Bayesian style dependent tests Greg Francis PSY 626

Bayesian style dependent tests Greg Francis PSY 626: Bayesian Statistics for Psychological Science Fall 2020 Purdue University

Weapon priming l Anderson, Benjamin, & Bartholow (1998): 32 subjects read an aggression-related word (injure, shatter) out loud as fast as possible w After presentation of a weapon word (shotgun, grenade) w After presentation of a neutral word (rabbit, fish) l l l There were additional trials with non-aggression-related words being read (we ignore them) Our data is the mean response time for wording reading based on whether the first (prime) word is a weapon word or a neutral word Follow along by downloading “Weapon. Prime. csv” and “Weapon. Prime 1. R” from the class web site > WPdata. Weapon <-subset(WPdata, WPdata$Prime=="Weapon") > WPdata. Neutral <-subset(WPdata, WPdata$Prime=="Neutral") > cat("Mean RT for Weapon prime: ", mean(WPdata. Weapon$Time)) Mean RT for Weapon prime: 409. 1875 > cat("Mean RT for Neutral prime: ", mean(WPdata. Neutral$Time)) Mean RT for Neutral prime: 416. 3438 > cat("Correlation between Weapon and Neutral primes: ", cor(subset(WPdata, WPdata$Prime=="Weapon")$Time, subset(WPdata, WPdata$Prime=="Neutral")$Time)) Correlation between Weapon and Neutral primes: 0. 9393478 >

Standard analysis l # Run traditional dependent t-test l traditional 2 <- t. test(Time ~ Prime, data=WPdata, paired=TRUE) l print(traditional 2) Paired t-test data: Time by Prime t = 2. 2239, df = 31, p-value = 0. 03358 alternative hypothesis: true difference in means is not equal to 0 95 percent confidence interval: 0. 593243 13. 719257 sample estimates: mean of the differences 7. 15625

Standard analysis l The dependent t-test is the same as a one-sample t-test of difference scores compared to 0 (just a sign flip) WPdata. Diff<- subset(WPdata, WPdata$Prime=="Weapon") WPdata. Diff$Difference = WPdata. Weapon$Time WPdata. Neutral$Time > t. test(WPdata. Diff$Difference, mu=0) One Sample t-test data: WPdata. Diff$Difference t = -2. 2239, df = 31, p-value = 0. 03358 alternative hypothesis: true mean is not equal to 0 95 percent confidence interval: -13. 719257 -0. 593243 sample estimates: mean of x -7. 15625 >

Bayesian Null model l We can build models using differences l Sample Difference scores from a normal distribution with a mean of 0 l WPmodel 0 b <- map( l alist( Difference ~ dnorm(0, sigma), l sigma ~ dunif(0, 2000) l ), data= WPdata. Diff , control=list(maxit=100000) ) l Quadratic approximate posterior distribution l Formula: l Difference ~ dnorm(0, sigma) l sigma ~ dunif(0, 2000) l Posterior means: l sigma l 19. 29298 l Log-likelihood: -140. 12 l >

Bayesian Alternative model l l We can build models using differences Sample Difference scores from a normal distribution with an unknown WPmodel 1 b <- map( mean l l alist( Difference ~ dnorm(mu, sigma), l mu ~ dnorm(0, 500), l sigma ~ dunif(0, 2000) l ), data= WPdata. Diff , control=list(maxit=100000) ) l Quadratic approximate posterior distribution l Formula: l Difference ~ dnorm(mu, sigma) l mu ~ dnorm(0, 500) l sigma ~ dunif(0, 2000) l Posterior means: mu l sigma l -7. 158999 17. 915740 l Log-likelihood: -137. 75 l >

Model comparison l l l Does the null (WPmodel 0 b, 1 parameter) or alternative (WPmodel 1 b, 2 parameters) model better fit the data? Try: compare(WPmodel 0 b, WPmodel 1 b) Ø Ø WPmodel 1 b Ø WPmodel 0 b WAIC SE d. WAIC d. SE p. WAIC weight 280. 1 7. 67 0. 0 NA 2. 1 0. 79 282. 7 8. 47 2. 7 4. 19 1. 2 0. 21

More direct model l l Oftentimes we do not really care about the difference scores, per sea. We are really interested in the mean Times. The difference scores are just a way of comparing the Times for different conditions. We do not want to ignore that paired scores come from the same subject w A subject fast for a Weapon prime will also be fast for a Neutral prime, just because they are “fast” w We want to factor variability across subjects so that we can focus on variability due to the conditions l We can more directly model such effects

Null model l l Each subject has a unique mean Quadratic approximate posterior distribution No effect of Prime Formula: Time ~ dnorm(mu, sigma) mu <- a[Subject] ~ dnorm(500, # Different baseline 500) (intercept) sigma ~ dunif(0, <-1000) WPmodel 0 map( for each participant alist( Time ~ dnorm(mu, sigma), Posterior means: mua[1] <- a[Subject], a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[13] a[Subject] ~ dnorm(500, 500), 443. 510533 432. 012676 417. 515376 359. 526191 457. 008005 408. 017157 444. 010436 417. 515362 sigma ~ dunif(0, 1000) 433. 512393 379. 522449 380. 522271 385. 021439 429. 013228 ), data= , control=list(maxit=100000) ) a[14] WPdata a[15] a[16] a[17] a[18] a[19] a[20] a[21] a[22] a[23] a[24] a[25] a[26] 370. 524142 338. 030200 438. 011562 519. 496358 298. 537556 339. 529923 388. 520777 407. 017328 536. 993110 433. 512390 433. 512401 437. 011737 408. 517055 a[27] a[28] a[29] a[30] a[31] a[32] sigma 406. 517427 322. 533070 429. 513138 485. 502701 386. 021243 443. 010611 9. 646535 Log-likelihood: -235. 87 >

Different Null model l What happens if we ignore subject dependencies? l Common mean across subjects, no effect of Prime WPmodel 0 c <- map( alist( Time ~ dnorm(mu, sigma), Quadratic approximate posterior distribution mu <- a, a ~ dnorm(500, 500), Formula: sigma ~ dunif(0, 1000) Time ~ dnorm(mu, sigma) ), data= WPdata , control=list(maxit=100000) ) mu <- a a ~ dnorm(500, 500) sigma ~ dunif(0, 1000) Posterior means: a sigma 412. 73955 51. 38347 Log-likelihood: -342. 93 >

Different Null model l l Ignoring dependencies (subject differences) is not a good idea Although WPmodel 0 has many more parameters, they help explain data (increase likelihood) because subjects seem to be different > compare(WPmodel 0, WPmodel 0 c) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 0 567. 6 19. 97 0 NA 45. 7 1 WPmodel 0 c 690. 6 13. 86 123 19. 21 2. 5 0 >

Alternative model Quadratic approximate posterior distribution l Different mean for each subject, and an effect of Prime Formula: Time ~ dnorm(mu, sigma) mu <- a[Subject] + b[Condition. Index] a[Subject] ~ dnorm(500, 500) b[Condition. Index] ~ dnorm(0, 100) # Alternative model 1: different mean for each condition sigma ~ dunif(0, 1000) WPmodel 1 <- map( alist( Time ~ dnorm(mu, sigma), Posterior means: mu <a[Subject] a[1] a[2] a[3] +b[Condition. Index], a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] ~ dnorm(500, 500), a[13] a[Subject] a[14] 476. 45851 464. 96027 450. 46277 392. 47229 b[Condition. Index] ~ dnorm(0, 100), 489. 95701 440. 96465 476. 95842 450. 46251 466. 46051 sigma 412. 46887 ~ dunif(0, 413. 46901 1000) 417. 96831 461. 96131 403. 47002 a[15] a[16]WPdata a[17] , control=list(maxit=100000) a[18] a[19] a[20] a[21] ) a[22] a[23] a[24] a[25] ), data= a[26] a[27] a[28] 370. 97560 470. 95898 552. 44589 331. 48238 372. 47581 421. 46769 439. 96443 569. 94250 466. 46001 466. 46021 469. 95968 441. 46415 439. 46426 355. 47816 a[29] a[30] a[31] a[32] b[1] b[2] sigma 462. 46085 518. 45197 418. 96800 475. 95884 -36. 53240 -29. 37841 8. 95843 Log-likelihood: -231. 14 >

Effect of prime l l Comparing a model with subject effects but no prime effect against a model with subject effects and a prime effect Favors the model with a prime effect > compare(WPmodel 0, WPmodel 1) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 1 558. 9 17. 25 0. 0 NA 45. 9 0. 98 WPmodel 0 567. 2 19. 67 8. 3 10. 41 45. 5 0. 02

Mispecified model > WPmodel 1 Quadratic approximate posterior distribution l WPmodel 1 has too many parameters Quadratic approximate posterior distribution Formula: Time ~ dnorm(mu, l Each mean sigma) is mu_ij = a_i + b_j Formula: mu ~ <-dnorm(mu, a[Subject] + b[Condition. Index] Time sigma) a[Subject] ~ dnorm(500, mu <a[Subject] +ab[Condition. Index] l We have free 500) parameter, because we could have (for b[Condition. Index] ~ dnorm(0, a[Subject] ~ dnorm(500, 500) 100) sigma dunif(0, w ~490 = mu_1 Neutral = 490 +0 and 483 = mu_1 Weapon = 490 - 7 b[Condition. Index] ~1000) dnorm(0, 100) example) sigma ~ dunif(0, 1000) w Ormeans: Posterior a[1] means: a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[13] Posterior w 490 = mu_1 Neutral = 460+30 and 483 = mu_1 Weapon = 460 +23 470. 375627 458. 881464 386. 389222 434. 882178 a[1] a[2] a[3] 444. 383091 a[4] a[5] a[6] 483. 877134 a[7] a[8] a[9]470. 879584 a[10] 444. 382290 a[11] 460. 377737 406. 386409 407. 389108 411. 886637 455. 881657 440. 0725439 428. 5775471 414. 0777735 356. 0888822 453. 5695930 404. 5741834 440. 5725709 l Re-running the model cana[18] lead a[19] to quite different parameters a[14] a[15] a[16] a[17] a[20] a[21] a[22] a[23] a[24] a[25] 414. 0770179 430. 0730368 376. 0815018 377. 0841033 a[26] a[12] a[13] a[14] a[15] a[16] a[17] a[18] a[19] a[20] a[21] a[22] 397. 391936 364. 898002 464. 879588 546. 367923 325. 402337 366. 397446 415. 384745 433. 887751 381. 5787698 425. 5780089 367. 0835175 334. 5895985 434. 5745023 516. 0625932 295. 0948480 563. 863232385. 0840370 460. 378584 460. 379470 463. 877789 435. 382410 336. 0893905 403. 5798503 533. 5617297 a[27] a[24] a[28] a[29] a[30] a[23] a[25] a[26] a[31] a[27] a[32] a[28] b[1] a[29] b[2]a[30]sigmaa[31] a[32] b[1] 433. 387669 349. 397457 456. 383038 512. 368982 412. 887357 469. 879958 -30. 453677 -23. 299174 430. 0764897 430. 0748232 433. 5754550 405. 0756580 403. 0782065 319. 0926067 426. 0774240 8. 958446 382. 5845213 439. 5722461 -0. 1545788 482. 0661102 b[2] sigma Log-likelihood: -231. 14 6. 9990280 8. 9573732 Log-likelihood: -231. 14 >

Another Alternative model > WPmodel 2 Quadratic approximate posterior distribution l You can reduce one parameter by letting the a_i terms handle the Formula: Neutral prime condition Time ~ dnorm(mu, sigma) Time ~ dnorm(mu, mu <- a[Subject] + b sigma) * Is. Weapon l The is 500) then deviation due to Weapon vs. Neutral mu <a[Subject] + bb* Is. Weapon a[Subject] ~variable dnorm(500, 500) b a[Subject] ~ dnorm(0, ~100) b ~ dnorm(0, 100) sigma ~ dunif(0, 1000) Posterior means: a[1] a[2] model a[3] 2: a[4] a[6] from a[7]neutral a[8] a[9] a[10] a[11] # Alternative model a[5] differences a[1]a[13]a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[10] a[11] a[12] WPdata$Is. Weapon<ifelse(WPdata$Prime =="Weapon", 1, a[9] 0) a[12] a[13] 447. 097760 435. 599807 421. 102712 363. 111929 460. 595595 411. 603557 447. 597959 WPmodel 2 <map( 447. 098313437. 099760 435. 600081383. 108577 421. 102161384. 108211 363. 111045388. 607263 460. 596215432. 600994 411. 603547 447. 598266 421. 102102 alist( Time ~ dnorm(mu, sigma), 421. 102175 437. 099904 384. 107880 432. 600454 a[14] a[15] a[16] 383. 107957 a[17] a[18] a[19]388. 607129 a[20] a[21] a[22] a[23] mu <a[Subject] +b*Is. Weapon, a[17] a[18] a[19] a[20] a[21] a[22] a[23] a[24]a[14]a[25]a[15]a[26]a[16] a[Subject] ~ dnorm(500, 500), 523. 085916 302. 121441 343. 115166 392. 107435 a[24] a[25] a[26] 441. 599103 374. 109439 341. 615004 374. 109337 341. 614334 441. 599013437. 100061 523. 086704440. 599217 302. 120377412. 103527 343. 114072 392. 106656 b ~ dnorm(0, 100), 437. 099311 410. 604193 540. 582903 410. 603731 540. 584042 437. 099687 437. 099785 412. 103642 sigma ~a[28] dunif(0, 1000) a[27] a[29] a[30] a[31] a[32]440. 599293 b sigma a[28] a[29] a[30] 489. 091331 a[31] a[32] b 446. 597798 sigma 410. 104181 433. 100464 389. 607587 -7. 179394 ), a[27] data= 326. 117571 WPdata , control=list(maxit=100000) ) 410. 103895 326. 116854 433. 100326 489. 091922 389. 606944 446. 598373 -7. 179240 8. 958166 8. 958942 Log-likelihood: -231. 14
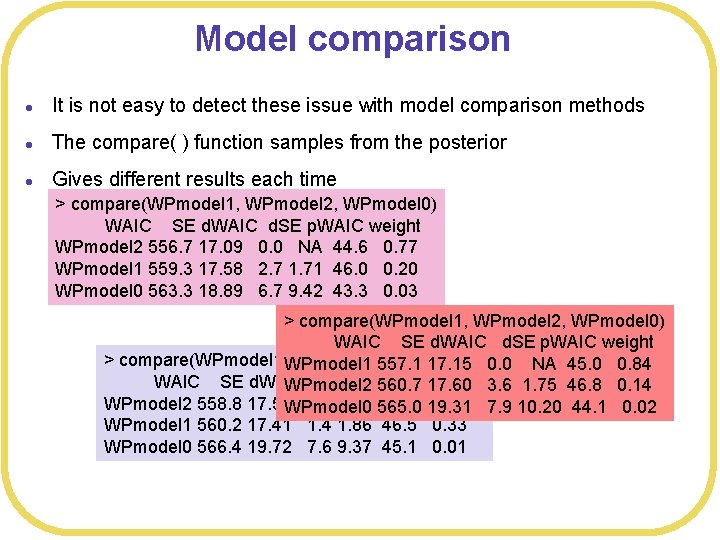
Model comparison l It is not easy to detect these issue with model comparison methods l The compare( ) function samples from the posterior l Gives different results each time > compare(WPmodel 1, WPmodel 2, WPmodel 0) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 2 556. 7 17. 09 0. 0 NA 44. 6 0. 77 WPmodel 1 559. 3 17. 58 2. 7 1. 71 46. 0 0. 20 WPmodel 0 563. 3 18. 89 6. 7 9. 42 43. 3 0. 03 > compare(WPmodel 1, WPmodel 2, WPmodel 0) WAIC SE d. WAIC d. SE p. WAIC weight > compare(WPmodel 1, WPmodel 1 WPmodel 2, WPmodel 0) 557. 1 17. 15 0. 0 NA 45. 0 0. 84 WAIC SE d. WAIC d. SE p. WAIC WPmodel 2 560. 7 weight 17. 60 3. 6 1. 75 46. 8 0. 14 WPmodel 2 558. 8 17. 59 WPmodel 0 0. 0 NA 565. 0 45. 8 19. 31 0. 66 7. 9 10. 20 44. 1 0. 02 WPmodel 1 560. 2 17. 41 1. 4 1. 86 46. 5 0. 33 WPmodel 0 566. 4 19. 72 7. 6 9. 37 45. 1 0. 01
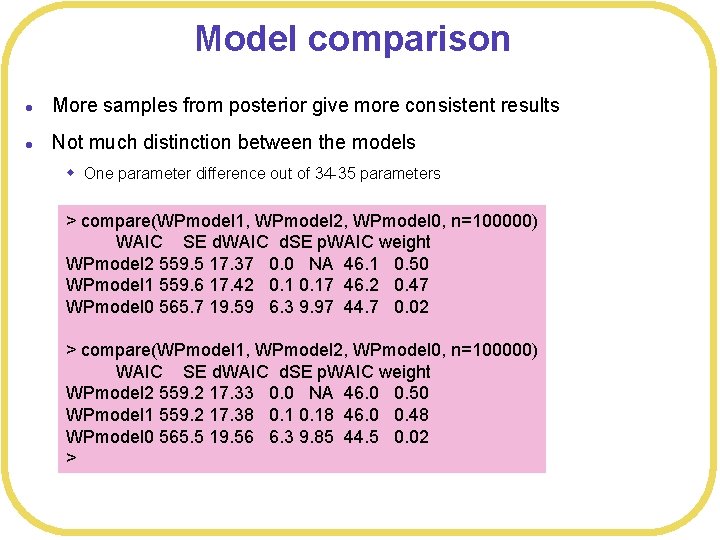
Model comparison l More samples from posterior give more consistent results l Not much distinction between the models w One parameter difference out of 34 -35 parameters > compare(WPmodel 1, WPmodel 2, WPmodel 0, n=100000) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 2 559. 5 17. 37 0. 0 NA 46. 1 0. 50 WPmodel 1 559. 6 17. 42 0. 17 46. 2 0. 47 WPmodel 0 565. 7 19. 59 6. 3 9. 97 44. 7 0. 02 > compare(WPmodel 1, WPmodel 2, WPmodel 0, n=100000) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 2 559. 2 17. 33 0. 0 NA 46. 0 0. 50 WPmodel 1 559. 2 17. 38 0. 18 46. 0 0. 48 WPmodel 0 565. 5 19. 56 6. 3 9. 85 44. 5 0. 02 >
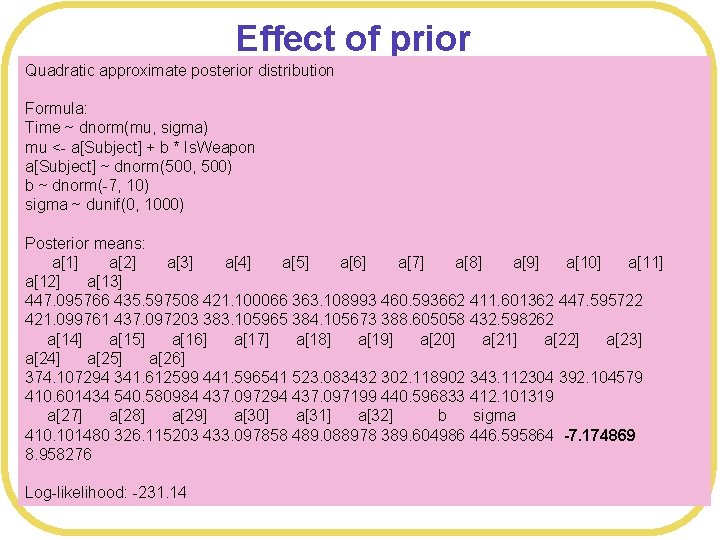
Effect of prior Quadratic approximate posterior distribution WAIC considers not just the number of parameters, but the full posteror Time ~ dnorm(mu, sigma) distribution l Formula: mu <- a[Subject] + b * Is. Weapon a[Subject] dnorm(500, 500) l Thus, ~the prior matters b ~ dnorm(-7, 10) sigma ~ dunif(0, you 1000) l Suppose had additional information that motivated you to put a precise prior on b Posterior means: a[1] a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] # Precise a[12] a[13]prior WPmodel 3 <- map( 421. 100066 363. 108993 460. 593662 411. 601362 447. 595722 447. 095766 435. 597508 alist( Time ~ dnorm(mu, sigma), 384. 105673 388. 605058 432. 598262 421. 099761 437. 097203 383. 105965 a[14] a[15] a[16] a[17] a[18] a[19] a[20] a[21] a[22] a[23] mu <- a[Subject] +b*Is. Weapon, a[24] a[25] ~ dnorm(500, a[26] a[Subject] 500), 374. 107294 341. 612599 b ~ dnorm(-7, 10), 441. 596541 523. 083432 302. 118902 343. 112304 392. 104579 410. 601434 540. 580984 437. 097294 437. 097199 440. 596833 412. 101319 sigma ~ dunif(0, 1000) a[27] a[28] a[29] a[30] a[31] a[32] b sigma ), data= WPdata , control=list(maxit=100000) ) 410. 101480 326. 115203 433. 097858 489. 088978 389. 604986 446. 595864 -7. 174869 8. 958276 Log-likelihood: -231. 14
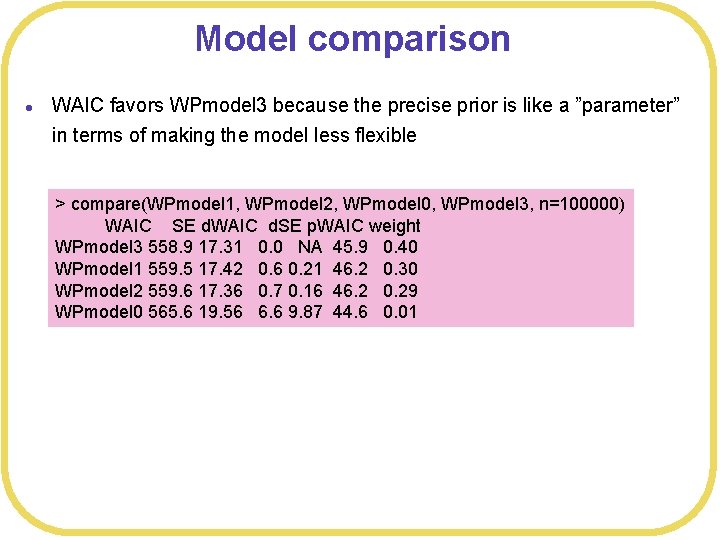
Model comparison l WAIC favors WPmodel 3 because the precise prior is like a ”parameter” in terms of making the model less flexible > compare(WPmodel 1, WPmodel 2, WPmodel 0, WPmodel 3, n=100000) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 3 558. 9 17. 31 0. 0 NA 45. 9 0. 40 WPmodel 1 559. 5 17. 42 0. 6 0. 21 46. 2 0. 30 WPmodel 2 559. 6 17. 36 0. 7 0. 16 46. 2 0. 29 WPmodel 0 565. 6 19. 56 6. 6 9. 87 44. 6 0. 01
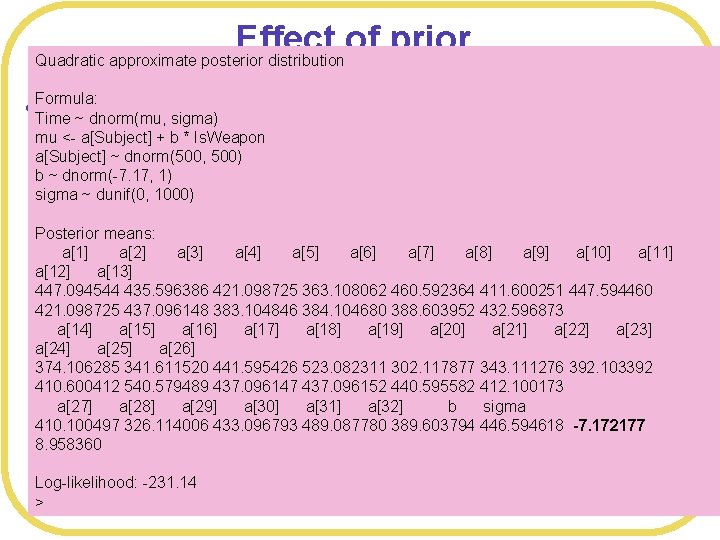
Effect of prior Quadratic approximate posterior distribution l Formula: Even more precise Time ~ dnorm(mu, sigma) mu <- a[Subject] + b * Is. Weapon a[Subject] ~ dnorm(500, 500) b ~ dnorm(-7. 17, 1) sigma ~ dunif(0, 1000) Posterior means: a[1] a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[13] # Precise prior 447. 094544 WPmodel 4435. 596386 <- map( 421. 098725 363. 108062 460. 592364 411. 600251 447. 594460 421. 098725 437. 096148 383. 104846 384. 104680 388. 603952 432. 596873 alist( Time ~ dnorm(mu, sigma), a[14] a[15] a[16] a[17] a[18] a[19] a[20] a[21] a[22] a[23] mu <a[Subject] +b*Is. Weapon, a[24] a[25] a[26] a[Subject] ~ dnorm(500, 500), 523. 082311 302. 117877 343. 111276 392. 103392 374. 106285 341. 611520 441. 595426 b ~ dnorm(-7. 17, 1), 437. 096147 437. 096152 440. 595582 412. 100173 410. 600412 540. 579489 sigma a[27] ~ dunif(0, a[28] 1000) a[29] a[30] a[31] a[32] b sigma 410. 100497 326. 114006 433. 096793 489. 087780 389. 603794 446. 594618 -7. 172177 ), data= WPdata , control=list(maxit=100000) ) 8. 958360 Log-likelihood: -231. 14 >

Model comparison l WAIC favors WPmodel 4 because the precise prior is like a ”parameter” in terms of making the model less flexible compare(WPmodel 0, WPmodel 1, WPmodel 2, WPmodel 3, WPmodel 4, n=100000) WAIC SE d. WAIC d. SE p. WAIC weight WPmodel 4 556. 6058 17. 20791 0. 000000 NA 44. 80485 0. 550424025 WPmodel 3 558. 9973 17. 32539 2. 391490 0. 1915307 45. 89405 0. 166491482 WPmodel 1 559. 3015 17. 36494 2. 695619 0. 2395604 46. 04485 0. 143005004 WPmodel 2 559. 4336 17. 37786 2. 827743 0. 2566478 46. 12307 0. 133863083 WPmodel 0 565. 5728 19. 55089 8. 966994 9. 7955239 44. 56592 0. 006216406
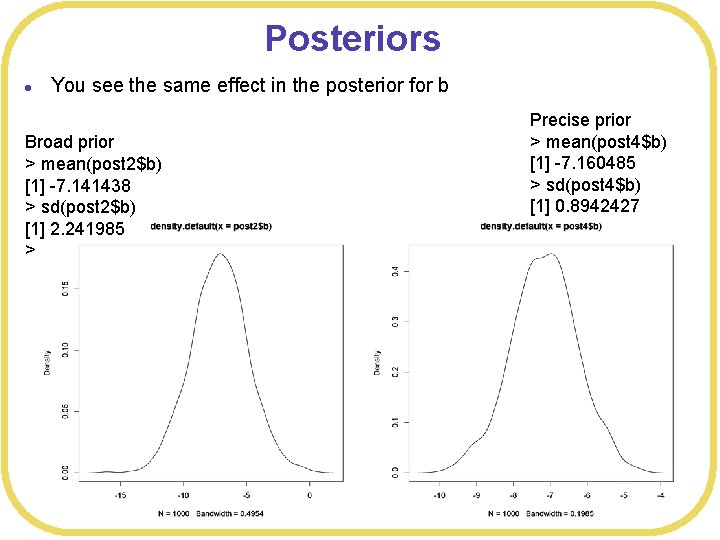
Posteriors l You see the same effect in the posterior for b Broad prior > mean(post 2$b) [1] -7. 141438 > sd(post 2$b) [1] 2. 241985 > Precise prior > mean(post 4$b) [1] -7. 160485 > sd(post 4$b) [1] 0. 8942427

Facial feedback l l Is your emotional state influenced by your facial muscles? If I ask you to smile, you may report feeling happier w But this could just be because you guess I want you to report feeling happier w Or because intentional smiling is associated with feeling happier l You can ask people to use smiling muscles without them realizing it

Facial feedback l Within subjects design (n=21) w Judge “happiness” in a piece of abstract art while w Holding a pen in your teeth (smiling) w Holding a pen in your lips (frowning/pouting) w No pen l l 11 trials for each condition Different art on each trial

Facial feedback l Within subjects design (n=21) w Judge “happiness” in a piece of abstract art while w Holding a pen in your teeth (smiling) w Holding a pen in your lips (frowning/pouting) w No pen l l 11 trials for each condition Different art on each trial

Facial feedback l Within subjects design (n=21) w Judge “happiness” in a piece of abstract art while w Holding a pen in your teeth (smiling) w Holding a pen in your lips (frowning/pouting) w No pen l l 11 trials for each condition Different art on each trial
Data l l l The Happiness. Rating is a number between 0 (no happiness) to 100 (lots of happiness) The facial feedback hypothesis suggests that the mean Happiness. Rating values should be larger when the pen is held in the teeth and lower when the pen is held in the lips File Facial. Feedback. csv contains all the data

Loading data l # load full data file l FFdata<read. csv(file="Facial. Feedback. csv", header=TRUE, strings. As. Factors=FALSE) l # load the rethinking library l library(rethinking) l # Set up dummy variables l FFdata$Pen. In. Teeth <- ifelse(FFdata$Condition =="Pen. In. Teeth", 1, 0) l FFdata$No. Pen <- ifelse(FFdata$Condition =="No. Pen", 1, 0) l FFdata$Pen. In. Lips <- ifelse(FFdata$Condition =="Pen. In. Lips", 1, 0)

Null model l FFmodel 0 <- map( l alist( Happiness. Rating ~ dnorm(mu, sigma), l mu <- a, l Quadratic a ~ dnorm(50, approximate posterior distribution l Formula: 50), sigma ~ dunif(0, 100) Happiness. Rating ~ dnorm(mu, sigma) l mu <- a[Participant] ), data= a[Participant] ~ dnorm(50, 50) sigma ~ dunif(0, 100) FFdata ) Posterior means: a[1] a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[13] a[14] a[15] 47. 73694 41. 46279 38. 26466 53. 52993 51. 99068 39. 83253 61. 76589 53. 58987 50. 72454 39. 04921 59. 41270 48. 94404 35. 36877 53. 04697 48. 31046 a[16] a[17] a[18] a[19] a[20] a[21] sigma 42. 72970 44. 44817 15. 00510 43. 99732 53. 59007 37. 08845 19. 30753 Log-likelihood: -3034. 95 >

Null model w Let’s look at the posterior distributions w post<-extract. samples(FFmodel 0, n= 1000) w > str(post) w List of 2 w $ sigma: num [1: 1000] 19 18. 5 18. 8 18. 5 18. 6. . . w $a : num [1: 1000, 1: 21] 46. 8 46. 6 47. 2 49. 1 45. 1. . . w - attr(*, "source")= chr "quap posterior: 1000 samples from FFmodel 0" w Usually, we do not care about the mean value for each participant w We might care about the mean value across participants
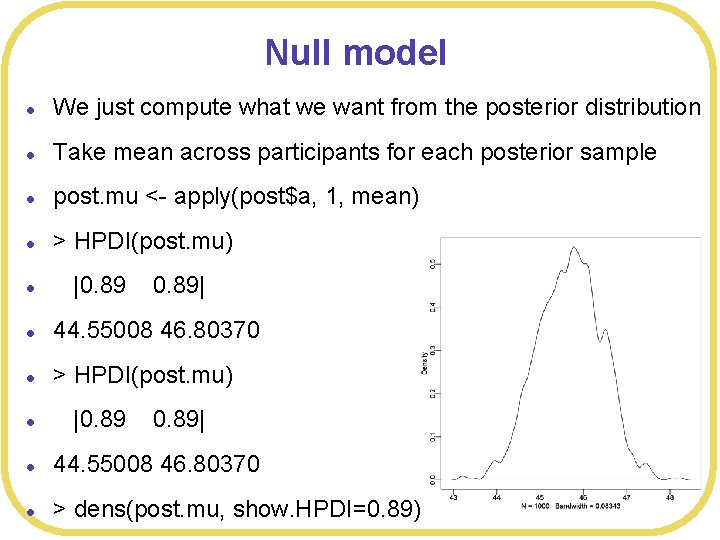
Null model l We just compute what we want from the posterior distribution l Take mean across participants for each posterior sample l post. mu <- apply(post$a, 1, mean) l > HPDI(post. mu) l |0. 89| l 44. 55008 46. 80370 l > dens(post. mu, show. HPDI=0. 89)

Alternative model l Dummy variables set up different values for mu l To avoid a mis-specified model, we let a[Participant] correspond to the “No. Pen” condition l Parameters b 1 and b 2 indicate differences from the No. Pen condition mean l FFmodel 1 <- map( l alist( Happiness. Rating ~ dnorm(mu, sigma), l mu <- a[Participant] + b 1*Pen. In. Teeth + b 2*Pen. In. Lips, l a[Participant] ~ dnorm(50, 50), l b 1 ~ dnorm(0, 20), l b 2 ~ dnorm(0, 20), l sigma ~ dunif(0, 100) l ), data= FFdata , control=list(maxit=100000))
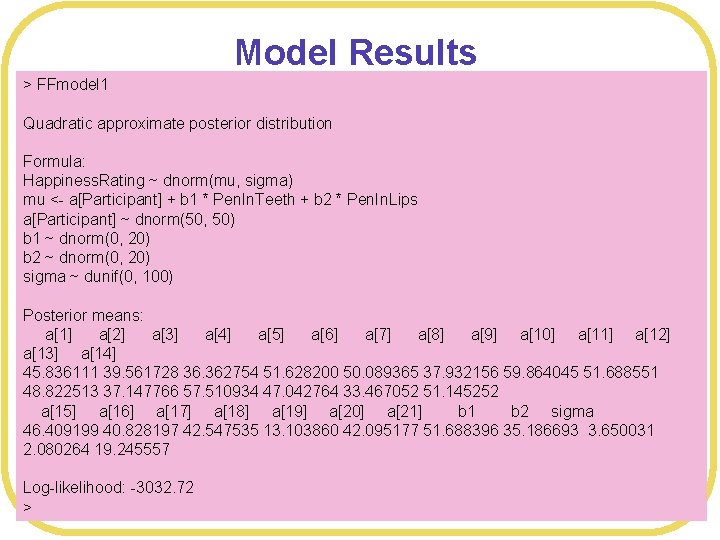
Model Results > FFmodel 1 Quadratic approximate posterior distribution Formula: Happiness. Rating ~ dnorm(mu, sigma) mu <- a[Participant] + b 1 * Pen. In. Teeth + b 2 * Pen. In. Lips a[Participant] ~ dnorm(50, 50) b 1 ~ dnorm(0, 20) b 2 ~ dnorm(0, 20) sigma ~ dunif(0, 100) Posterior means: a[1] a[2] a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[13] a[14] 45. 836111 39. 561728 36. 362754 51. 628200 50. 089365 37. 932156 59. 864045 51. 688551 48. 822513 37. 147766 57. 510934 47. 042764 33. 467052 51. 145252 a[15] a[16] a[17] a[18] a[19] a[20] a[21] b 1 b 2 sigma 46. 409199 40. 828197 42. 547535 13. 103860 42. 095177 51. 688396 35. 186693 3. 650031 2. 080264 19. 245557 Log-likelihood: -3032. 72 >

Posterior l post<-extract. samples(FFmodel 1, n= 1000) > str(post) List of 4 $ b 1 : num [1: 1000] 4. 09 1. 66 3. 4 5. 57 2. 18. . . $ b 2 : num [1: 1000] 3. 116 1. 713 3. 609 4. 366 0. 223. . . $ sigma: num [1: 1000] 19. 1 19 19 19. 2 19. 3. . . $ a : num [1: 1000, 1: 21] 45. 9 47. 2 42. 6 41. 6 52. 7. . . - attr(*, "source")= chr "quap posterior: 1000 samples from FFmodel 1" > l It’s a bit complicated to get the posterior for different means > post. mu. No. Pen <- apply(post$a, 1, mean) > HPDI(post. mu. No. Pen) |0. 89| 41. 68881 45. 69882 > > mean(post. mu. No. Pen) [1] 43. 79379 >
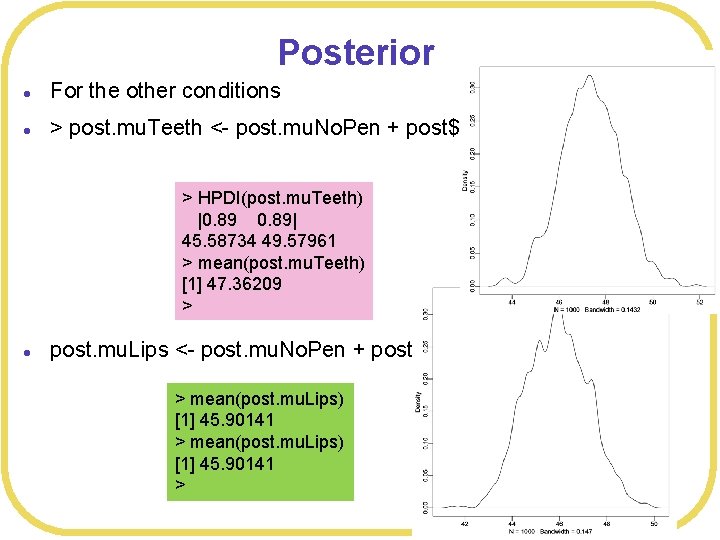
Posterior l For the other conditions l > post. mu. Teeth <- post. mu. No. Pen + post$b 1 > HPDI(post. mu. Teeth) |0. 89| 45. 58734 49. 57961 > mean(post. mu. Teeth) [1] 47. 36209 > l post. mu. Lips <- post. mu. No. Pen + post$b 2 > mean(post. mu. Lips) [1] 45. 90141 >

Posterior l Now, we can ask questions, like, what is the probability that mean happiness ratings are higher with the pen in teeth compared to the pen in the mouth > length(post. mu. Teeth[post. mu. Teeth> post. mu. Lips])/length(post. mu. Teeth) [1] 0. 783 > l What is the probability that the mean happiness ratings are higher with the pen in teeth compared to no pen > length(post. mu. Teeth[post. mu. Teeth> post. mu. No. Pen])/length(post. mu. Teeth) [1] 0. 98 >

Model comparison l Which model is expected to better predict future data? > compare(FFmodel 0, FFmodel 1, n=10000) WAIC SE d. WAIC d. SE p. WAIC weight FFmodel 1 6114. 622 39. 17174 0. 0000000 NA 24. 41115 0. 591148 FFmodel 0 6115. 359 39. 67474 0. 7374264 4. 1745 22. 61288 0. 408852> l Slightly favors alternative model
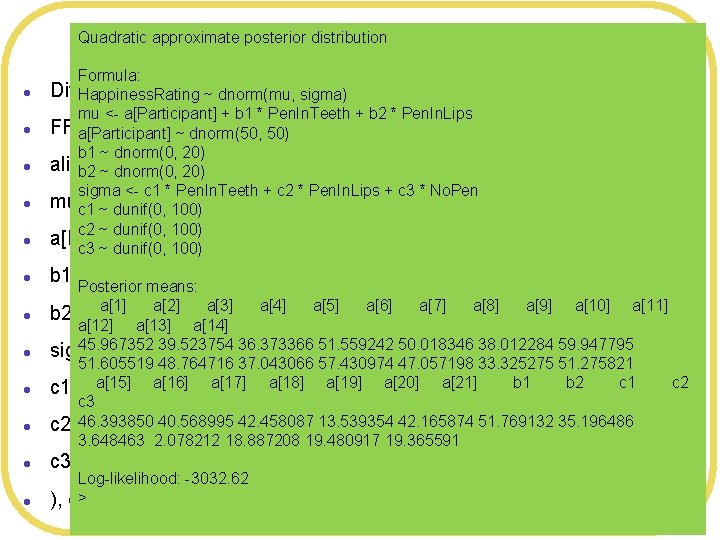
Quadratic approximate posterior distribution More flexible model l l Formula: Different means and sd’s forsigma) each condition Happiness. Rating ~ dnorm(mu, mu <- a[Participant] + b 1 * Pen. In. Teeth + b 2 * Pen. In. Lips FFmodel 2 <- map( a[Participant] ~ dnorm(50, 50) b 1 ~ dnorm(0, 20) alist( ~ dnorm(mu, sigma), b 2 Happiness. Rating ~ dnorm(0, 20) sigma <- c 1 * Pen. In. Teeth + c 2 * Pen. In. Lips + c 3 * No. Pen muc 1 <-~a[Participant] + b 1*Pen. In. Teeth + b 2*Pen. In. Lips, dunif(0, 100) c 2 ~ dunif(0, 100) a[Participant] ~ dnorm(50, 50), c 3 ~ dunif(0, 100) b 1 ~ dnorm(0, 20), Posterior means: a[1] a[2] b 2 ~ dnorm(0, 20), a[3] a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[13] a[14] 45. 967352 39. 523754 36. 373366 51. 559242 50. 018346 38. 012284 59. 947795 sigma <- c 1*Pen. In. Teeth + c 2*Pen. In. Lips + c 3*No. Pen, 51. 605519 48. 764716 37. 043066 57. 430974 47. 057198 33. 325275 51. 275821 a[15] a[16] b 1 b 2 c 1 ~ dunif(0, 100), a[17] a[18] a[19] a[20] a[21] c 3 46. 393850 c 2 ~ dunif(0, 40. 568995 100), 42. 458087 13. 539354 42. 165874 51. 769132 35. 196486 3. 648463 2. 078212 18. 887208 19. 480917 19. 365591 l c 3 ~ dunif(0, 100) l ), Log-likelihood: -3032. 62 > data= FFdata , control=list(maxit=100000) ) c 2

Model comparison l Which model is expected to better predict future data? > compare(FFmodel 0, FFmodel 1, FFmodel 2, n=10000) WAIC SE d. WAIC d. SE p. WAIC weight FFmodel 1 6114. 791 39. 20242 0. 000000 NA 24. 49803 0. 50833173 FFmodel 0 6115. 074 39. 64060 0. 282826 4. 156558 22. 45589 0. 44129839 FFmodel 2 6119. 415 39. 83348 4. 623482 1. 286490 26. 85913 0. 05036988 l l Barely favors alternative model A model with different standard deviations for each condition does not much help with predicting future data

Another more flexible model l We could include trial number Formula: dnorm(mu, sigma) l Happiness. Rating FFmodel 3 <-~map( mu <- a[Participant] + b 1 * Pen. In. Teeth + b 2 * Pen. In. Lips + d * l alist( Trial Happiness. Rating ~ dnorm(mu, sigma), a[Participant] ~ dnorm(50, 50) l b 1 mu <- a[Participant] + b 1*Pen. In. Teeth + b 2*Pen. In. Lips ~ dnorm(0, 20) b 2 ~ dnorm(0, 20) ld ~ a[Participant] dnorm(0, 20) ~ dnorm(50, 50), sigma ~ dunif(0, 100) l + d*Trial, b 1 ~ dnorm(0, 20), Posterior means: b 2 20), a[4] a[5] a[6] a[7] a[8] a[9] a[10] a[11] a[12] a[1]~ dnorm(0, a[2] a[3] a[14] l d ~ dnorm(0, 20), 39. 265013 54. 532068 52. 993409 40. 834000 62. 769072 54. 592484 48. 739225 42. 463387 51. 726237 40. 049640 60. 415650 49. 946193 36. 368751 54. 049611 l sigma dunif(0, a[15] ~ a[16] a[17]100) a[18] a[19] a[20] a[21] b 1 b 2 d sigma 49. 312509 43. 730704 45. 450350 16. 002393 44. 997837 54. 592287 38. 088426 3. 266235 l ), data= FFdata , control=list(maxit=1000000) ) 2. 034827 -0. 163134 19. 186684 l Log-likelihood: -3030. 57 >

Comparing models l Somewhat favors model 3 (taking into account trials) > compare(FFmodel 0, FFmodel 1, FFmodel 2, FFmodel 3, n=10000) WAIC SE d. WAIC d. SE p. WAIC weight FFmodel 3 6112. 7 38. 80 0. 0 NA 25. 6 0. 57 FFmodel 1 6114. 7 39. 23 2. 0 4. 52 24. 5 0. 21 FFmodel 0 6114. 8 39. 60 2. 1 6. 17 22. 3 0. 20 FFmodel 2 6119. 4 39. 83 6. 7 4. 76 26. 8 0. 02 l Does this support the Facial Feedback hypothesis? l Let’s look at the posterior > > post<-extract. samples(FFmodel 3, n= 1000) > str(post) List of 5 $ b 1 : num [1: 1000] -1. 994 1. 343 3. 233 2. 816 0. 937. . . $ b 2 : num [1: 1000] 0. 163 2. 893 3. 684 1. 502 3. 582. . . $ d : num [1: 1000] -0. 141 -0. 225 -0. 17 -0. 241 -0. 14. . . $ sigma: num [1: 1000] 18. 5 18. 8 18. 6 19 19. 1. . . $ a : num [1: 1000, 1: 21] 50. 1 45. 2 47. 3 53 50. . . - attr(*, "source")= chr "quap posterior: 1000 samples from FFmodel 3" >
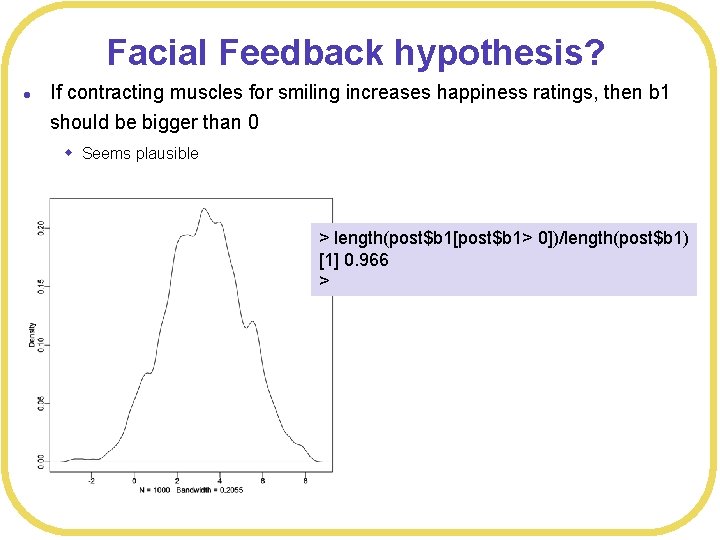
Facial Feedback hypothesis? l If contracting muscles for smiling increases happiness ratings, then b 1 should be bigger than 0 w Seems plausible > length(post$b 1[post$b 1> 0])/length(post$b 1) [1] 0. 966 >
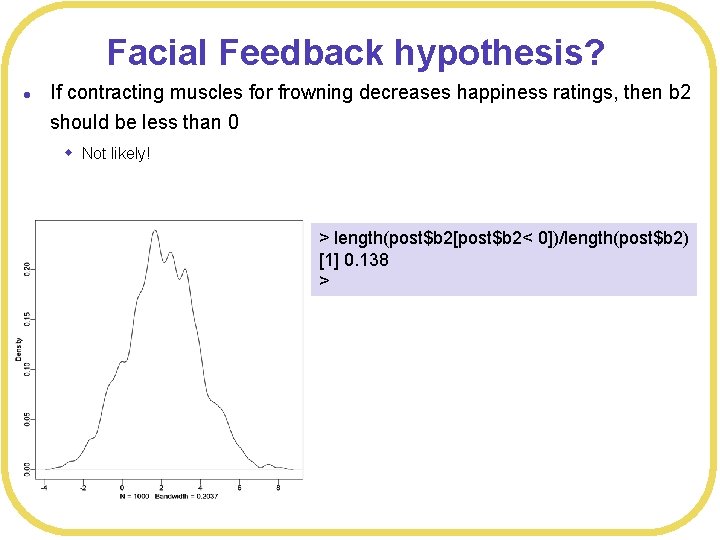
Facial Feedback hypothesis? l If contracting muscles for frowning decreases happiness ratings, then b 2 should be less than 0 w Not likely! > length(post$b 2[post$b 2< 0])/length(post$b 2) [1] 0. 138 >

What does it all mean? l l Relative to the variability in the Happiness. Ratings, the differences in means are pretty small There’s not much difference between the null model and a model with different means w With equal sd, they are expected do about equally well at predicting future data l What do you do? w Gather more data w Develop a better model w Give up w Use what you’ve got

Conclusions l Dependent tests l Pretty straightforward once you get the notation straight w Which is really just the notation of regression l Something similar to contrasts is done by looking at the posteriors w No messy hypothesis testing
- Slides: 45