Cyberinfrastructure for Top Down Proteomics RYAN FELLERS PROTEOMICS

Cyberinfrastructure for Top Down Proteomics RYAN FELLERS, PROTEOMICS CENTER OF EXCELLENCE OSG ALL HANDS MEETING, 3/24/2015

Agenda Introduction to Proteomics ◦ What is it? ◦ Top Down Proteomics specifics Tools and Support ◦ Legacy of Tool Development ◦ New Project Future Directions

Proteins and Disease Alzheimer’s Disease • Amyloid Beta Parkinson’s Disease • Alpha synuclein Sickle Cell Anemia • Hemoglobin Ronald Reagan Muhammad Ali and Michael J. Fox Miles Davis

What is Proteomics? Proteomics as coined by Wilkins and Williams: “The study of proteins as a result from genomics. ”
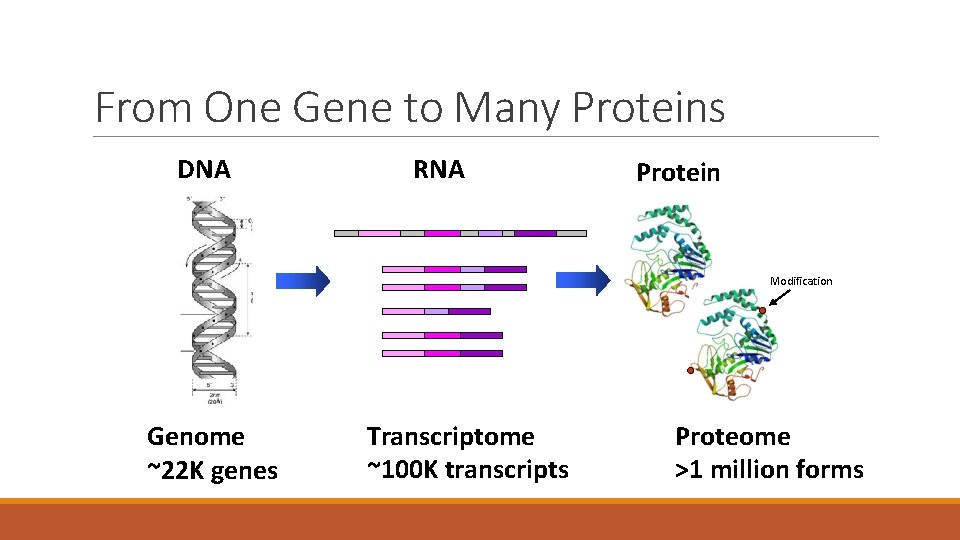
From One Gene to Many Proteins DNA RNA Protein Modification Genome ~22 K genes Transcriptome ~100 K transcripts Proteome >1 million forms
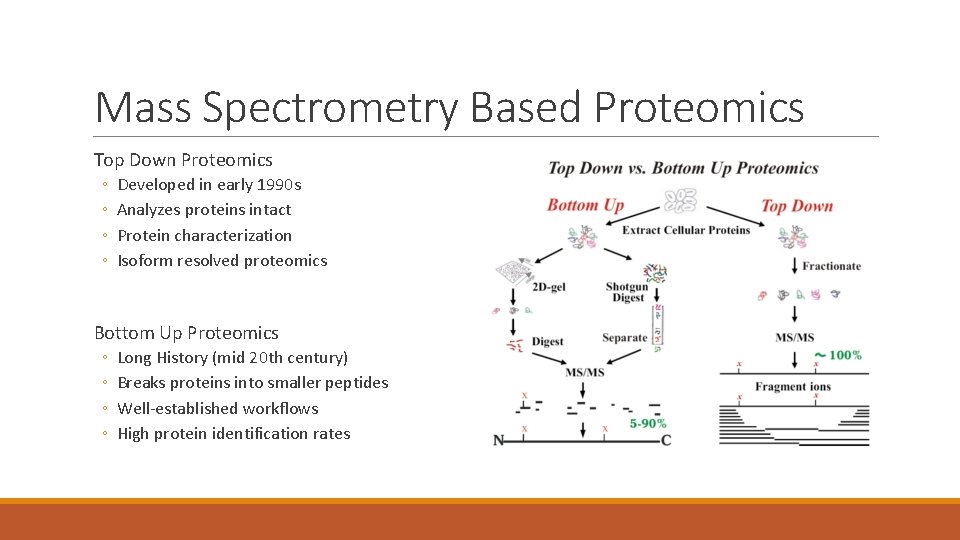
Mass Spectrometry Based Proteomics Top Down Proteomics ◦ ◦ Developed in early 1990 s Analyzes proteins intact Protein characterization Isoform resolved proteomics Bottom Up Proteomics ◦ ◦ Long History (mid 20 th century) Breaks proteins into smaller peptides Well-established workflows High protein identification rates
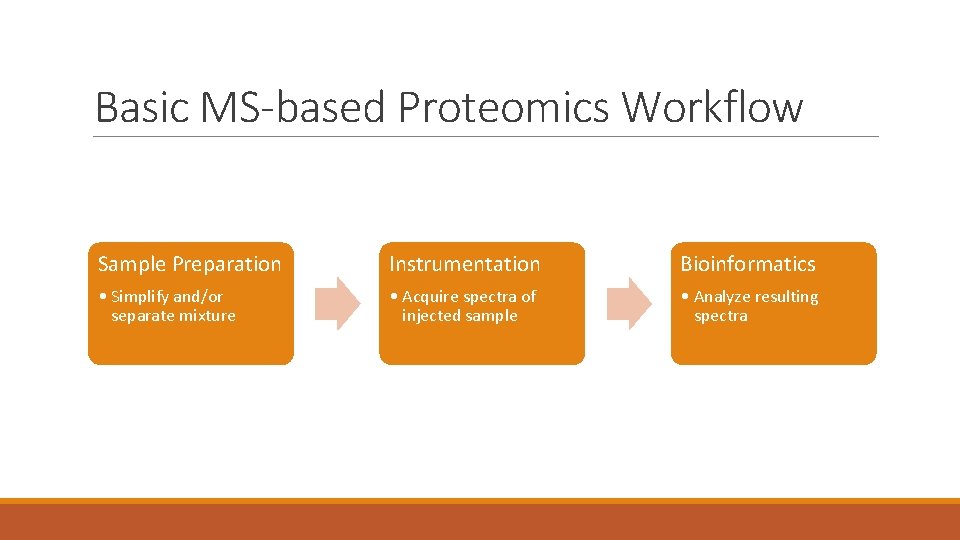
Basic MS-based Proteomics Workflow Sample Preparation Instrumentation Bioinformatics • Simplify and/or separate mixture • Acquire spectra of injected sample • Analyze resulting spectra
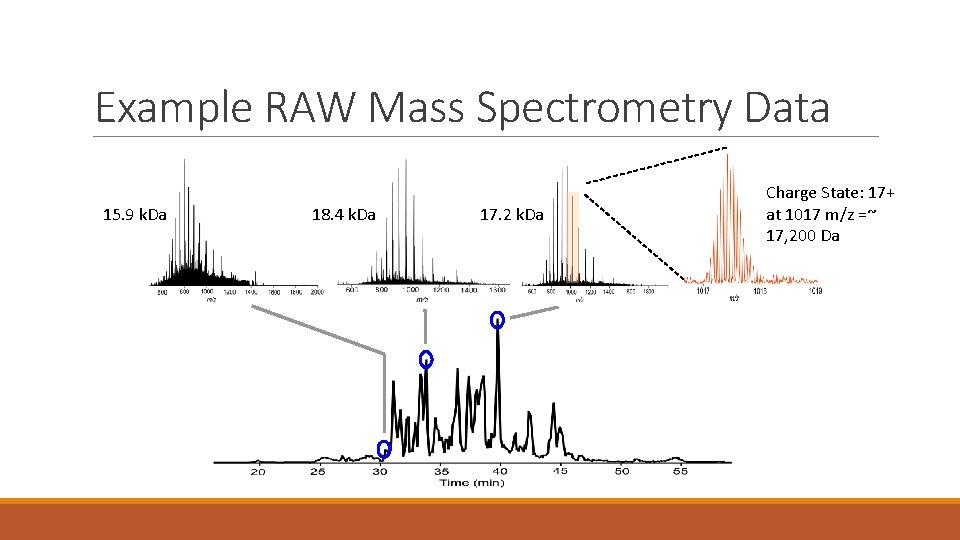
Example RAW Mass Spectrometry Data 15. 9 k. Da 18. 4 k. Da 17. 2 k. Da Charge State: 17+ at 1017 m/z =~ 17, 200 Da

Tools and Support HOW DO WE SERVE OUR USERS?

Development History Commercial products in partnership with Thermo Fisher Scientific ◦ Pro. Sight. PC ◦ Version 1. 0 (2006) ◦ Version 2. 0 (2009) ◦ Version 3. 0 (2012) ◦ Pro. Sight. PD (2015) Freeware released through the university ◦ Pro. Sight. PTM (2002) ◦ Pro. Sight Lite (2014) Internal, customized HPC ◦ Rocks 5. 3 (2010)

Limitations of our Toolset. NET is a Windows only technology ◦ We use Mono Foundation to run on Linux Compute is difficult to scale ◦ We would like to be able to move our compute between different resources as needed It has been difficult to standardize ◦ Users are met with too many choices and a complicated user experience (UX)

Our Solution - Galaxy Why Galaxy? ◦ ◦ ◦ It is ubiquitous in other “-omics” Makes search accessible through a web portal Allows us to utilize multiple HPC systems There exist some proteomics tools (Galaxy. P) It has a very active community Started project 3 months ago and plan to be live internally by late Spring / early Summer
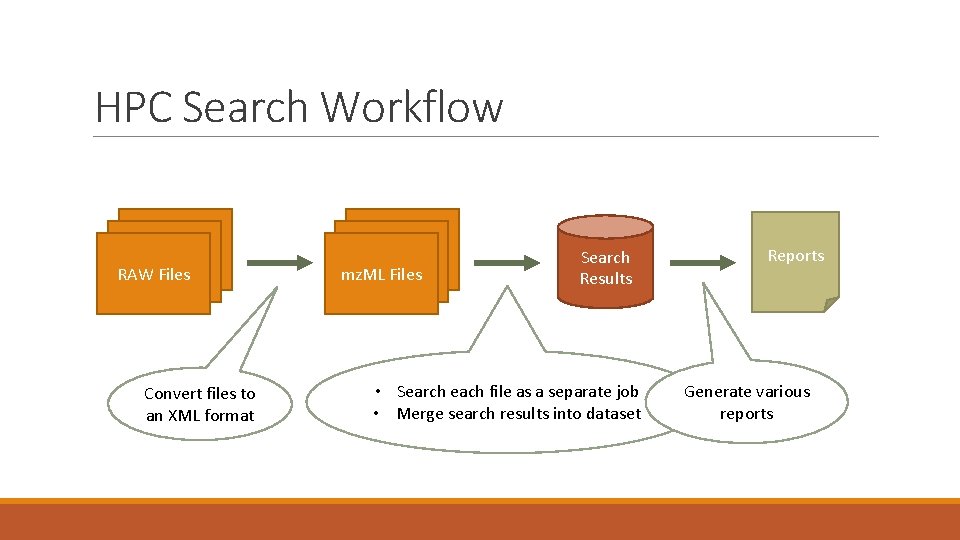
HPC Search Workflow RAW Files Convert files to an XML format mz. ML Files Search Results • Search each file as a separate job • Merge search results into dataset Reports Generate various reports
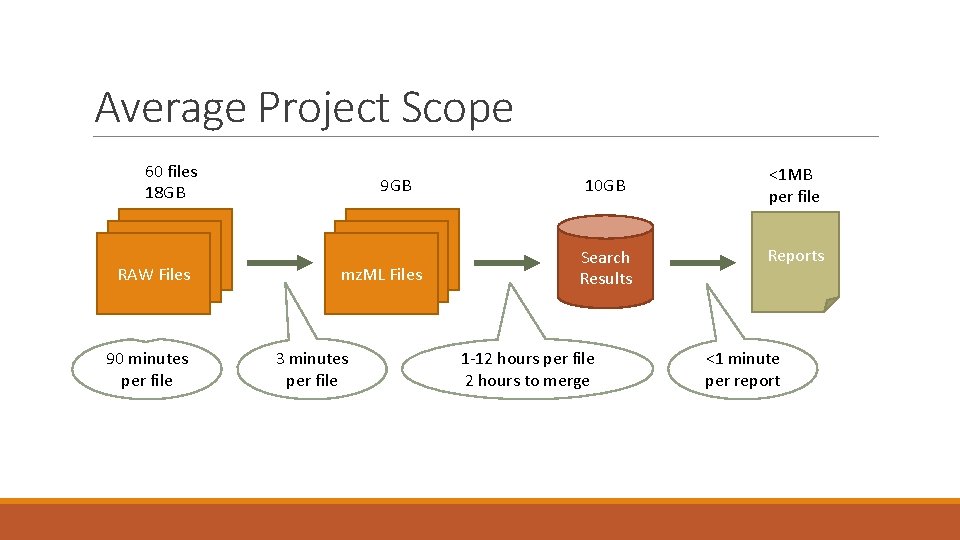
Average Project Scope 60 files 18 GB RAW Files 90 minutes per file 9 GB mz. ML Files 3 minutes per file 10 GB Search Results 1 -12 hours per file 2 hours to merge <1 MB per file Reports <1 minute per report

Tools in the Search Workflow mz. ML Converter ◦ Converts RAW instrument files to mz. ML files ◦. NET console application required to run on Windows HPC Processing ◦ Compares spectra in mz. ML files against a candidate protein database to generate a list of protein results ◦. NET console application running under mono 3. x Result Merger ◦ Merges multiple protein result sets into one large result set ◦. NET console application running under mono 3. x Report Generator ◦ Executes SQL queries against a result set and outputs a comma separated value (CSV) files ◦ Python-based Galaxy tool
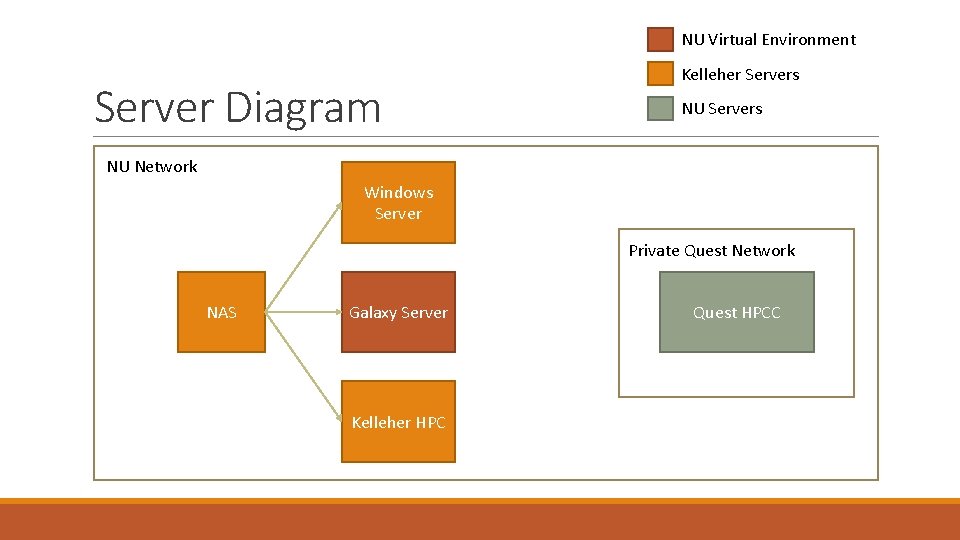
NU Virtual Environment Server Diagram Kelleher Servers NU Network Windows Server Private Quest Network NAS Galaxy Server Kelleher HPC Quest HPCC

Pulsar Server Problem ◦ We want to run our Galaxy search jobs on Quest ◦ Traditionally, Galaxy requires a shared file system to run jobs on an HPC Solution: Pulsar Server ◦ Light-weight server running on a remote resource ◦ Parameters and inputs are transferred to the remote server and results are sent back to Galaxy ◦ Requirements ◦ Rights to log onto the remote system and pull down Pulsar git repository ◦ One open port for communication
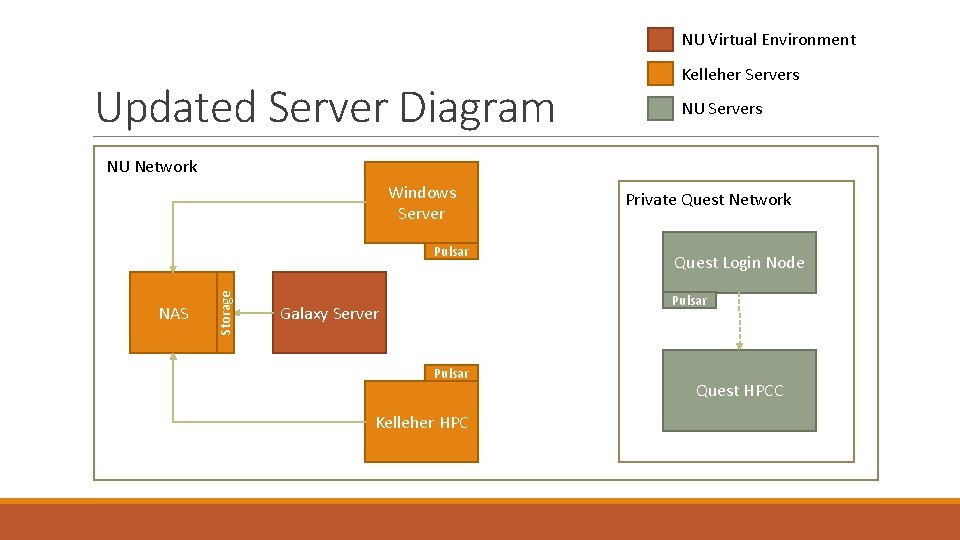
NU Virtual Environment Updated Server Diagram Kelleher Servers NU Network Windows Server NAS Storage Pulsar Private Quest Network Quest Login Node Pulsar Galaxy Server Pulsar Kelleher HPC Quest HPCC
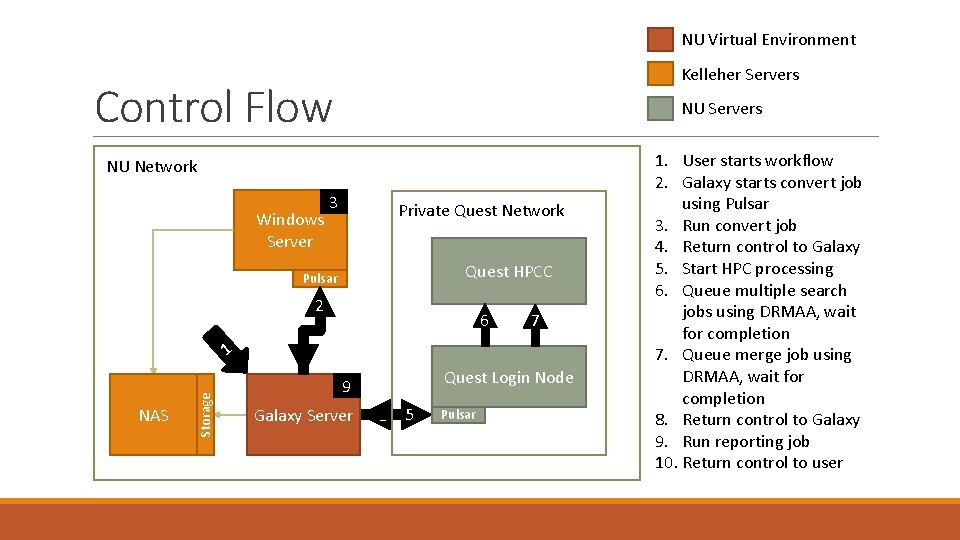
NU Virtual Environment Kelleher Servers Control Flow NU Network NU Servers 3 Windows Server 3 Private Quest Network Quest HPCC Pulsar 2 NAS Storage 1 6 7 4 Quest Login Node 9 Galaxy Server 8 5 Pulsar 1. User starts workflow 2. Galaxy starts convert job using Pulsar 3. Run convert job 4. Return control to Galaxy 5. Start HPC processing 6. Queue multiple search jobs using DRMAA, wait for completion 7. Queue merge job using DRMAA, wait for completion 8. Return control to Galaxy 9. Run reporting job 10. Return control to user
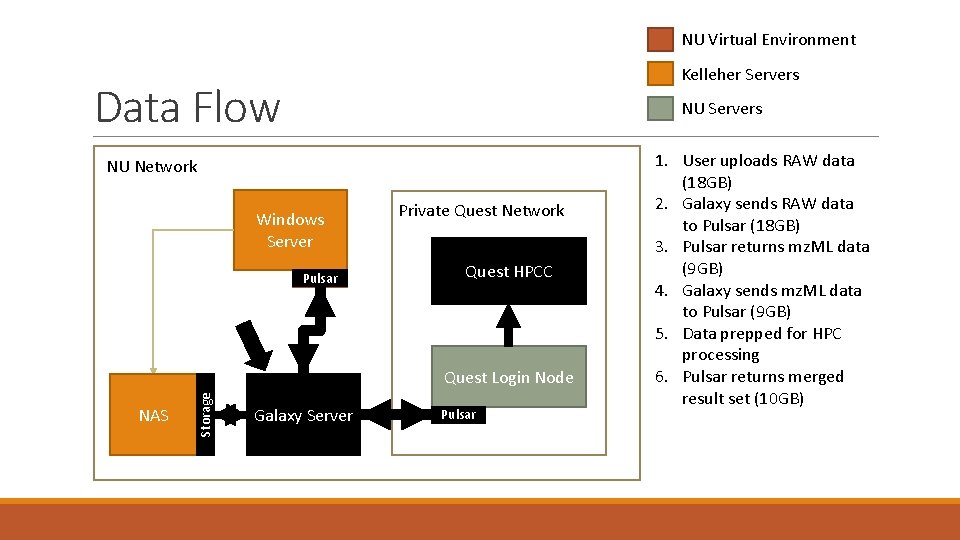
NU Virtual Environment Kelleher Servers Data Flow NU Network NU Servers 3 Windows Server Pulsar Private Quest Network Quest HPCC NAS Storage Quest Login Node Galaxy Server Pulsar 1. User uploads RAW data (18 GB) 2. Galaxy sends RAW data to Pulsar (18 GB) 3. Pulsar returns mz. ML data (9 GB) 4. Galaxy sends mz. ML data to Pulsar (9 GB) 5. Data prepped for HPC processing 6. Pulsar returns merged result set (10 GB)

Still a work in progress … Control Flow improvements ◦ Implement everything in one Galaxy workflow ◦ Make the Quest interaction more robust (progress, more parameters, etc. ) Data Flow improvements ◦ Be more efficient when moving data around NAS storage ◦ Use more efficient protocol to move data to/from Pulsar endpoints General improvements ◦ Develop more reports and useful visualizations

Long-Term Vision What we can do ◦ Expand this functionality to tools other than search ◦ Integrate with external Protein Repositories (e. g. Uni. Prot, Proteoform Repo, etc. ) The size and frequency of large projects are expected to increase ◦ Balance load according to availability ◦ Scale up to additional compute resources as needed

Acknowledgements Dr. Neil Kelleher Members of the Informatics Team ◦ Dr. Rich Le. Duc ◦ Bryan Early ◦ Joe Greer All of the members of the Kelleher Research Group and Proteomics Center of Excellence Dr. Bill Barnett and Rob Quick of the OSG

Questions?

What is Proteomics? Proteomics as coined by Wilkins and Williams: “The study of proteins as a result from genomics. ” Ruedi Aebersold …defining the mandate of proteomics : “(P)roteomics represents the effort to establish the identities, quantities, structures and biochemical and cellular functions of all proteins in an organism, organ, or organelle, and how these properties vary in space, time and physiological state. ”


Mass Spectrometry Based Proteomics BOTTOM UP PROTEOMICS TOP DOWN PROTEOMICS Long History (mid 20 th century) Developed in early 1990 s High protein identification rates Protein characterization PTM specific proteomics Isoform resolved proteomics Well-established quantitation workflows High throughput methods in development
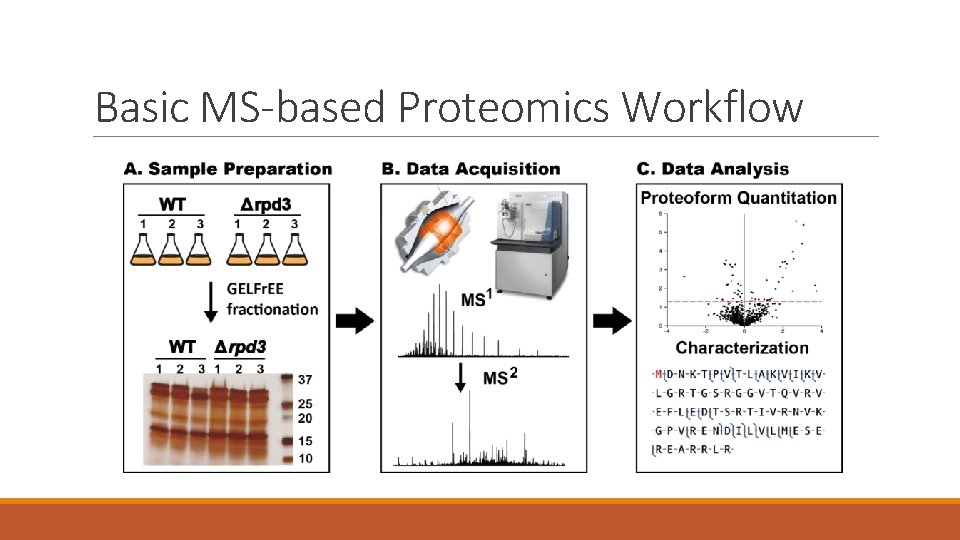
Basic MS-based Proteomics Workflow

Average Project Scope Data Size ◦ ◦ Instrument generates RAW files, which generate roughly 300 MB in a 90 minute run The average number of RAW files per project is 60, roughly 18 GB of data generated in <5 days. Conversion to mz. ML files reduces the size by ~50% Result sets are typically ~10 GB SQLite files Run time ◦ Conversion of a RAW file to mz. ML takes 3 minutes ◦ Searching a single mz. ML file varies according to search complexity (1 -12 hours) ◦ Merging multiple result sets into one takes approximately 2 hours

Additional Connection Details Mounting the NAS ◦ All Kelleher servers mount some or all of the NAS using CIFS User authentication to client portal ◦ Users will use their NU credentials (verified using LDAP) to log into Galaxy Moving data to/from Pulsar ◦ We have setup Pulsar to use a simple RESTful API (HTTP) on custom port Quest HPCC interaction ◦ We are using DRMAA via a Python library to queue jobs
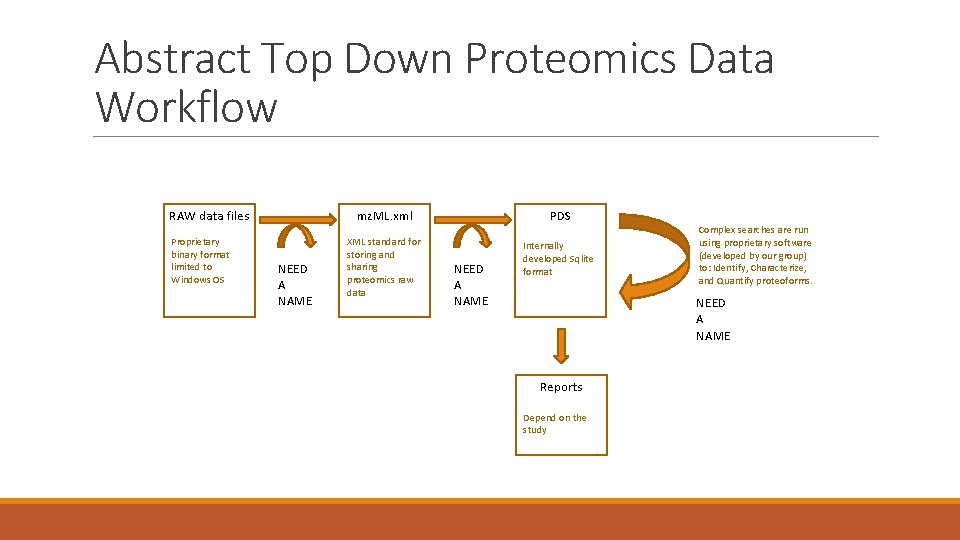
Abstract Top Down Proteomics Data Workflow RAW data files Proprietary binary format limited to Windows OS NEED A NAME mz. ML. xml PDS XML standard for storing and sharing proteomics raw data Internally developed Sqlite format NEED A NAME Complex searches are run using proprietary software (developed by our group) to: Identify, Characterize, and Quantify proteoforms. NEED A NAME Reports Depend on the study
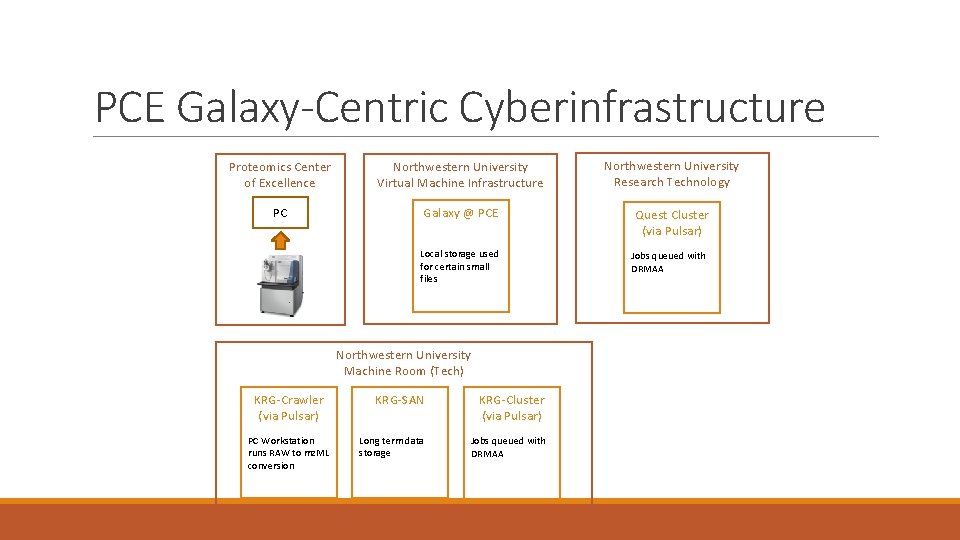
PCE Galaxy-Centric Cyberinfrastructure Proteomics Center of Excellence Northwestern University Virtual Machine Infrastructure Northwestern University Research Technology PC Galaxy @ PCE Quest Cluster (via Pulsar) Local storage used for certain small files Northwestern University Machine Room (Tech) KRG-Crawler (via Pulsar) PC Workstation runs RAW to mz. ML conversion KRG-SAN KRG-Cluster (via Pulsar) Long term data storage Jobs queued with DRMAA
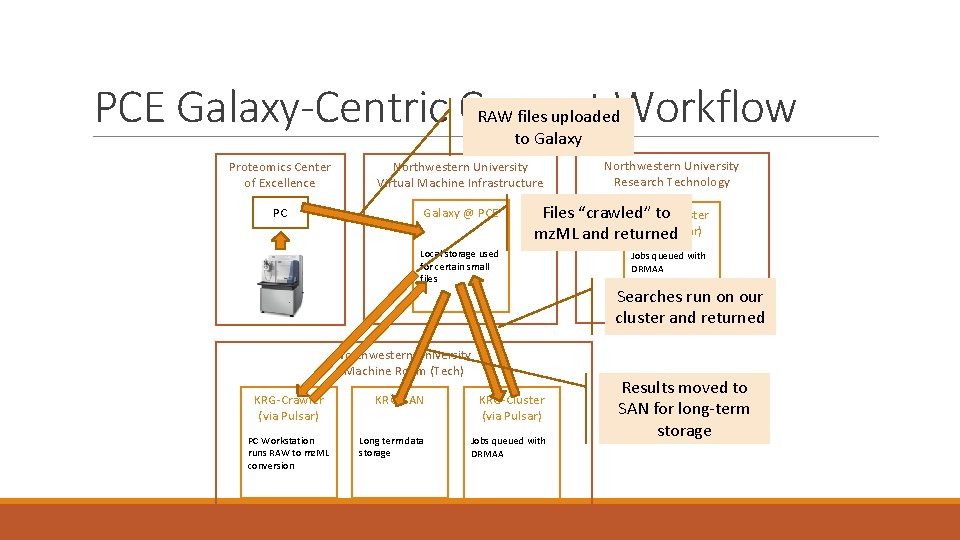
PCE Galaxy-Centric Current Workflow RAW files uploaded to Galaxy Proteomics Center of Excellence Northwestern University Virtual Machine Infrastructure PC Galaxy @ PCE Files “crawled” to. Cluster Quest (via Pulsar) mz. ML and returned Local storage used for certain small files Northwestern University Machine Room (Tech) KRG-Crawler (via Pulsar) PC Workstation runs RAW to mz. ML conversion Northwestern University Research Technology KRG-SAN KRG-Cluster (via Pulsar) Long term data storage Jobs queued with DRMAA Searches run on our cluster and returned Results moved to SAN for long-term storage
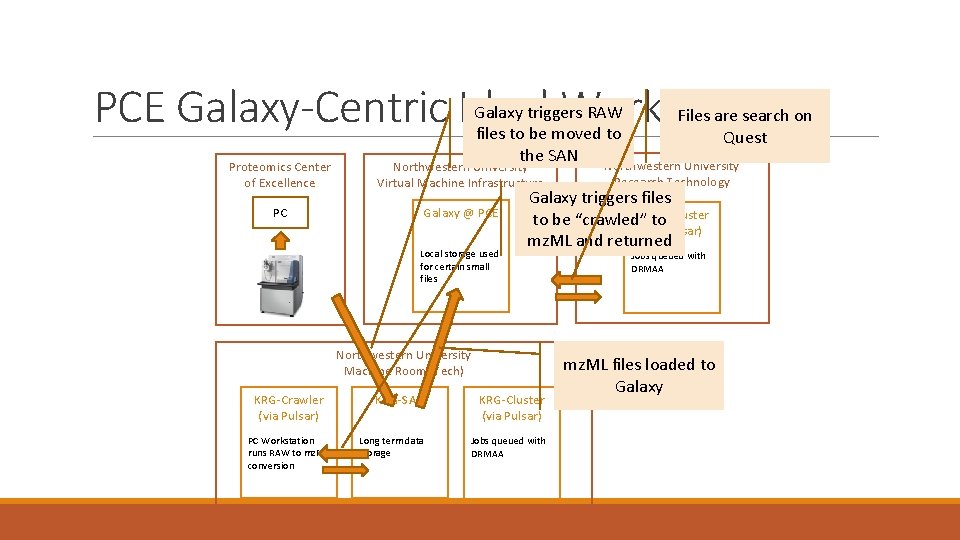
PCE Galaxy-Centric Ideal Workflow Galaxy triggers RAW files to be moved to the SAN Proteomics Center of Excellence Northwestern University Virtual Machine Infrastructure PC Galaxy @ PCE Local storage used for certain small files PC Workstation runs RAW to mz. ML conversion Northwestern University Research Technology Galaxy triggers files Quest to be “crawled” to Cluster (via Pulsar) mz. ML and returned Northwestern University Machine Room (Tech) KRG-Crawler (via Pulsar) Files are search on Quest KRG-SAN KRG-Cluster (via Pulsar) Long term data storage Jobs queued with DRMAA mz. ML files loaded to Galaxy
- Slides: 35