Mass Spectrometry based proteomic quantitation LC-MS Shotgun proteomics 1. Records M/Z Targeted MS 1. Select precursor ion MS 2. Selects peptides based on abundance and fragments MS/MS 3. Protein database search for peptide identification Data Dependent Acquisition (DDA) Digestion Fractionation Lysis MS 2. Precursor fragmentation MS/MS 3. Use Precursor-Fragment pairs for identification Uses predefined set of peptides

Multiple Reaction Monitoring (MRM) • • Triple Quadrupole acts as ion filters Precursor selected in first mass analyzer (Q 1) Fragmented by collision activated dissociation (Q 2) One or several of the fragments are specifically measured in the second mass analyzer (Q 3)
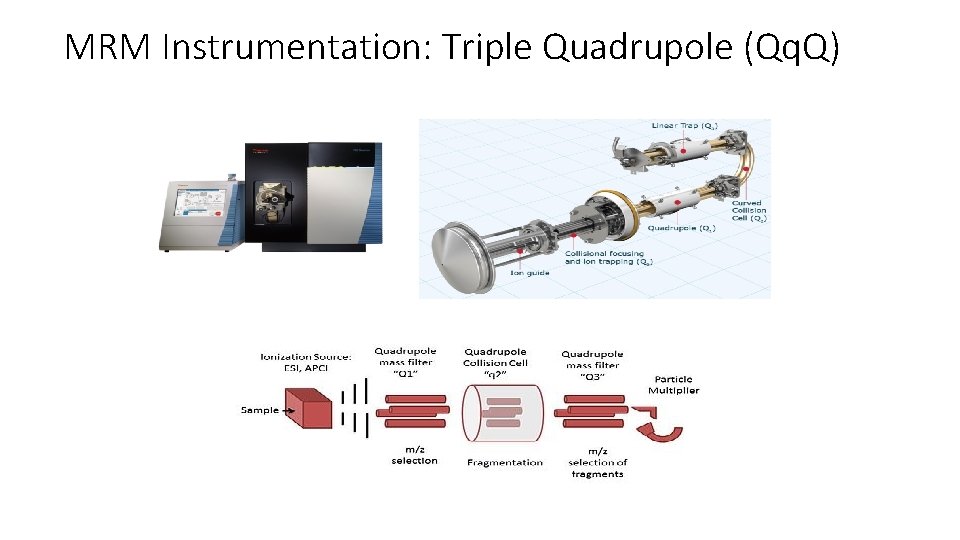
MRM Instrumentation: Triple Quadrupole (Qq. Q)

Fragment Ion Detection
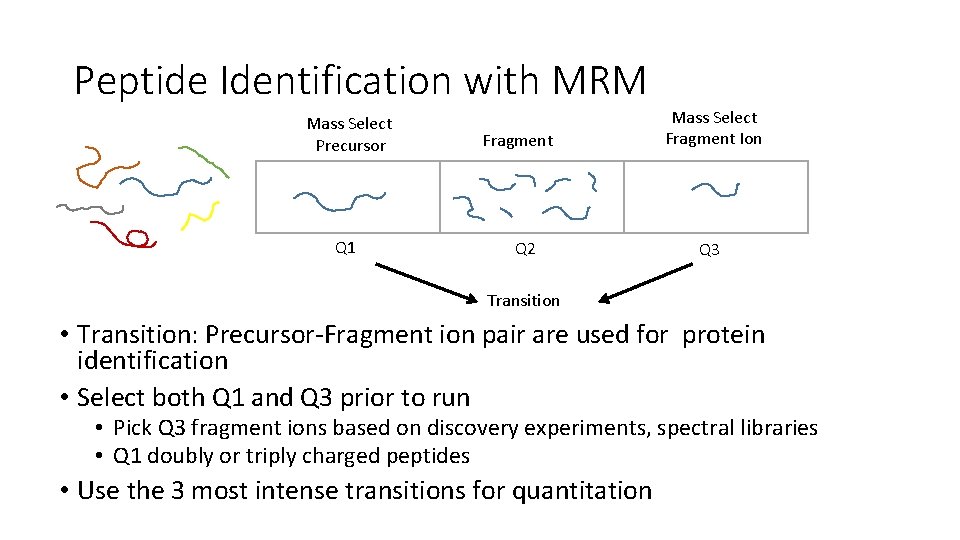
Peptide Identification with MRM Mass Select Precursor Q 1 Fragment Q 2 Mass Select Fragment Ion Q 3 Transition • Transition: Precursor-Fragment ion pair are used for protein identification • Select both Q 1 and Q 3 prior to run • Pick Q 3 fragment ions based on discovery experiments, spectral libraries • Q 1 doubly or triply charged peptides • Use the 3 most intense transitions for quantitation
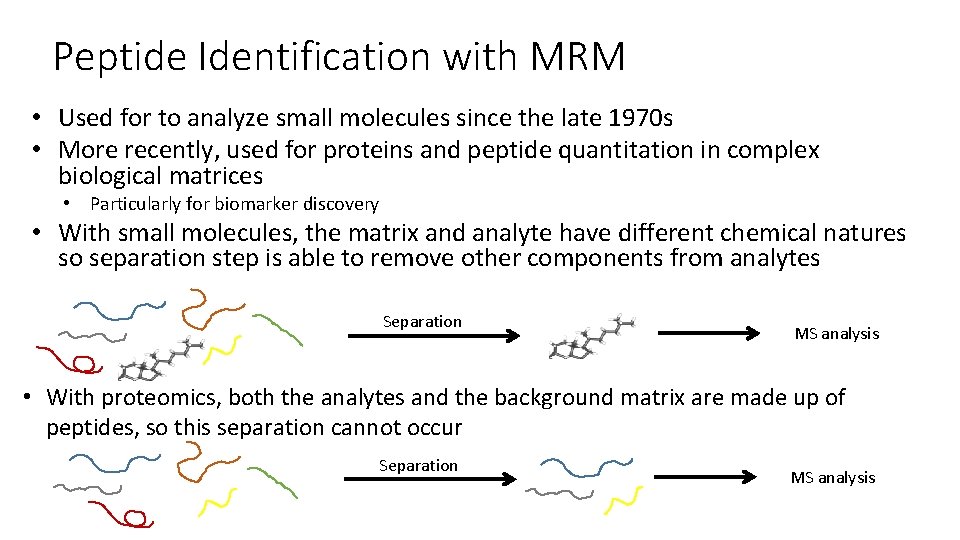
Peptide Identification with MRM • Used for to analyze small molecules since the late 1970 s • More recently, used for proteins and peptide quantitation in complex biological matrices • Particularly for biomarker discovery • With small molecules, the matrix and analyte have different chemical natures so separation step is able to remove other components from analytes Separation MS analysis • With proteomics, both the analytes and the background matrix are made up of peptides, so this separation cannot occur Separation MS analysis

Strengths of MRM • Can detect multiple transitions on the order of 10 msec per transition • Can analyze many peptides (100 s) per assay and the monitoring of many transitions per peptide • High sensitivity • High reproducibility • Detects low level analytes even in complex matrix • Golden standard for quantitation!

Weaknesses of MRM • Focuses on defined set of peptide candidates • Need to know charge state, retention time and relative product ion intensities before experimentation • Physical limit to the number of transitions that can be measured at once • Can get around this by using time-scheduled MRM, monitor transitions for a peptide in small window near retention time
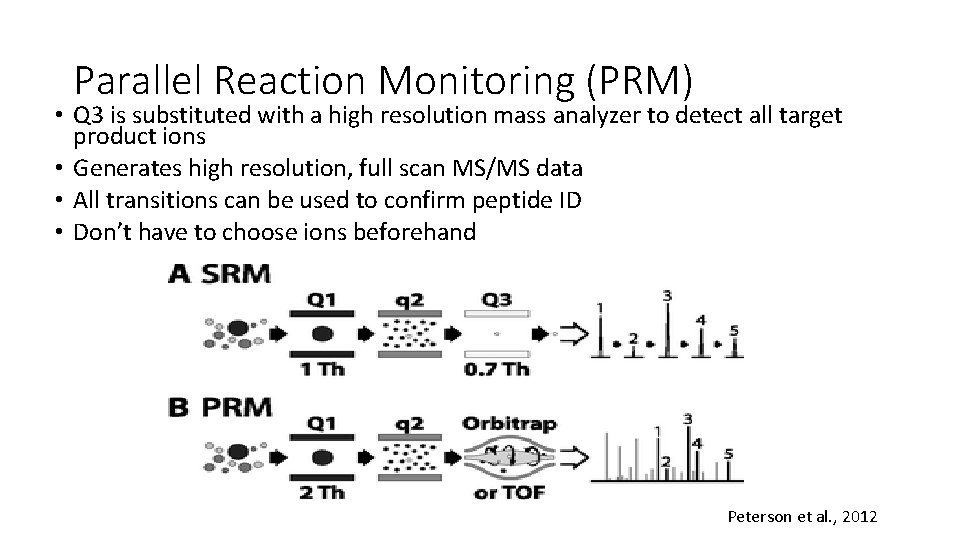
Parallel Reaction Monitoring (PRM) • Q 3 is substituted with a high resolution mass analyzer to detect all target product ions • Generates high resolution, full scan MS/MS data • All transitions can be used to confirm peptide ID • Don’t have to choose ions beforehand Peterson et al. , 2012

PRM Instrumentation: Quadrupole Time of Flight (Qqtof) The third quadrupole is replaced with a time of flight (TOF) mass analyzer offering high sensitivity, mass resolution and mass accuracy for both precursor and product ion spectra
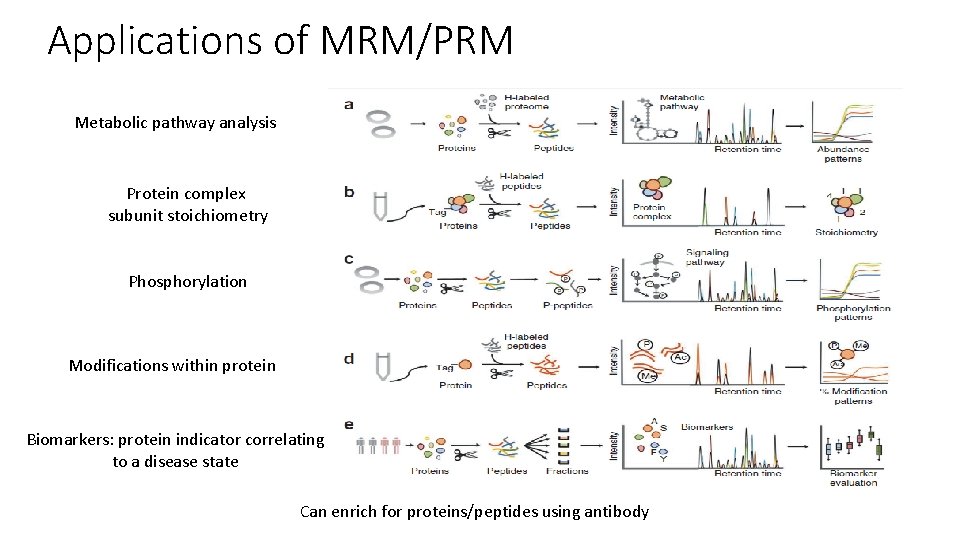
Applications of MRM/PRM Metabolic pathway analysis Protein complex subunit stoichiometry Phosphorylation Modifications within protein Biomarkers: protein indicator correlating to a disease state Can enrich for proteins/peptides using antibody

Motivating example: AKT and breast cancer
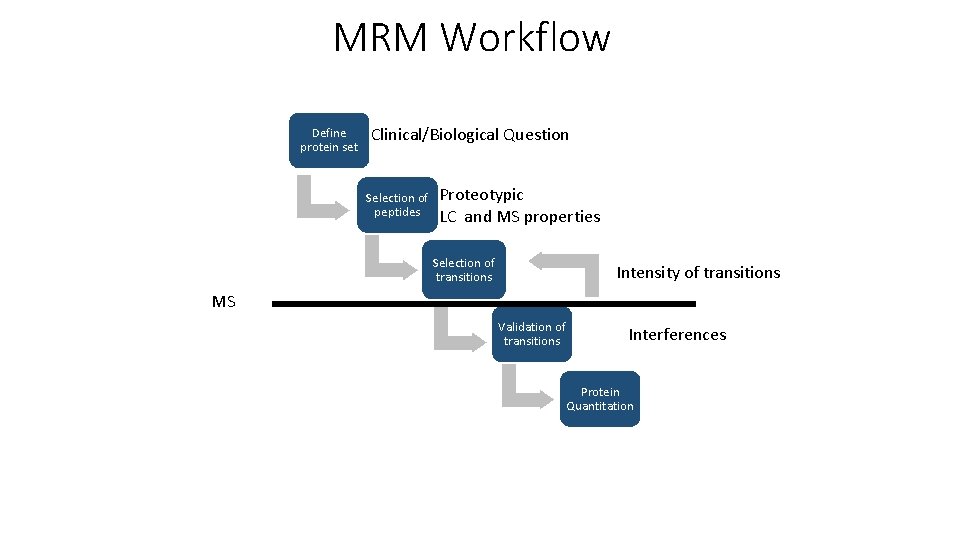
MRM Workflow Define protein set Clinical/Biological Question Selection of peptides Proteotypic LC and MS properties Selection of transitions Intensity of transitions MS Validation of transitions Interferences Protein Quantitation
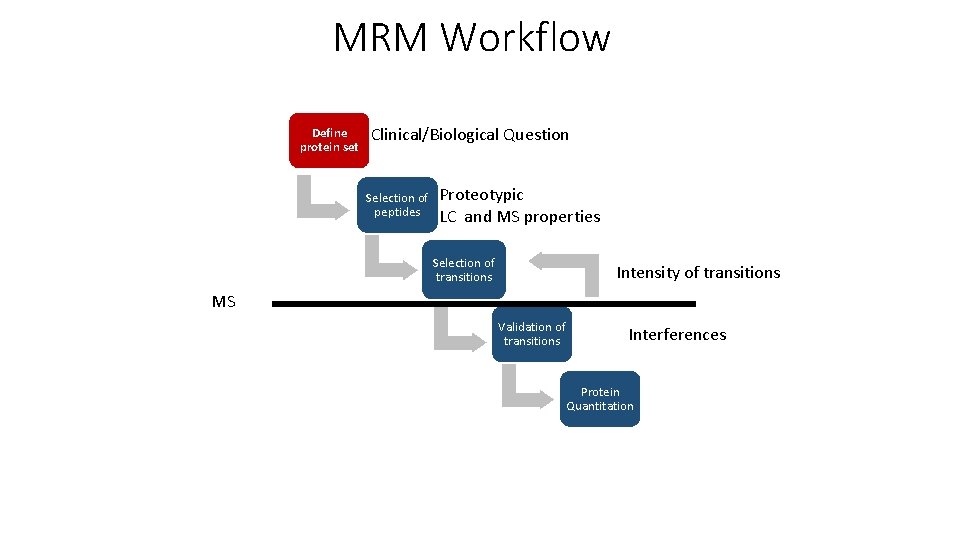
MRM Workflow Define protein set Clinical/Biological Question Selection of peptides Proteotypic LC and MS properties Selection of transitions Intensity of transitions MS Validation of transitions Interferences Protein Quantitation
Define the set of proteins based on a biological question
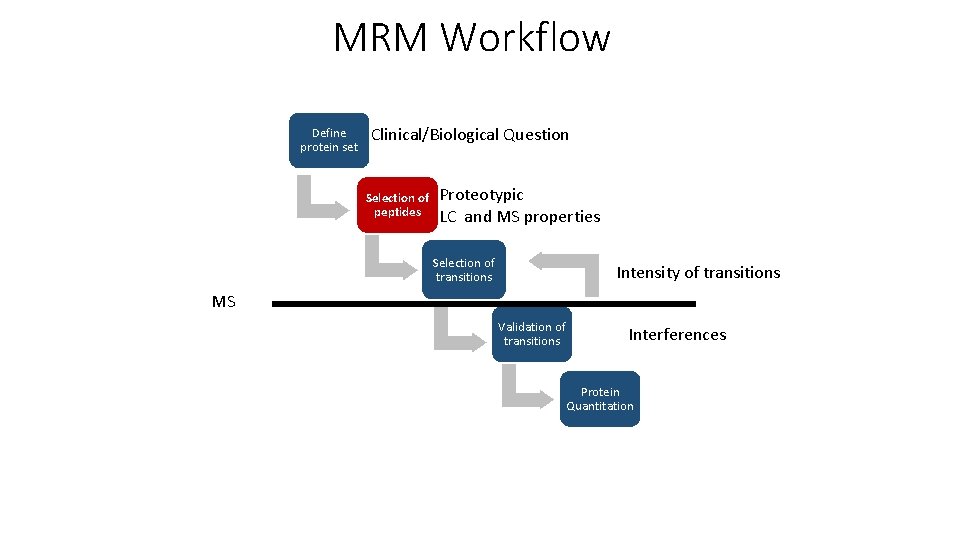
MRM Workflow Define protein set Clinical/Biological Question Selection of peptides Proteotypic LC and MS properties Selection of transitions Intensity of transitions MS Validation of transitions Interferences Protein Quantitation

Selecting Peptides • A few representative peptides will be used to quantify each protein • Need to fulfill certain characteristics • • Have an unique sequence Consistently observed by LC-MS methods 8 -25 amino acids Good ionization efficiency m/z within the range of the instrument No missed cleavages Not too hydrophillic (poorly retained) or hydrophobic (may stick to column)
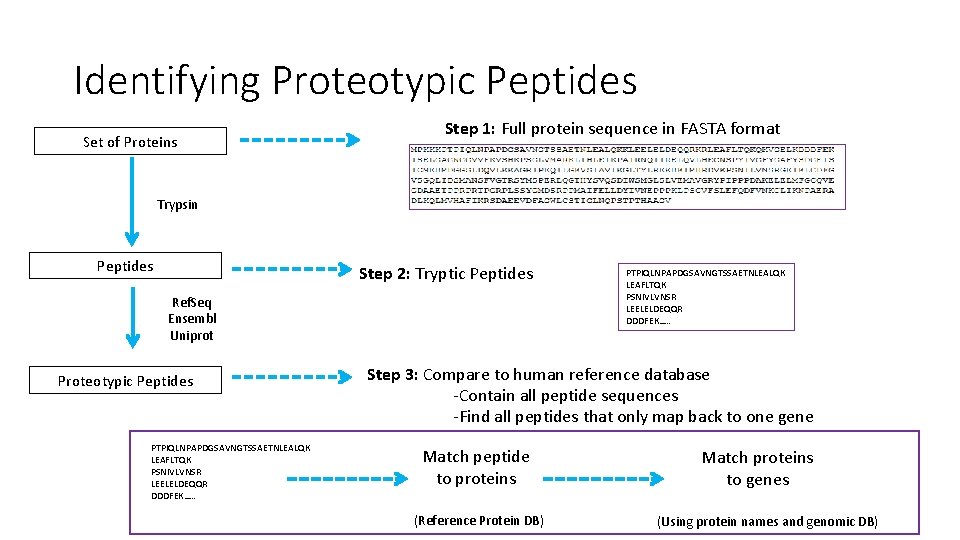
Identifying Proteotypic Peptides Set of Proteins Step 1: Full protein sequence in FASTA format Trypsin Peptides Step 2: Tryptic Peptides Ref. Seq Ensembl Uniprot Proteotypic Peptides PTPIQLNPAPDGSAVNGTSSAETNLEALQK LEAFLTQK PSNIVLVNSR LEELELDEQQR DDDFEK…. . Step 3: Compare to human reference database -Contain all peptide sequences -Find all peptides that only map back to one gene Match peptide to proteins (Reference Protein DB) Match proteins to genes (Using protein names and genomic DB)
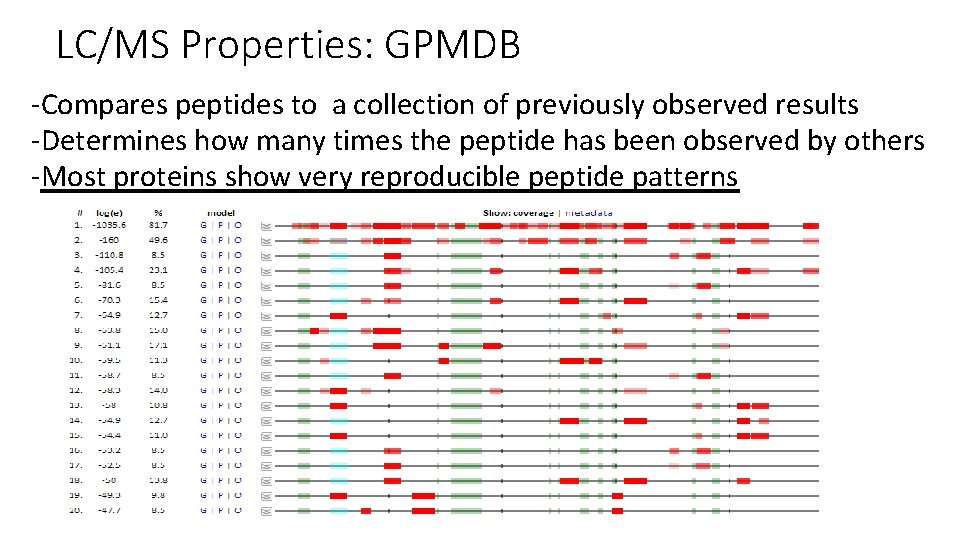
LC/MS Properties: GPMDB -Compares peptides to a collection of previously observed results -Determines how many times the peptide has been observed by others -Most proteins show very reproducible peptide patterns
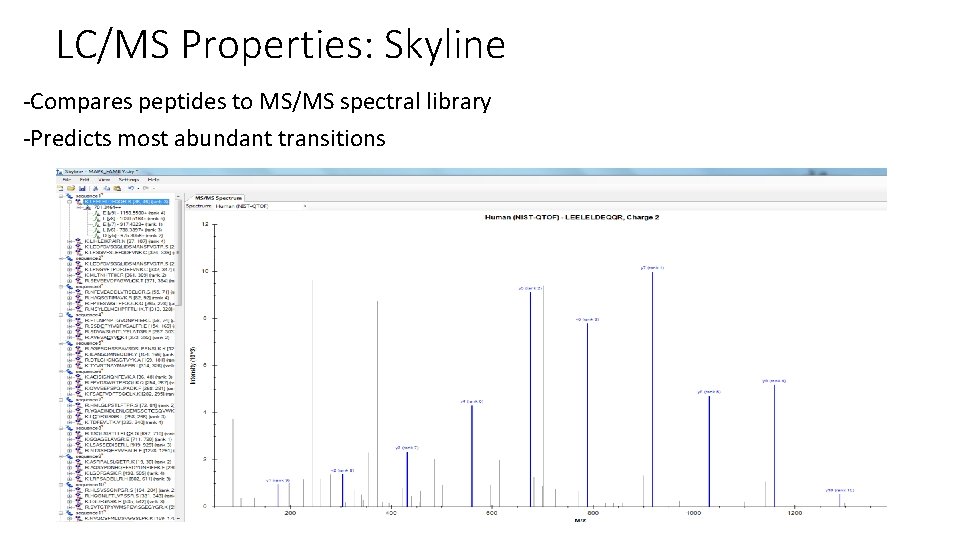
LC/MS Properties: Skyline -Compares peptides to MS/MS spectral library -Predicts most abundant transitions
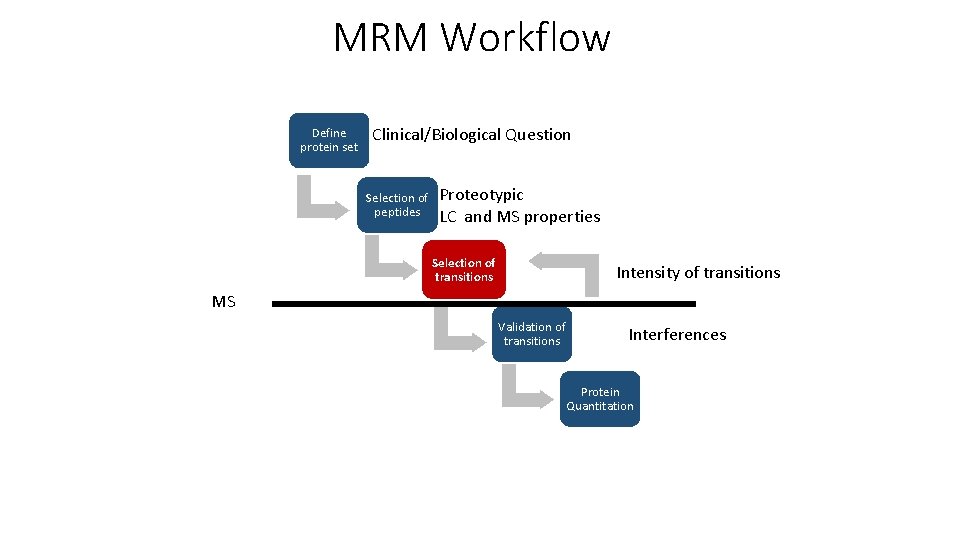
MRM Workflow Define protein set Clinical/Biological Question Selection of peptides Proteotypic LC and MS properties Selection of transitions Intensity of transitions MS Validation of transitions Interferences Protein Quantitation

Selecting Transitions • Limitation of MRM-MS: ~1 -2 m/z unit window for precursor and fragment ion occasionally let in interfering peptides with similar characteristics • If we want to use these transitions for quantitation, we need to be confident there are no interferences • Largest always largest, smallest always smallest etc. • b-fragments of high m/z are less represented on Qq. Q MRM

Selecting Transitions • Limitation of MRM-MS: ~1 -2 m/z unit window for precursor and fragment ion occasionally let in interfering peptides with similar characteristics • If we want to use these transitions for quantitation, we need to be confident there are no interferences • Largest always largest, smallest always smallest etc. • b-fragments of high m/z are less represented on Qq. Q MRM Peptide of interest Interfering peptide
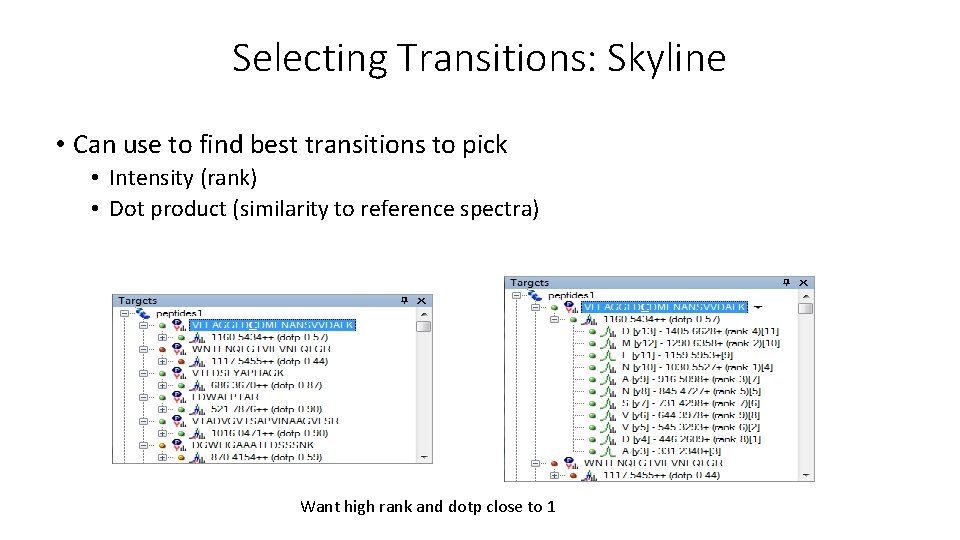
Selecting Transitions: Skyline • Can use to find best transitions to pick • Intensity (rank) • Dot product (similarity to reference spectra) Want high rank and dotp close to 1
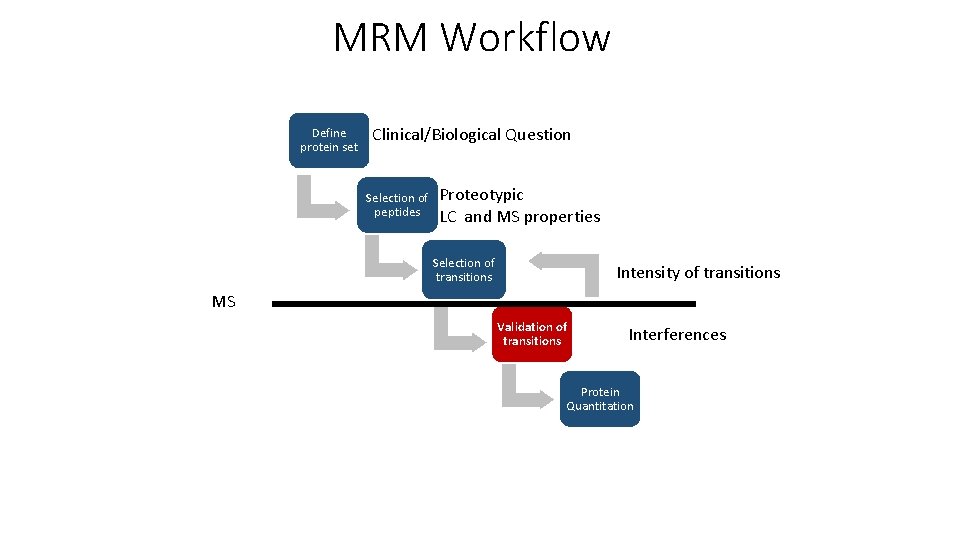
MRM Workflow Define protein set Clinical/Biological Question Selection of peptides Proteotypic LC and MS properties Selection of transitions Intensity of transitions MS Validation of transitions Interferences Protein Quantitation
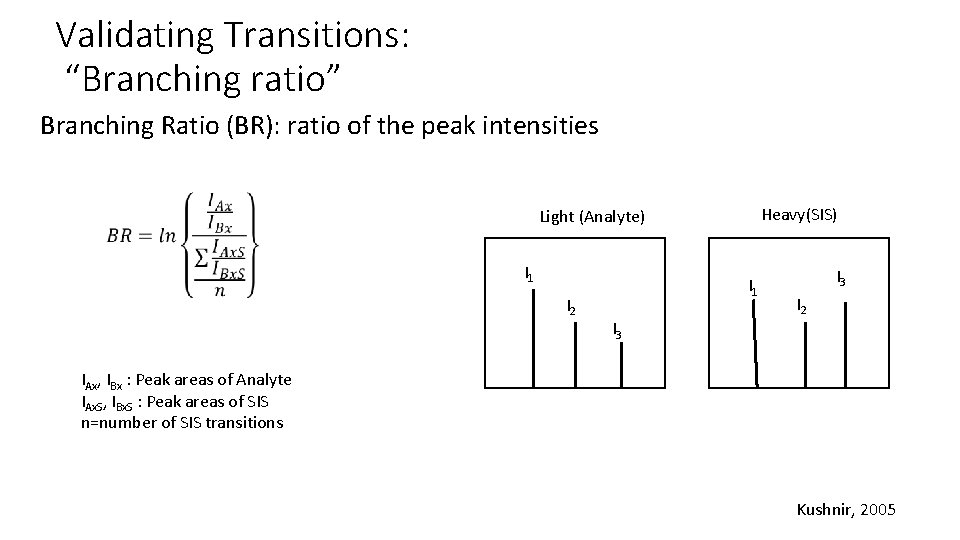
Validating Transitions: “Branching ratio” Branching Ratio (BR): ratio of the peak intensities Heavy(SIS) Light (Analyte) I 1 I 2 I 1 I 3 I 2 IAx, IBx : Peak areas of Analyte IAx. S, IBx. S : Peak areas of SIS n=number of SIS transitions Kushnir, 2005
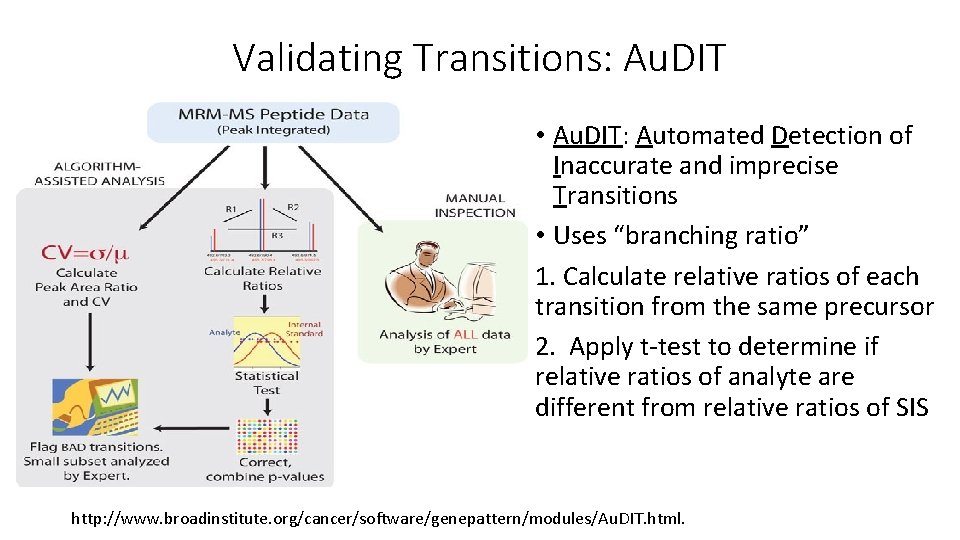
Validating Transitions: Au. DIT • Au. DIT: Automated Detection of Inaccurate and imprecise Transitions • Uses “branching ratio” 1. Calculate relative ratios of each transition from the same precursor 2. Apply t-test to determine if relative ratios of analyte are different from relative ratios of SIS http: //www. broadinstitute. org/cancer/software/genepattern/modules/Au. DIT. html.
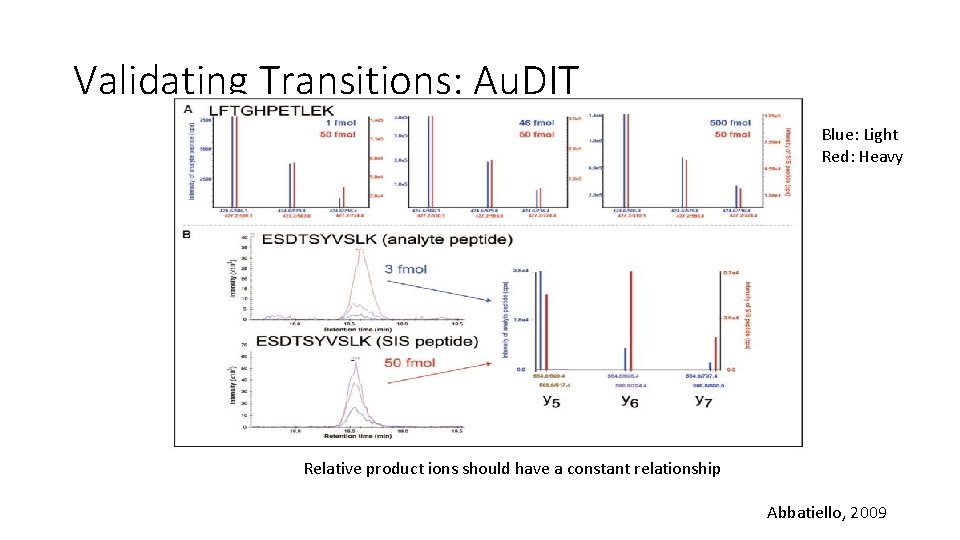
Validating Transitions: Au. DIT Blue: Light Red: Heavy Relative product ions should have a constant relationship Abbatiello, 2009
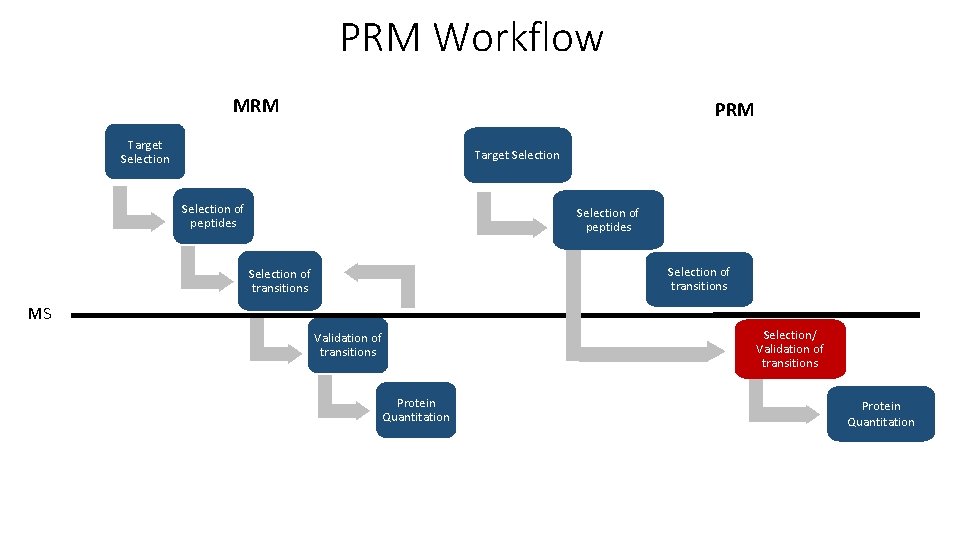
PRM Workflow MRM PRM Target Selection of peptides Selection of transitions MS Selection/ Validation of transitions Protein Quantitation
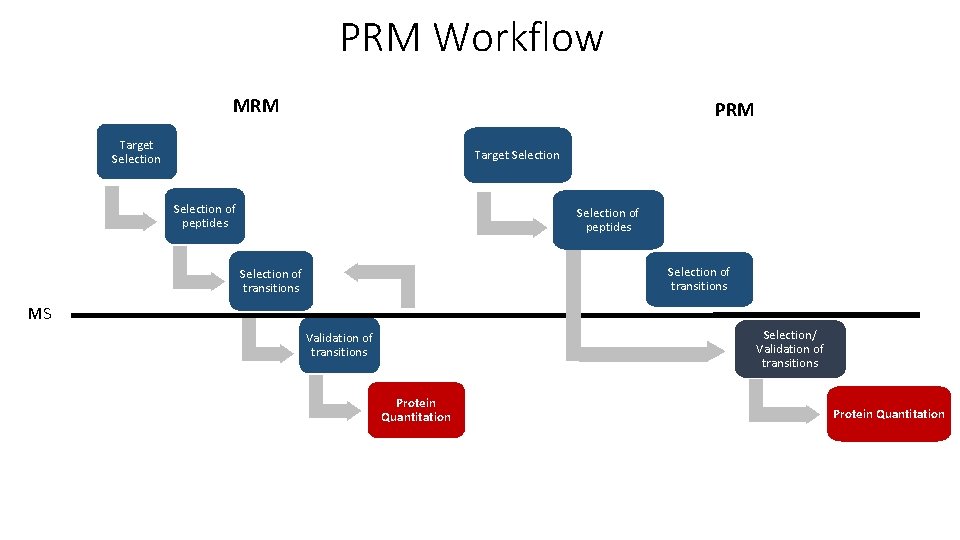
PRM Workflow MRM PRM Target Selection of peptides Selection of transitions MS Selection/ Validation of transitions Protein Quantitation

Protein Quantitation

Label-free quantitation • Usually use 3 or more precursor-product ion pairs (transitions) for quantitation • Relies on direct evaluation of MS signal intensities of naturally occurring peptides in a sample. • Simple and straightforward • Low precision • Several peptides for each protein should be quantified to avoid false quantitation
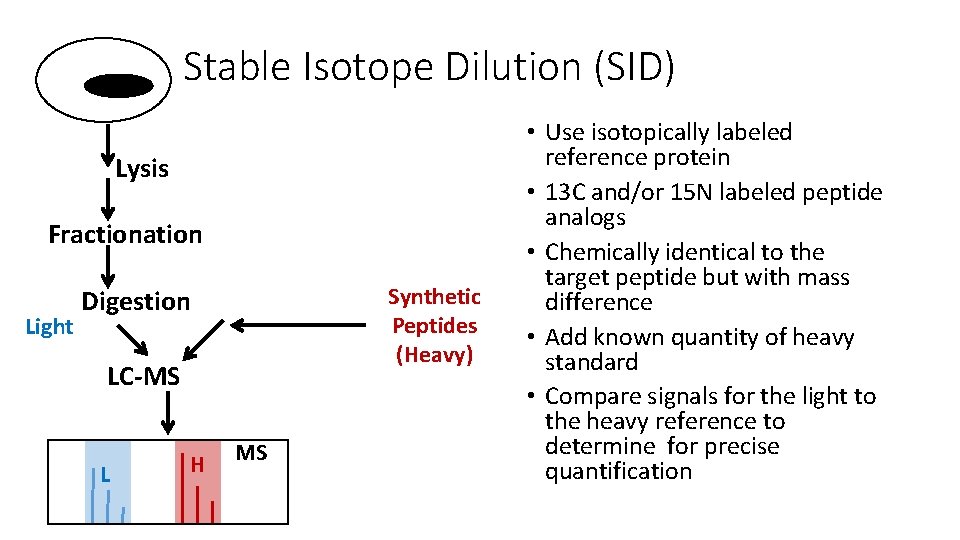
Stable Isotope Dilution (SID) Lysis Fractionation Light Synthetic Peptides (Heavy) Digestion LC-MS L H MS • Use isotopically labeled reference protein • 13 C and/or 15 N labeled peptide analogs • Chemically identical to the target peptide but with mass difference • Add known quantity of heavy standard • Compare signals for the light to the heavy reference to determine for precise quantification

Quantitation Details L Analyte MS H SIS: Stable Isotope Standard PAR: Peak Area Ratio SIS PAR = Light (Analyte) Peak Area Heavy (SIS) Peak Area Analyte concentration= PAR*SIS peptide concentration -Use at least 3 transitions -Have to make sure these transitions do not have interferences
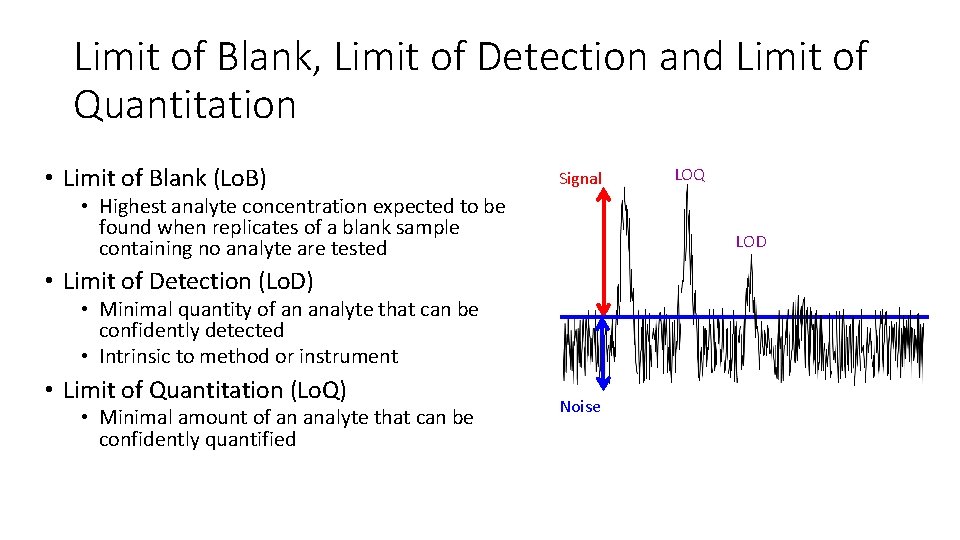
Limit of Blank, Limit of Detection and Limit of Quantitation • Limit of Blank (Lo. B) Signal • Highest analyte concentration expected to be found when replicates of a blank sample containing no analyte are tested LOD • Limit of Detection (Lo. D) • Minimal quantity of an analyte that can be confidently detected • Intrinsic to method or instrument • Limit of Quantitation (Lo. Q) • Minimal amount of an analyte that can be confidently quantified LOQ Noise
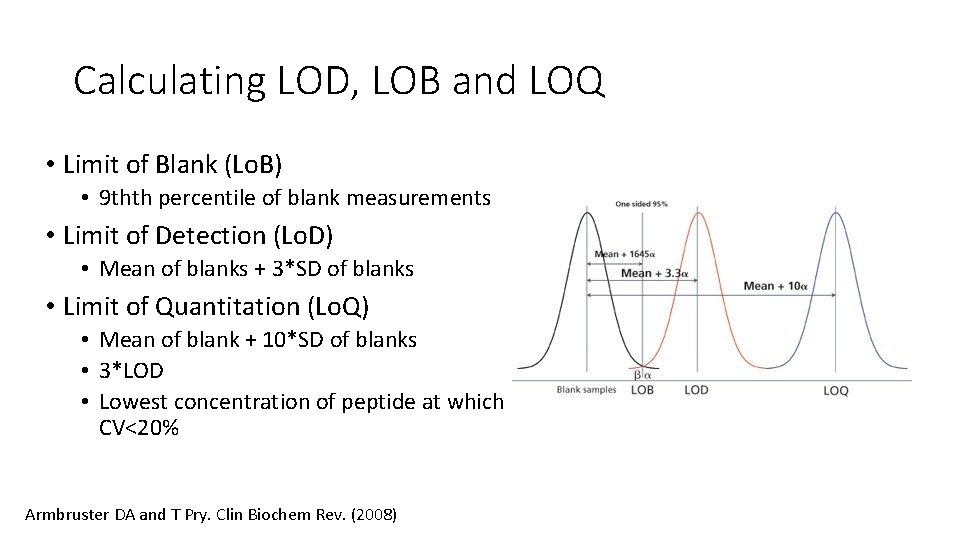
Calculating LOD, LOB and LOQ • Limit of Blank (Lo. B) • 9 thth percentile of blank measurements • Limit of Detection (Lo. D) • Mean of blanks + 3*SD of blanks • Limit of Quantitation (Lo. Q) • Mean of blank + 10*SD of blanks • 3*LOD • Lowest concentration of peptide at which CV<20% Armbruster DA and T Pry. Clin Biochem Rev. (2008)

Standard Curve • Standard concentration curve is generated using increasing concentrations of the “heavy peptide” in the presence of a constant amount of biological sample (aka matrix) • Matrix contains the “light” version of the peptide • Standard Curve typically created using: • 5 heavy peptide serial dilutions • Blank (no heavy) • Once the curve is generated, biological samples can be processed measuring the amount of endogenous “light” peptide in each sample • Using the calibration curve, we can compute the endogenous concentration from the ratio of endogenous/heavy (PAR)
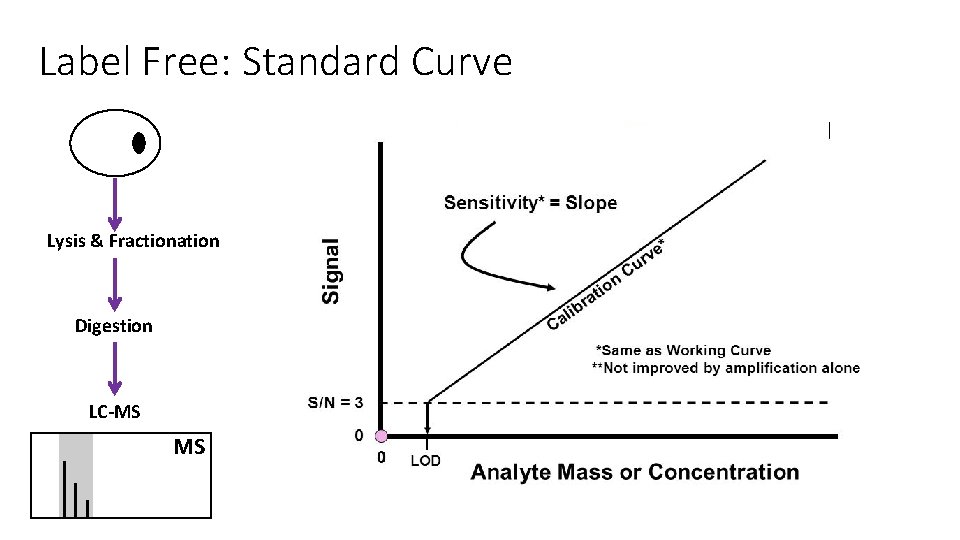
Label Free: Standard Curve Lysis & Fractionation Digestion LC-MS MS

Lower Limit of Detection The lowest amount of analyte that is statistically distinguishable from background or a negative control. Two methods to determine lower limit of detection: 1. Lowest concentration of the analyte where CV is less than for example 20%. 2. Determine level of blank by taking 95 th percentile of the blank measurements and add a constant times the standard deviation of the lowest concentration. K. Linnet and M. Kondratovich, Partly Nonparametric Approach for Determining the Limit of Detection, Clinical Chemistry 50 (2004) 732– 740.
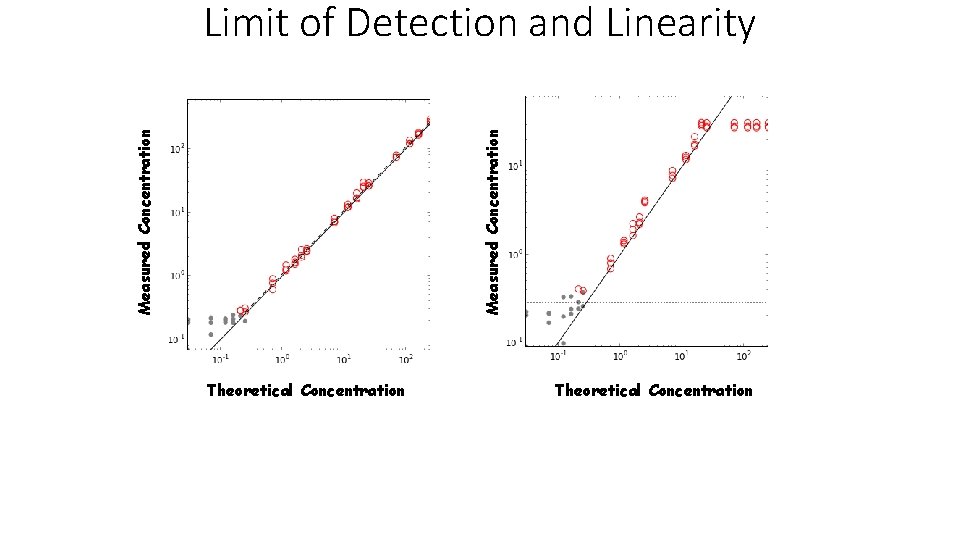
Measured Concentration Limit of Detection and Linearity Theoretical Concentration
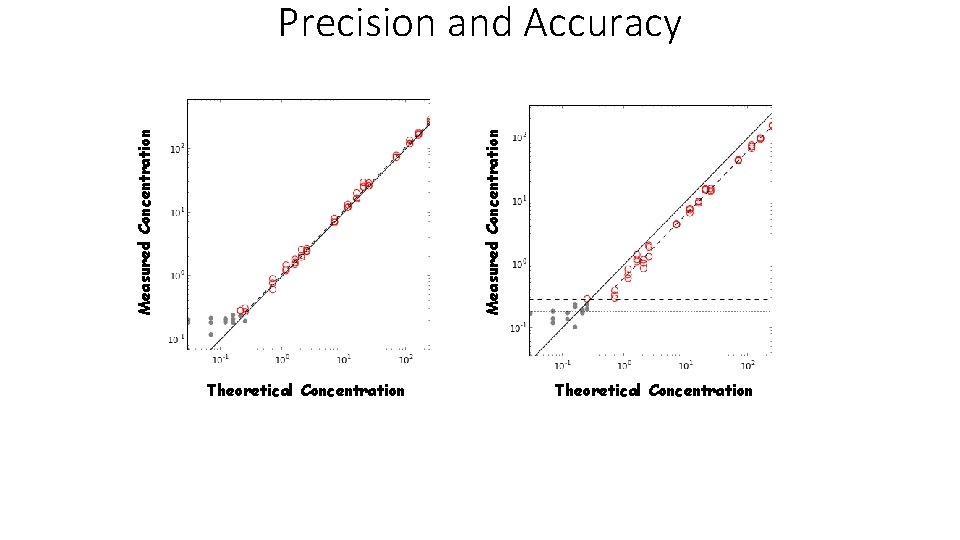
Measured Concentration Precision and Accuracy Theoretical Concentration
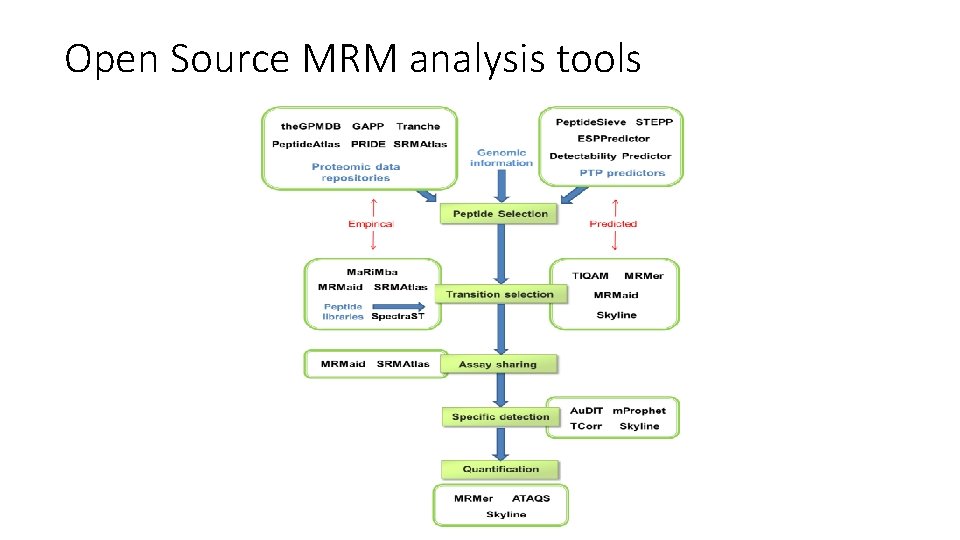
Open Source MRM analysis tools

Data Independent Acquisition
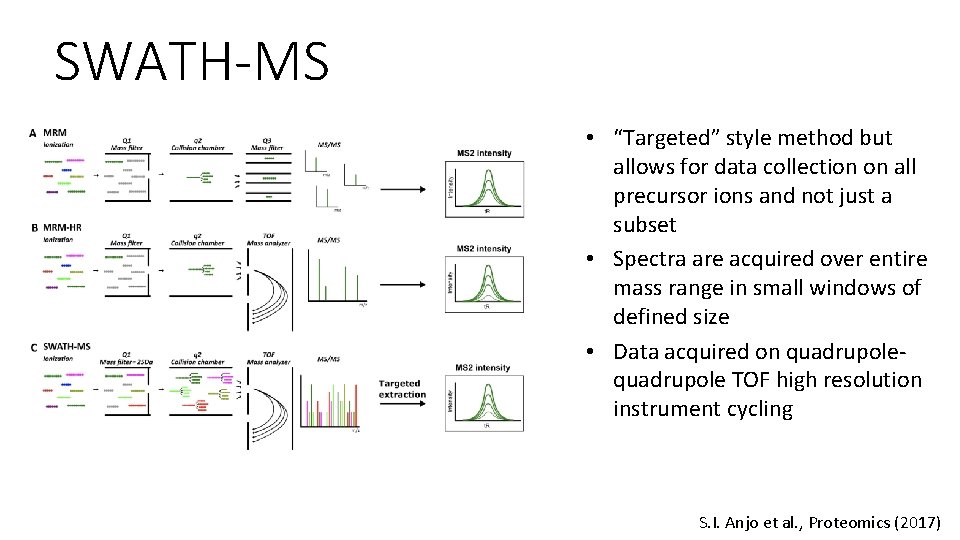
SWATH-MS • “Targeted” style method but allows for data collection on all precursor ions and not just a subset • Spectra are acquired over entire mass range in small windows of defined size • Data acquired on quadrupole TOF high resolution instrument cycling S. I. Anjo et al. , Proteomics (2017)
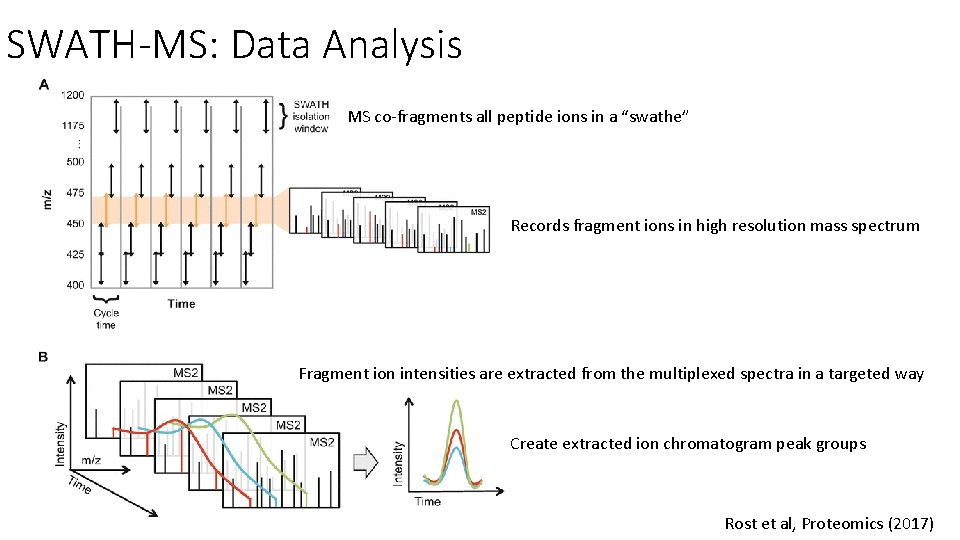
SWATH-MS: Data Analysis MS co-fragments all peptide ions in a “swathe” Records fragment ions in high resolution mass spectrum Fragment ion intensities are extracted from the multiplexed spectra in a targeted way Create extracted ion chromatogram peak groups Rost et al, Proteomics (2017)
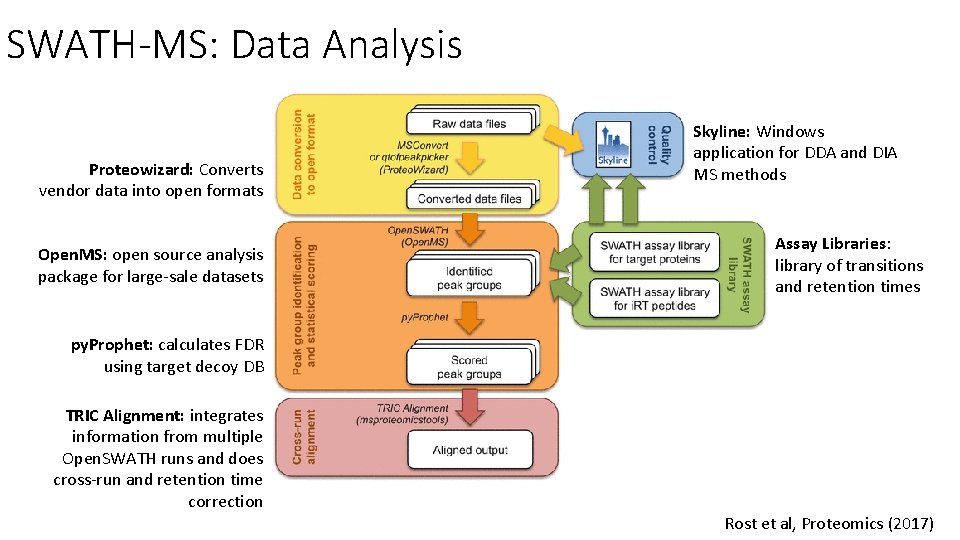
SWATH-MS: Data Analysis Proteowizard: Converts vendor data into open formats Open. MS: open source analysis package for large-sale datasets Skyline: Windows application for DDA and DIA MS methods Assay Libraries: library of transitions and retention times py. Prophet: calculates FDR using target decoy DB TRIC Alignment: integrates information from multiple Open. SWATH runs and does cross-run and retention time correction Rost et al, Proteomics (2017)

Summary • Targeted proteomics can be used as an alternative to antibody-based protein assays in hypothesis driven biological experiments. • Allows for multiplexed analysis of many proteins in different conditions. • PRM removes the step of transition validation and allows for all computational analysis post-acquisition. • SWATH removes the step of peptide selection and generates transitions for all precursor ions in the defined precursor retention time and m/z • Transition interference must be appropriately identified prior to protein quantitation. • Several computational tools to predict and identify interferences in MRM and PRM have been developed.

Questions?
- Slides: 54