Bayesian Shrinkage Greg Francis PSY 626 Bayesian Statistics

Bayesian Shrinkage Greg Francis PSY 626: Bayesian Statistics for Psychological Science Fall 2020 Purdue University

Shrinkage l In Lecture 5, we noted that shrinking means toward a fixed value improved model fits for new data w There is an error in the equations there (the numerator should be variance instead of standard deviation) l Shrink to the average w Morris-Efron estimator
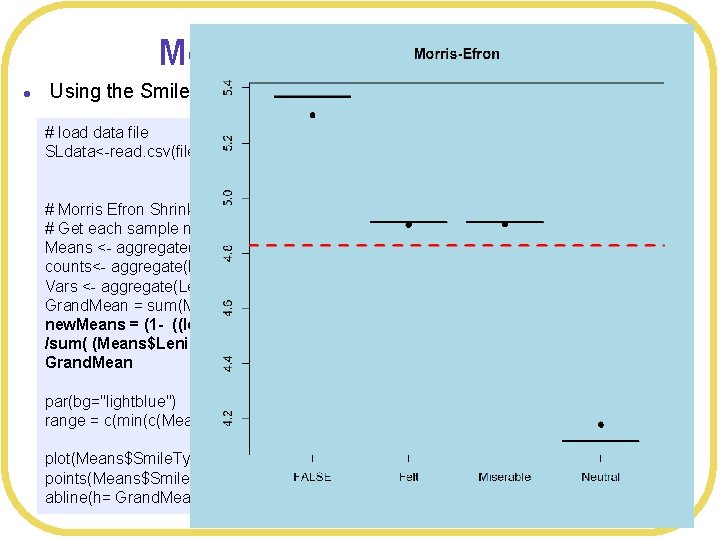
Morris-Efron shrinkage l Using the Smiles and Leniency data set (independent means ANOVA) # load data file SLdata<-read. csv(file="Smiles. Leniency. csv", header=TRUE, strings. As. Factors=TRUE) # Morris Efron Shrinkage # Get each sample mean Means <- aggregate(Leniency~Smile. Type, FUN=mean, data=SLdata) counts<- aggregate(Leniency~Smile. Type, FUN=length, data=SLdata) Vars <- aggregate(Leniency~Smile. Type, FUN=var, data=SLdata) Grand. Mean = sum(Means$Leniency)/4 new. Means = (1 - ((length(Means$Leniency)-3))*Vars$Leniency/counts$Leniency /sum( (Means$Leniency - Grand. Mean)^2 )) *(Means$Leniency - Grand. Mean) + Grand. Mean par(bg="lightblue") range = c(min(c(Means$Leniency, new. Means)), max(c(Means$Leniency, new. Means))) plot(Means$Smile. Type, Means$Leniency, main="Morris-Efron", ylim=range) points(Means$Smile. Type, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)

Bayesian ANOVA l l Using the Smiles and Leniency data set (independent means ANOVA) We used linear regression to produce the Bayesian equivalent of an ANOVA Quadratic approximate posterior distribution Formula: Leniency ~ dnorm(mu, sigma) mu <- a[Condition. Index] ~ dnorm(5, 5) sigma ~ dunif(0, 10) Posterior means: a[1] a[2] a[3] a[4] sigma 5. 366508 4. 912020 4. 912080 4. 120311 1. 603446 Log-likelihood: -257. 19 >
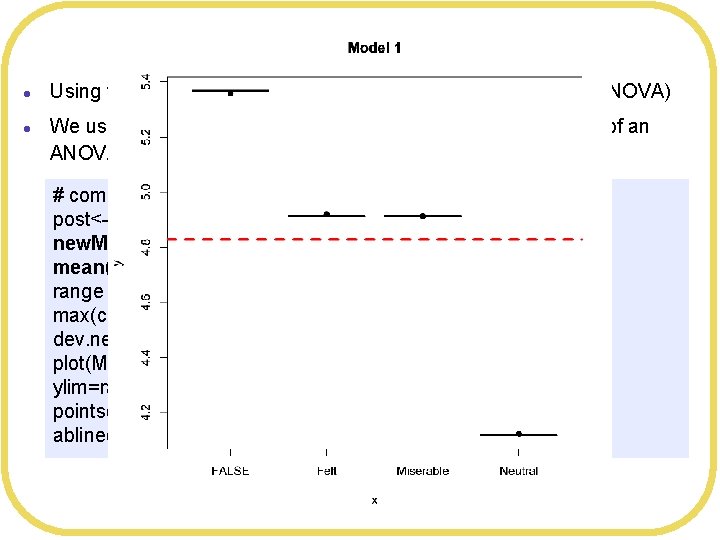
Bayesian ANOVA l l Using the Smiles and Leniency data set (independent means ANOVA) We used linear regression to produce the Bayesian equivalent of an ANOVA (a little hard to interpret) # compute means of posteriors post<-extract. samples(SLmodel 3, n= 2000) new. Means <- c(mean(post$a[, 1]), mean(post$a[, 2]), mean(post$a[, 3]), mean(post$a[, 4] ) ) range = c(min(c(Means$Leniency, new. Means)), max(c(Means$Leniency, new. Means))) dev. new() plot(Means$Smile. Type, Means$Leniency, main="Model 1", ylim=range) points(Means$Smile. Type, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)

Bayesian Shrinkage l We get something very much like shrinkage by using a prior l Let’s set up a prior to pull values toward the grand mean # Set prior on means to shrink toward Grand. Mean SLmodel 4 <- map( alist( Leniency ~ dnorm(mu, sigma), Quadratic approximate posterior distribution mu <- a[Condition. Index], Formula: a[Condition. Index] ~ dnorm(4. 827206, 5), Leniency ~ dnorm(mu, sigma) sigma ~ dunif(0, 10) mu <- a[Condition. Index] ~ dnorm(4. 827206, 5) ), data= SLdata ) sigma ~ dunif(0, 10) Posterior means: a[1] a[2] a[3] a[4] sigma 5. 366045 4. 911563 4. 911468 4. 119826 1. 603436 Log-likelihood: -257. 19 >
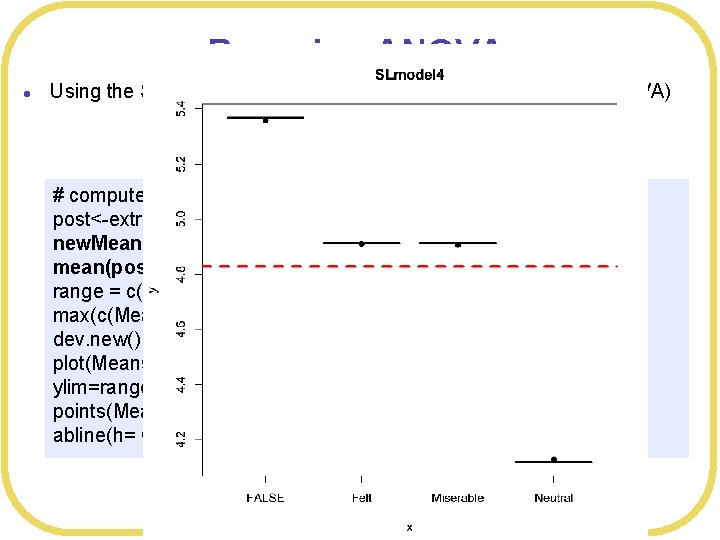
Bayesian ANOVA l Using the Smiles and Leniency data set (independent means ANOVA) # compute means of posteriors post<-extract. samples(SLmodel 4, n= 2000) new. Means <- c(mean(post$a[, 1]), mean(post$a[, 2]), mean(post$a[, 3]), mean(post$a[, 4] ) ) range = c(min(c(Means$Leniency, new. Means)), max(c(Means$Leniency, new. Means))) dev. new() plot(Means$Smile. Type, Means$Leniency, main="SLmodel 4", ylim=range) points(Means$Smile. Type, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)

Model comparison l Shrinkage should lead to better prediction > compare(SLmodel 3, SLmodel 4) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 3 524. 8 15. 08 0. 0 NA 5. 1 0. 52 SLmodel 4 524. 9 15. 04 0. 15 5. 1 0. 48 l l Hardly any effect. Big standard deviation of prior. Shrinkage never really hurts for prediction, is that true of the effect of priors?
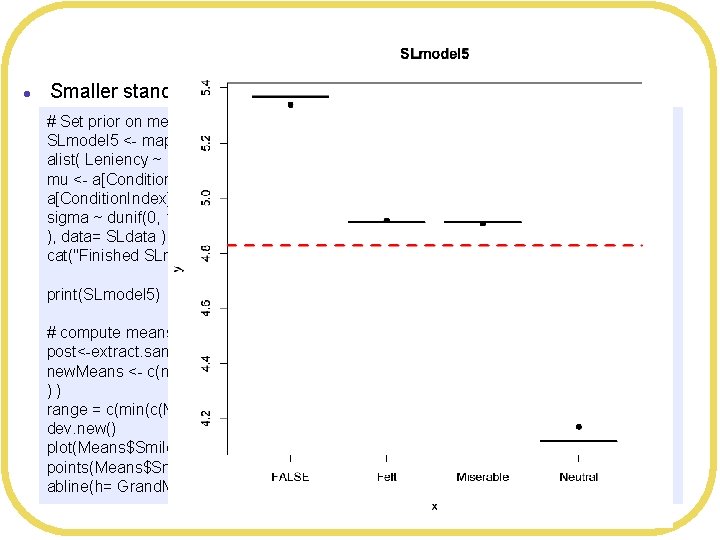
Bayesian Shrinkage l Smaller standard deviation # Set prior on means to shrink toward Grand. Mean, with smaller standard deviation SLmodel 5 <- map( alist( Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], a[Condition. Index] ~ dnorm(4. 827206, 1), sigma ~ dunif(0, 10) ), data= SLdata ) cat("Finished SLmodel 5n") print(SLmodel 5) # compute means of posteriors post<-extract. samples(SLmodel 5, n= 2000) new. Means <- c(mean(post$a[, 1]), mean(post$a[, 2]), mean(post$a[, 3]), mean(post$a[, 4] ) ) range = c(min(c(Means$Leniency, new. Means)), max(c(Means$Leniency, new. Means))) dev. new() plot(Means$Smile. Type, Means$Leniency, main="SLmodel 5", ylim=range) points(Means$Smile. Type, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)
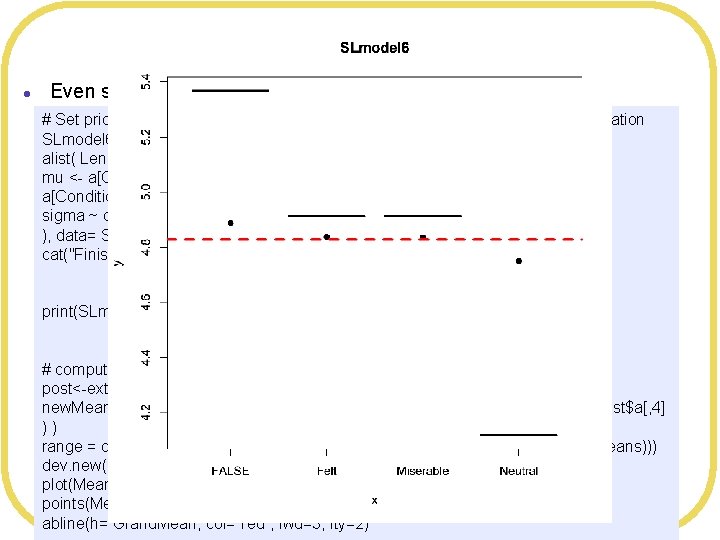
Bayesian Shrinkage l Even smaller standard deviation # Set prior on means to shrink toward Grand. Mean, with very small standard deviation SLmodel 6 <- map( alist( Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], a[Condition. Index] ~ dnorm(4. 827206, 0. 1), sigma ~ dunif(0, 10) ), data= SLdata ) cat("Finished SLmodel 6n") print(SLmodel 6) # compute means of posteriors post<-extract. samples(SLmodel 6, n= 2000) new. Means <- c(mean(post$a[, 1]), mean(post$a[, 2]), mean(post$a[, 3]), mean(post$a[, 4] ) ) range = c(min(c(Means$Leniency, new. Means)), max(c(Means$Leniency, new. Means))) dev. new() plot(Means$Smile. Type, Means$Leniency, main="SLmodel 6", ylim=range) points(Means$Smile. Type, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)

Model comparison l l Contrary to shrinkage, the prior can hurt if it is too constraining Makes intuitive sense > compare(SLmodel 3, SLmodel 4, SLmodel 5, SLmodel 6) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 5 523. 9 15. 14 0. 0 NA 4. 6 0. 38 SLmodel 3 524. 9 15. 06 1. 0 0. 45 5. 1 0. 23 SLmodel 4 525. 0 15. 26 1. 1 0. 54 5. 2 0. 22 SLmodel 6 525. 4 14. 73 1. 5 5. 52 1. 3 0. 18 > l l l A standard deviation of 1 (model 5) helps, but a standard deviation of 0. 1 (model 6) hurts A standard deviation of 5 (model 4) helps some Model 3 shrinks to the middle of the scale, 5, rather than to the grand mean

Ad hoc approach l l Going too big for the standard deviation is not going to hurt (but it might not help as much as it could) Grand. SE = sqrt(var(Means$Leniency) ) # SD of means # Set prior on means to shrink toward Grand. Mean, with prior standard deviation estimated from data Try using a standard deviation estimated from the data itself SLmodel 7 <- map( w Standard error of the mean (0. 2791214) alist( Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], a[Condition. Index] ~ dnorm(4. 827206, 0. 5195674), sigma ~ dunif(0, 10) ), data= SLdata ) cat("Finished SLmodel 7n") print(SLmodel 7) # compute means of posteriors post<-extract. samples(SLmodel 7, n= 2000) new. Means <- c(mean(post$a[, 1]), mean(post$a[, 2]), mean(post$a[, 3]), mean(post$a[, 4] ) ) range = c(min(c(Means$Leniency, new. Means)), max(c(Means$Leniency, new. Means))) dev. new() plot(Means$Smile. Type, Means$Leniency, main="SLmodel 7", ylim=range) points(Means$Smile. Type, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)

Model comparison l Does a decent job here compared to the “no shrinkage” model > compare(SLmodel 3, SLmodel 7) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 7 523. 7 14. 94 0. 0 NA 4. 2 0. 66 SLmodel 3 525. 1 15. 23 1. 4 1. 61 5. 2 0. 34 > l Does a decent job compared to other models > compare(SLmodel 3, SLmodel 4, SLmodel 5, SLmodel 6, SLmodel 7) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 7 523. 6 14. 92 0. 0 NA 4. 2 0. 31 SLmodel 5 524. 4 15. 22 0. 8 1. 13 4. 8 0. 21 SLmodel 4 524. 7 15. 16 1. 1 1. 51 5. 0 0. 18 SLmodel 3 524. 7 15. 16 1. 1 1. 64 5. 0 0. 17 SLmodel 6 525. 3 14. 64 1. 7 4. 32 1. 2 0. 13 l No guarantees

Hierarchical model l l Set up hyper-parameters that indicate grand mean and standard deviation Use broad priors for hyper-parameters because the data should constrain them # Heirarchical model SLmodel 8 <- map 2 stan( alist( Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], a[Condition. Index] ~ dnorm(Gmean, Gsd), Gmean ~ dnorm(5, 5), Gsd ~ dunif(0, 10), sigma ~ dunif(0, 10) ), data= SLdata )
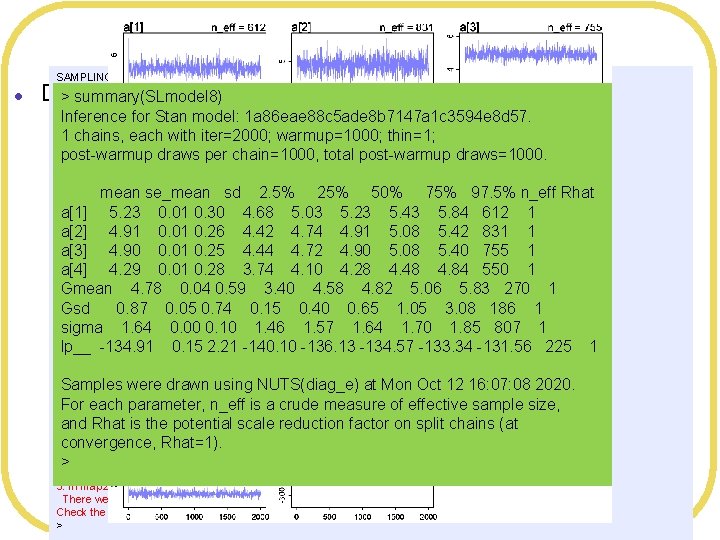
Problems l SAMPLING FOR MODEL '1 a 86 eae 88 c 5 ade 8 b 7147 a 1 c 3594 e 8 d 57' NOW (CHAIN 1). Chain 1: > summary(SLmodel 8) Chain 1: Gradient evaluation took 2. 9 e-05 seconds Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0. 29 seconds. Inference for Stan model: 1 a 86 eae 88 c 5 ade 8 b 7147 a 1 c 3594 e 8 d 57. Chain 1: Adjust your expectations accordingly! 1 chains, each with iter=2000; warmup=1000; thin=1; Chain 1: post-warmup draws per chain=1000, total post-warmup draws=1000. Chain 1: Iteration: 1 / 2000 [ 0%] (Warmup) Chain 1: Iteration: 200 / 2000 [ 10%] (Warmup) mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat Chain 1: Iteration: 400 / 2000 [ 20%] (Warmup) Chain 1: Iteration: 600 / 2000 [ 30%] (Warmup) a[1] 5. 23 0. 01 0. 30 4. 68 5. 03 5. 23 5. 43 5. 84 612 1 Chain 1: Iteration: 800 / 2000 [ 40%] (Warmup) a[2] 4. 91 0. 01 0. 26 4. 42 4. 74 4. 91 5. 08 5. 42 831 1 Chain 1: Iteration: 1000 / 2000 [ 50%] (Warmup) Chain 1: Iteration: 1001 / 2000 [ 50%] (Sampling) a[3] 4. 90 0. 01 0. 25 4. 44 4. 72 4. 90 5. 08 5. 40 755 1 Chain 1: Iteration: 1200 / 2000 [ 60%] (Sampling) a[4] 4. 29 0. 01 0. 28 3. 74 4. 10 4. 28 4. 48 4. 84 550 1 Chain 1: Iteration: 1400 / 2000 [ 70%] (Sampling) Chain 1: Iteration: 1600 / 2000 [ 80%] (Sampling) Gmean 4. 78 0. 04 0. 59 3. 40 4. 58 4. 82 5. 06 5. 83 270 1 Chain 1: Iteration: 1800 / 2000 [ 90%] (Sampling) Gsd 0. 87 0. 05 0. 74 0. 15 0. 40 0. 65 1. 05 3. 08 186 1 Chain 1: Iteration: 2000 / 2000 [100%] (Sampling) Chain 1: sigma 1. 64 0. 00 0. 10 1. 46 1. 57 1. 64 1. 70 1. 85 807 1 Chain 1: Elapsed Time: 0. 169378 seconds (Warm-up) lp__ -134. 91 0. 15 2. 21 -140. 10 -136. 13 -134. 57 -133. 34 -131. 56 225 1 Chain 1: 0. 183306 seconds (Sampling) Chain 1: 0. 352684 seconds (Total) Chain 1: Samples were drawn using NUTS(diag_e) at Mon Oct 12 16: 07: 08 2020. Computing WAIC For each parameter, n_eff is a crude measure of effective sample size, Warning messages: 1: There were 5 divergent transitions after warmup. See and Rhat is the potential scale reduction factor on split chains (at http: //mc-stan. org/misc/warnings. html#divergent-transitions-after-warmup convergence, Rhat=1). to find out why this is a problem and how to eliminate them. 2: Examine the pairs() plot to diagnose sampling problems > 3: In map 2 stan(alist(Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], : There were 5 divergent iterations during sampling. Check the chains (trace plots, n_eff, Rhat) carefully to ensure they are valid. > Divergence
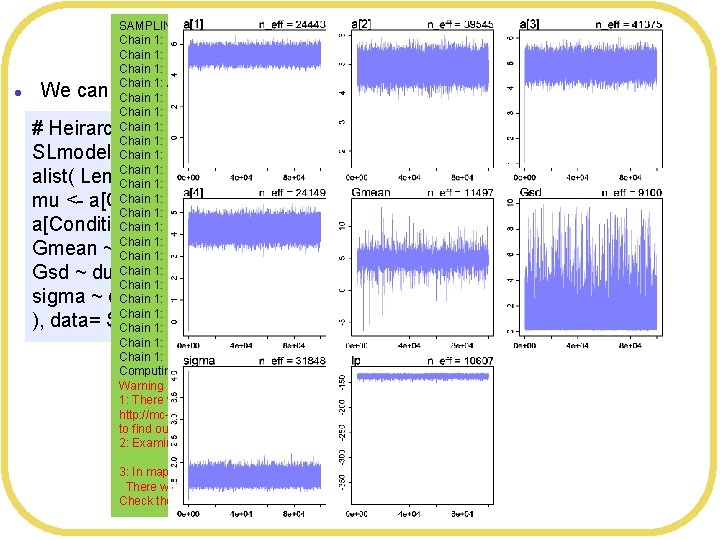
SAMPLING FOR MODEL '9 bf 75 bab 9 bbfb 0826 ce 921608414 ac 4 a' NOW (CHAIN 1). Chain 1: Gradient evaluation took 3. 1 e-05 seconds Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0. 31 seconds. Chain 1: Adjust your expectations accordingly! Chain 1: Iteration: 1 / 100000 [ 0%] (Warmup) Chain 1: Iteration: 10000 / 100000 [ 10%] (Warmup) Chain 1: Iteration: 20000 / 100000 [ 20%] (Warmup) Chain 1: Iteration: 30000 / 100000 [ 30%] (Warmup) Chain 1: Iteration: 40000 / 100000 [ 40%] (Warmup) Chain 1: Iteration: 50000 / 100000 [ 50%] (Warmup) Chain 1: Iteration: 50001 / 100000 [ 50%] (Sampling) Chain 1: Iteration: 60000 / 100000 [ 60%] (Sampling) Chain 1: Iteration: 70000 / 100000 [ 70%] (Sampling) Chain 1: Iteration: 80000 / 100000 [ 80%] (Sampling) Chain 1: Iteration: 90000 / 100000 [ 90%] (Sampling) Chain 1: Iteration: 100000 / 100000 [100%] (Sampling) Chain 1: Elapsed Time: 7. 26029 seconds (Warm-up) Chain 1: 9. 13692 seconds (Sampling) Chain 1: 16. 3972 seconds (Total) Chain 1: Computing WAIC Warning messages: 1: There were 227 divergent transitions after warmup. See http: //mc-stan. org/misc/warnings. html#divergent-transitions-after-warmup to find out why this is a problem and how to eliminate them. 2: Examine the pairs() plot to diagnose sampling problems 3: In map 2 stan(alist(Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], : There were 227 divergent iterations during sampling. Check the chains (trace plots, n_eff, Rhat) carefully to ensure they are valid. Iterations and warm up l We can modify Stan’s algorithm to try to avoid divergent transitions # Heirarchical model SLmodel 8 <- map 2 stan( alist( Leniency ~ dnorm(mu, sigma), mu <- a[Condition. Index], a[Condition. Index] ~ dnorm(Gmean, Gsd), Gmean ~ dnorm(5, 5), Gsd ~ dunif(0, 10), sigma ~ dunif(0, 10) ), data= SLdata, iter=1 e 4, warmup=2000 )

Convergence l Other statistics indicate the model is not bad > summary(SLmodel 8) Inference for Stan model: 9 bf 75 bab 9 bbfb 0826 ce 921608414 ac 4 a. 1 chains, each with iter=1 e+05; warmup=50000; thin=1; post-warmup draws per chain=50000, total post-warmup draws=50000. mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] 5. 25 0. 00 0. 28 4. 71 5. 05 5. 25 5. 44 5. 81 24443 1 a[2] 4. 89 0. 00 0. 26 4. 39 4. 71 4. 89 5. 06 5. 41 39545 1 a[3] 4. 89 0. 00 0. 26 4. 39 4. 72 4. 89 5. 06 5. 41 41375 1 a[4] 4. 28 0. 00 0. 29 3. 70 4. 08 4. 28 4. 48 4. 84 24149 1 Gmean 4. 83 0. 01 0. 61 3. 65 4. 59 4. 82 5. 06 6. 00 11497 1 Gsd 0. 90 0. 01 0. 89 0. 14 0. 41 0. 65 1. 05 3. 27 9100 1 sigma 1. 65 0. 00 0. 10 1. 46 1. 57 1. 64 1. 71 1. 86 31848 1 lp__ -134. 99 0. 02 2. 32 -140. 58 -136. 24 -134. 60 -133. 31 -131. 63 10607 1 Samples were drawn using NUTS(diag_e) at Mon Oct 12 16: 17: 55 2020. For each parameter, n_eff is a crude measure of effective sample size, and Rhat is the potential scale reduction factor on split chains (at convergence, Rhat=1). >

Convergence l I ran the “default” settings again and got no divergent transitions > summary(SLmodel 8) Inference for Stan model: c 31 ee 3886 b 9 d 796 ff 1 f 7 b 3 f 283 cf 351 c. 1 chains, each with iter=2000; warmup=1000; thin=1; post-warmup draws per chain=1000, total post-warmup draws=1000. mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] 5. 26 0. 01 0. 28 4. 70 5. 07 5. 26 5. 44 5. 81 498 1 a[2] 4. 91 0. 01 0. 25 4. 42 4. 75 4. 90 5. 07 5. 39 705 1 a[3] 4. 90 0. 01 0. 26 4. 40 4. 74 4. 89 5. 06 5. 41 641 1 a[4] 4. 26 0. 01 0. 30 3. 68 4. 05 4. 27 4. 49 4. 83 666 1 Gmean 4. 82 0. 02 0. 45 3. 88 4. 59 4. 81 5. 05 5. 78 392 1 Gsd 0. 83 0. 03 0. 67 0. 16 0. 41 0. 64 1. 00 2. 64 369 1 sigma 1. 64 0. 00 0. 10 1. 46 1. 57 1. 64 1. 71 1. 86 666 1 lp__ -134. 90 0. 12 2. 09 -139. 54 -136. 14 -134. 67 -133. 35 -131. 69 285 1 Samples were drawn using NUTS(diag_e) at Mon Oct 12 16: 24: 03 2020. For each parameter, n_eff is a crude measure of effective sample size, and Rhat is the potential scale reduction factor on split chains (at convergence, Rhat=1).
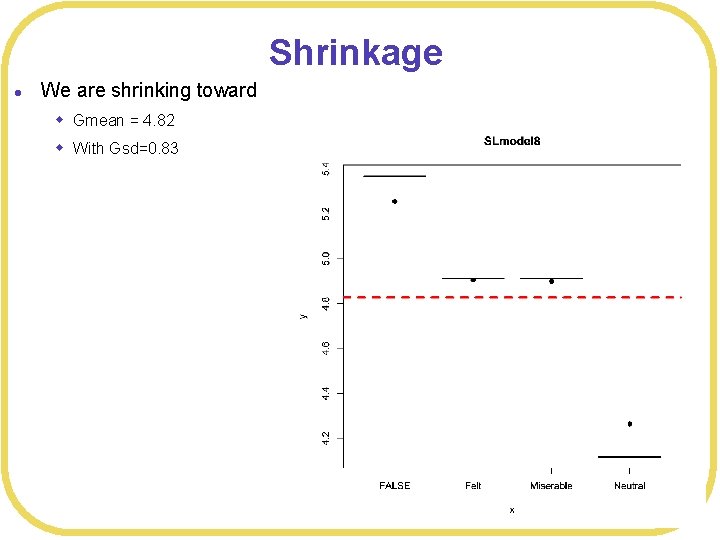
Shrinkage l We are shrinking toward w Gmean = 4. 82 w With Gsd=0. 83

Shrinkage l We are shrinking toward w Gmean = 4. 82 w With Gsd=0. 83 > compare(SLmodel 3, SLmodel 4, SLmodel 5, SLmodel 6, SLmodel 7, SLmodel 8) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 7 523. 3 14. 97 0. 0 NA 4. 1 0. 27 SLmodel 8 524. 2 14. 27 0. 9 0. 76 4. 4 0. 18 SLmodel 5 524. 2 15. 10 0. 9 1. 14 4. 8 0. 17 SLmodel 3 524. 6 15. 21 1. 3 1. 63 5. 0 0. 14 SLmodel 4 524. 7 15. 12 1. 3 1. 64 5. 0 0. 14 SLmodel 6 525. 3 14. 51 1. 9 4. 39 1. 2 0. 10 Warning message: In compare(SLmodel 3, SLmodel 4, SLmodel 5, SLmodel 6, SLmodel 7, SLmodel 8) : Not all model fits of same class. This is usually a bad idea, because it implies they were fit by different algorithms. Check yourself, before you wreck yourself. >

Shrinkage l Bayesian models naturally do something like shrinkage through the prior l It’s not true shrinkage, but it has similar effects l If you have informed priors, you should use them l You could explicitly set the prior based on properties of the data, but there’s no guarantee that is going to help w And it risks overfitting your data l Hierarchical modeling is a better approach w Tends to work about as well w Priors do not matter very much
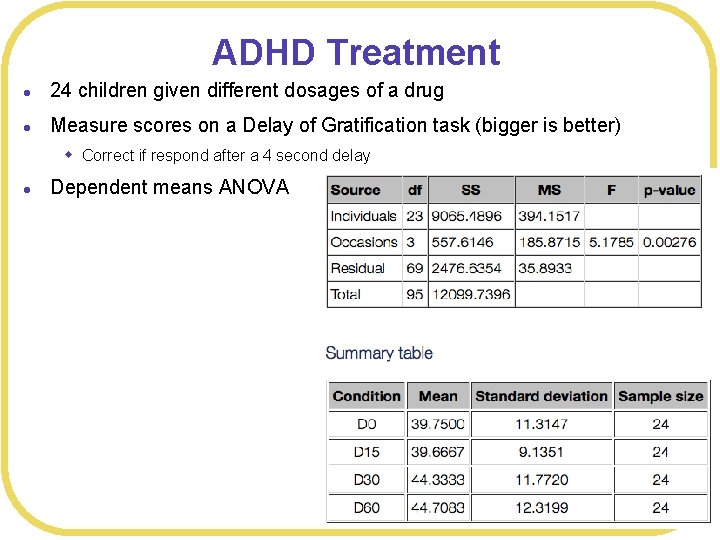
ADHD Treatment l 24 children given different dosages of a drug l Measure scores on a Delay of Gratification task (bigger is better) w Correct if respond after a 4 second delay l Dependent means ANOVA

Shrinkage for subjects l Our data has quite some variability across subjects # load data file ATdata<-read. csv(file="ADHDTreatment. csv", header=TRUE, strings. As. Factors=TRUE) # Set up dummy variables ATdata$Dosage 0 <- ifelse(ATdata$Dosage =="D 0", 1, 0) ATdata$Dosage 15 <- ifelse(ATdata$Dosage =="D 15", 1, 0) ATdata$Dosage 30 <- ifelse(ATdata$Dosage =="D 30", 1, 0) ATdata$Dosage 60 <- ifelse(ATdata$Dosage =="D 60", 1, 0) # Make unique index for each participant ATdata$Participant <- coerce_index(ATdata$Subject. ID) plot(ATdata$Dosage. Number, ATdata$Correct. Responses) for(i in c(1: 24)) { this. Set<- subset(ATdata, ATdata$Participant == i) lines(this. Set$Dosage. Number, this. Set$Correct. Responses, col=i, lty=i) }
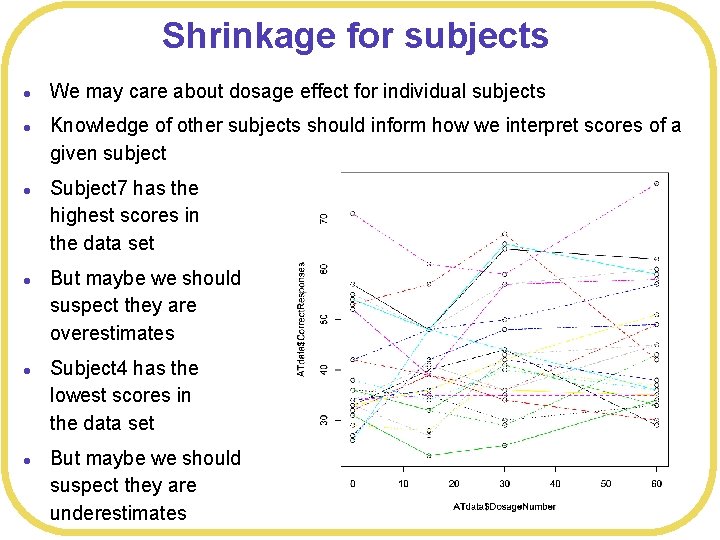
Shrinkage for subjects l l l We may care about dosage effect for individual subjects Knowledge of other subjects should inform how we interpret scores of a given subject Subject 7 has the highest scores in the data set But maybe we should suspect they are overestimates Subject 4 has the lowest scores in the data set But maybe we should suspect they are underestimates

Simple model l mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] 55. 32 0. 09 3. 35 48. 45 53. 19 55. 41 57. 58 61. 78 1285 1 a[2] 52. 44 0. 10 3. 17 46. 04 50. 40 52. 41 54. 66 58. 53 956 1 a[3] 32. 69 0. 09 3. 10 26. 82 30. 70 32. 60 34. 76 38. 62 1206 1 a[4] 46. 91 0. 09 3. 10 40. 78 44. 87 47. 05 49. 08 52. 88 1279 1 a[5] 36. 07 0. 09 3. 33 30. 20 33. 75 35. 97 38. 40 42. 54 1398 1 ATmodel 1 <- map 2 stan( a[6] 49. 25 0. 08 3. 22 43. 02 47. 09 49. 30 51. 24 55. 59 1436 1 alist( Correct. Responses ~ dnorm(mu, sigma), a[7] 33. 54 0. 09 3. 13 27. 13 31. 57 33. 43 35. 66 39. 75 1356 1 a[8] 52. 69 0. 09 3. 24 46. 05 50. 66 52. 73 54. 77 59. 19 1357 1 mu <- a[Participant] + b 2*Dosage 15 + b 3*Dosage 30 + b 4*Dosage 60, a[9] 31. 29 0. 10 3. 13 25. 19 29. 17 31. 27 33. 29 37. 39 1010 1 a[Participant] ~ dnorm(50, 50), a[10] 39. 22 0. 09 3. 24 32. 78 37. 13 39. 26 41. 32 45. 55 1413 1 a[11] 33. 35 0. 10 3. 18 27. 42 31. 06 33. 47 35. 66 39. 12 1076 1 b 2 ~ dnorm(0, 20), a[12] 39. 36 0. 09 3. 15 33. 01 37. 28 39. 32 41. 34 45. 60 1263 1 b 3 ~ dnorm(0, 20), a[13] 54. 20 0. 10 3. 18 48. 22 51. 94 54. 21 56. 23 60. 55 1064 1 a[14] 32. 52 0. 09 3. 23 26. 16 30. 44 32. 59 34. 48 39. 01 1286 1 b 4 ~ dnorm(0, 20), a[15] 31. 30 0. 10 3. 37 24. 63 29. 06 31. 13 33. 53 38. 07 1211 1 sigma ~ dunif(0, 100) a[16] 34. 75 0. 09 3. 21 28. 31 32. 58 34. 76 36. 96 40. 67 1181 1 a[17] 34. 24 0. 08 3. 12 28. 07 32. 16 34. 16 36. 30 40. 44 1500 1 ), data= ATdata) a[18] 31. 52 0. 09 3. 17 25. 29 29. 50 31. 43 33. 60 38. 15 1253 1 a[19] 26. 05 0. 09 3. 12 19. 90 23. 94 26. 10 28. 05 32. 04 1309 1 a[20] 35. 25 0. 08 3. 12 29. 33 33. 15 35. 21 37. 15 41. 83 1445 1 a[21] 34. 66 0. 08 3. 05 28. 73 32. 62 34. 61 36. 91 40. 54 1511 1 a[22] 64. 66 0. 10 3. 22 58. 28 62. 54 64. 58 66. 87 70. 71 1112 1 a[23] 38. 07 0. 08 3. 34 31. 85 35. 64 38. 08 40. 38 44. 71 1543 1 a[24] 36. 74 0. 09 3. 27 30. 54 34. 61 36. 83 38. 93 43. 10 1237 1 b 2 -0. 14 0. 08 1. 70 -3. 35 -1. 31 -0. 15 1. 06 3. 31 467 1 b 3 4. 47 0. 08 1. 74 1. 11 3. 34 4. 56 5. 61 7. 73 531 1 b 4 4. 82 0. 08 1. 72 1. 48 3. 61 4. 85 6. 03 8. 07 523 1 sigma 6. 09 0. 02 0. 53 5. 17 5. 71 6. 04 6. 40 7. 31 1142 1 lp__ -219. 80 0. 23 4. 36 -228. 86 -222. 68 -219. 53 -216. 59 -212. 11 348 1 Different mean for Dosage 0 for each subject, model deviations for other dosages
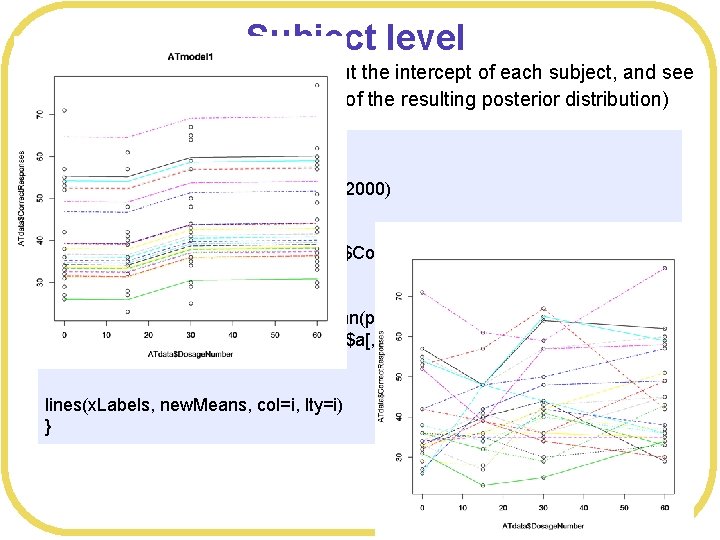
Subject level l From the posterior, we can pull out the intercept of each subject, and see the effect for each subject (mean of the resulting posterior distribution) dev. new() # Look at effect on each subject post<-extract. samples(ATmodel 1, n= 2000) plot(ATdata$Dosage. Number, ATdata$Correct. Responses, main="ATmodel 1") x. Labels<-c(0, 15, 30, 60) for(i in c(1: 24)) { new. Means <- c(mean(post$a[, i]), mean(post$a[, i]+post$b 2), mean(post$a[, i]+post$b 3), mean(post$a[, i]+post$b 4)) lines(x. Labels, new. Means, col=i, lty=i) }
![mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1]](http://slidetodoc.com/presentation_image_h/23fe6d2d391c54169d844a0a1135b272/image-27.jpg)
mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] 57. 05 0. 08 3. 13 51. 15 54. 75 57. 01 59. 23 63. 22 1459 1 a[2] 54. 12 0. 08 3. 14 48. 13 52. 04 54. 18 56. 11 60. 14 1478 1 a[3] 36. 37 0. 10 3. 40 29. 98 34. 20 36. 37 38. 39 43. 52 1052 1 Using default priors for subject intercepts a[4] 49. 21 0. 09 3. 50 42. 00 46. 96 49. 26 51. 44 56. 17 1590 1 a[5] 39. 40 0. 09 3. 24 33. 13 37. 25 39. 36 41. 43 46. 09 1347 1 a[6] 51. 30 0. 11 3. 20 44. 99 49. 21 51. 21 53. 23 57. 65 917 1 a[7] 36. 81 0. 08 3. 39 30. 53 34. 40 36. 76 38. 98 43. 52 1705 1 a[8] 54. 40 0. 09 3. 33 48. 06 52. 18 54. 35 56. 57 61. 15 1394 1 a[9] 34. 87 0. 08 2. 94 29. 09 32. 92 34. 79 36. 84 40. 57 1413 1 ATmodel 2 <- map 2 stan( a[10] 42. 07 0. 09 3. 19 36. 14 39. 80 42. 09 44. 36 48. 40 1193 1 a[11] 36. 67 0. 07 3. 08 30. 70 34. 49 36. 57 38. 63 43. 00 1742 1 alist( Correct. Responses ~ dnorm(mu, sigma), a[12] 42. 19 0. 09 3. 17 36. 19 40. 03 42. 20 44. 24 48. 36 1208 1 mu <- a[Participant] + b 2*Dosage 15 + b 3*Dosage 30 + b 4*Dosage 60, a[13] 55. 83 0. 09 2. 96 50. 02 53. 81 55. 76 57. 87 61. 80 1209 1 a[14] 36. 18 0. 09 3. 16 29. 96 34. 07 36. 12 38. 44 42. 47 1191 1 a[Participant] ~ dnorm(50, 10), a[15] 34. 90 0. 10 3. 11 28. 80 32. 80 34. 84 36. 83 41. 50 918 1 b 2 ~ dnorm(0, 20), a[16] 38. 05 0. 09 3. 08 31. 86 36. 01 37. 99 40. 08 44. 34 1154 1 a[17] 37. 73 0. 11 3. 26 31. 38 35. 59 37. 58 39. 79 44. 50 869 1 b 3 ~ dnorm(0, 20), a[18] 35. 19 0. 09 3. 20 29. 16 33. 10 35. 16 37. 19 41. 72 1197 1 b 4 ~ dnorm(0, 20), a[19] 30. 10 3. 14 24. 15 27. 88 29. 90 32. 25 36. 21 1087 1 sigma ~ dunif(0, 100) a[20] 38. 58 0. 10 3. 17 32. 43 36. 57 38. 58 40. 63 45. 04 1090 1 a[21] 38. 11 0. 09 3. 17 32. 29 35. 76 38. 03 40. 19 44. 51 1201 1 ), data= ATdata) a[22] 65. 30 0. 08 3. 15 58. 87 63. 08 65. 30 67. 35 71. 73 1375 1 a[23] 41. 28 0. 11 3. 23 34. 82 38. 98 41. 40 43. 42 47. 47 889 1 a[24] 39. 89 0. 10 3. 29 33. 83 37. 66 39. 72 41. 86 46. 76 1154 1 b 2 -3. 09 0. 07 1. 73 -6. 57 -4. 21 -3. 07 -1. 94 0. 29 536 1 b 3 1. 61 0. 08 1. 72 -2. 08 0. 47 1. 68 2. 83 4. 90 436 1 b 4 1. 93 0. 08 1. 70 -1. 69 0. 83 1. 94 3. 11 5. 16 455 1 sigma 6. 34 0. 02 0. 56 5. 37 5. 95 6. 30 6. 68 7. 60 726 1 lp__ -239. 54 0. 24 4. 38 -249. 79 -242. 05 -239. 10 -236. 52 -231. 98 340 1 Tighter prior l

Justified? l l In some sense, this is just modeling Extreme data are interpreted as noise, and the model predicts that testing an extreme subject again will result in something more “typical”

Compare models l Favors model with broader sd on prior > compare(ATmodel 1, ATmodel 2) WAIC SE d. WAIC d. SE p. WAIC weight ATmodel 1 645. 6 13. 86 0. 0 NA 23. 4 0. 96 ATmodel 2 651. 9 13. 97 6. 3 5. 33 22. 2 0. 04 >

Hierarchical model > print(summary(ATmodel 3)) Inference for Stan model: 1 db 8 f 5 b 04429 de 2343 e 811 c 594 d 58 f 90. 1 chains, each with iter=2000; warmup=1000; thin=1; post-warmup draws per chain=1000, total post-warmup draws=1000. w Set hyper-parameters for subject intercepts (Dosage 0) mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] 54. 13 0. 09 3. 10 47. 65 52. 05 54. 19 56. 34 59. 87 1089 1 a[2] 51. 40 0. 09 2. 91 45. 63 49. 45 51. 31 53. 35 57. 19 950 1 a[3] 33. 36 0. 09 2. 93 27. 83 31. 40 33. 27 35. 35 39. 18 976 1 a[4] 46. 36 0. 09 3. 00 40. 65 44. 35 46. 37 48. 27 52. 74 1216 1 a[5] 36. 49 0. 10 3. 26 30. 03 34. 32 36. 68 38. 89 42. 29 1001 1 a[6] 48. 52 0. 10 3. 08 42. 75 46. 35 48. 43 50. 62 54. 31 908 1 a[7] 34. 11 0. 10 3. 21 28. 08 32. 04 34. 01 36. 22 40. 35 950 1 a[8] 51. 58 0. 09 3. 07 45. 45 49. 56 51. 69 53. 69 57. 36 1107 1 a[9] 31. 96 0. 10 3. 05 26. 35 29. 92 31. 94 33. 92 38. 12 963 1 a[10] 39. 27 0. 10 3. 23 32. 94 37. 19 39. 24 41. 49 45. 28 1001 1 a[11] 33. 79 0. 10 3. 13 27. 70 31. 72 33. 72 35. 77 40. 48 982 1 a[12] 39. 29 0. 09 3. 04 33. 64 37. 28 39. 26 41. 25 45. 50 1196 1 a[13] 52. 91 0. 09 3. 16 46. 53 50. 94 52. 95 55. 05 58. 78 1178 1 a[14] 33. 10 0. 09 3. 12 27. 31 30. 92 33. 01 35. 19 39. 35 1189 1 a[15] 32. 05 0. 10 3. 25 25. 50 29. 95 31. 99 34. 10 38. 57 1131 1 a[16] 35. 23 0. 09 3. 28 29. 20 32. 93 35. 17 37. 49 41. 76 1321 1 a[17] 34. 75 0. 09 3. 12 28. 75 32. 72 34. 78 36. 81 40. 85 1115 1 a[18] 32. 23 0. 10 3. 24 25. 91 30. 07 32. 19 34. 38 38. 73 1017 1 a[19] 27. 33 0. 10 3. 17 21. 38 25. 28 27. 25 29. 33 33. 51 1065 1 a[20] 35. 66 0. 10 3. 11 29. 36 33. 53 35. 75 37. 77 41. 82 1022 1 a[21] 35. 19 0. 09 3. 19 29. 12 33. 05 35. 07 37. 36 41. 94 1140 1 a[22] 62. 55 0. 10 3. 25 55. 93 60. 39 62. 57 64. 64 68. 82 1119 1 a[23] 38. 31 0. 10 3. 12 32. 47 36. 13 38. 34 40. 37 44. 61 931 1 a[24] 37. 11 0. 10 3. 15 30. 94 35. 12 37. 19 39. 04 43. 64 938 1 Gmean 39. 88 0. 09 2. 38 35. 24 38. 35 39. 96 41. 53 44. 49 669 1 Gsd 9. 99 0. 05 1. 68 7. 25 8. 76 9. 74 11. 05 13. 63 1351 1 b 2 -0. 19 0. 08 1. 67 -3. 37 -1. 32 -0. 21 0. 91 3. 06 437 1 b 3 4. 47 0. 08 1. 72 0. 83 3. 39 4. 52 5. 57 7. 72 443 1 b 4 4. 83 0. 09 1. 71 1. 27 3. 82 4. 87 5. 92 8. 09 391 1 sigma 6. 08 0. 02 0. 54 5. 14 5. 72 6. 04 6. 41 7. 28 748 1 lp__ -283. 52 0. 25 4. 60 -293. 38 -286. 30 -283. 16 -280. 28 -275. 33 334 1 ATmodel 3 <- map 2 stan( alist( Correct. Responses ~ dnorm(mu, sigma), mu <- a[Participant] + b 2*Dosage 15 + b 3*Dosage 30 + b 4*Dosage 60, a[Participant] ~ dnorm(Gmean, Gsd), Gmean ~ dnorm(50, 50), Gsd ~ dunif(0, 50), b 2 ~ dnorm(0, 20), b 3 ~ dnorm(0, 20), b 4 ~ dnorm(0, 20), sigma ~ dunif(0, 100) ), data= ATdata) cat("Finished ATmodel 3n") print(summary(ATmodel 3))

Model comparison l From the posterior, we can pull out the intercept of each subject, and see the effect for each subject (mean of the resulting posterior distribution) > compare(ATmodel 1, ATmodel 2, ATmodel 3) WAIC SE d. WAIC d. SE p. WAIC weight ATmodel 3 644. 7 13. 02 0. 0 NA 22. 4 0. 60 ATmodel 1 645. 6 13. 86 0. 9 3. 53 23. 4 0. 38 ATmodel 2 651. 9 13. 97 7. 2 4. 33 22. 2 0. 02 > l This part is really kind of optional. You could just fit the hierarchical model and examine the poster distributions to interpret the analysis

> print(summary(ATmodel 4)) Inference for Stan model: a 79 c 6 d 6669 e 2900877957 d 0 bed 988 e 2 f. 1 chains, each with iter=2000; warmup=1000; thin=1; post-warmup draws per chain=1000, total post-warmup draws=1000. Shrink intercepts l mean se_mean sd 2. 5% 25% 50% 75% 97. 5% n_eff Rhat a[1] 53. 88 0. 14 3. 16 47. 96 51. 61 53. 98 55. 98 60. 10 544 1 a[2] 51. 32 0. 12 3. 14 45. 52 49. 15 51. 26 53. 43 57. 40 658 1 a[3] 33. 39 0. 13 3. 08 27. 37 31. 26 33. 36 35. 59 39. 44 523 1 a[4] 46. 49 0. 12 2. 97 40. 53 44. 61 46. 44 48. 38 52. 31 656 1 a[5] 36. 57 0. 13 3. 20 30. 44 34. 41 36. 54 38. 66 43. 01 625 1 a[6] 48. 41 0. 10 3. 08 42. 30 46. 48 48. 42 50. 42 54. 64 952 1 a[7] 34. 05 0. 13 3. 00 27. 95 32. 13 34. 11 35. 97 39. 89 545 1 a[8] 51. 59 0. 11 3. 02 45. 82 49. 55 51. 56 53. 47 57. 59 793 1 a[9] 32. 02 0. 12 3. 00 26. 08 30. 09 31. 95 34. 03 38. 01 620 1 a[10] 39. 34 0. 12 3. 32 32. 84 37. 15 39. 31 41. 50 45. 97 754 1 a[11] 33. 99 0. 11 3. 03 27. 93 31. 99 34. 02 36. 04 39. 61 733 1 a[12] 39. 41 0. 10 3. 04 33. 51 37. 37 39. 24 41. 38 45. 32 854 1 a[13] 52. 93 0. 14 3. 12 46. 79 50. 75 52. 99 55. 06 59. 29 484 1 a[14] 33. 18 0. 12 3. 14 26. 85 31. 04 33. 20 35. 21 39. 31 669 1 a[15] 32. 04 0. 11 3. 10 25. 72 30. 07 32. 11 34. 00 37. 76 848 1 a[16] 35. 22 0. 12 3. 08 29. 31 33. 10 35. 18 37. 47 41. 18 672 1 a[17] 34. 68 0. 11 2. 99 28. 87 32. 66 34. 63 36. 76 40. 48 712 1 a[18] 32. 32 0. 12 3. 12 26. 10 30. 26 32. 33 34. 45 38. 67 657 1 a[19] 27. 25 0. 13 3. 01 21. 32 25. 16 27. 23 29. 16 33. 22 564 1 a[20] 35. 68 0. 12 3. 18 29. 43 33. 65 35. 75 37. 70 42. 14 744 1 a[21] 35. 21 0. 12 3. 03 29. 14 33. 20 35. 14 37. 27 40. 97 675 1 a[22] 62. 48 0. 10 3. 08 56. 97 60. 20 62. 48 64. 54 68. 58 888 1 a[23] 38. 38 0. 13 3. 05 32. 59 36. 34 38. 27 40. 58 44. 03 596 1 a[24] 37. 07 0. 10 3. 05 30. 99 35. 05 37. 06 39. 11 43. 04 855 1 Gmean 39. 90 0. 11 2. 40 35. 09 38. 27 39. 90 41. 60 44. 48 466 1 Gsd 9. 99 0. 06 1. 71 7. 28 8. 78 9. 78 11. 05 13. 83 775 1 b 2 0. 15 0. 10 1. 76 -3. 26 -1. 03 0. 14 1. 34 3. 69 287 1 b 3 4. 27 0. 09 1. 67 1. 16 3. 13 4. 19 5. 38 7. 60 375 1 b 4 4. 61 0. 10 1. 70 1. 35 3. 51 4. 56 5. 78 7. 94 308 1 Gbmean 3. 14 0. 21 4. 16 -5. 08 1. 10 2. 99 5. 12 12. 08 399 1 Gbsd 6. 76 0. 41 5. 68 1. 19 2. 79 4. 82 8. 66 23. 29 187 1 sigma 6. 10 0. 02 0. 55 5. 10 5. 72 6. 07 6. 44 7. 20 991 1 lp__ -288. 02 0. 27 4. 54 -297. 15 -290. 90 -287. 71 -284. 84 -280. 16 289 1 We can also shrink things at the mean (averaging across subjects) ATmodel 4 <- map 2 stan( alist( Correct. Responses ~ dnorm(mu, sigma), mu <- a[Participant] + b 2*Dosage 15 + b 3*Dosage 30 + b 4*Dosage 60, a[Participant] ~ dnorm(Gmean, Gsd), Gmean ~ dnorm(50, 50), Gsd ~ dunif(0, 50), b 2 ~ dnorm(Gbmean, Gbsd), b 3 ~ dnorm(Gbmean, Gbsd), b 4 ~ dnorm(Gbmean, Gbsd), Gbmean ~ dnorm(0, 20), Gbsd ~ dunif(0, 30), sigma ~ dunif(0, 100) ), data= ATdata)
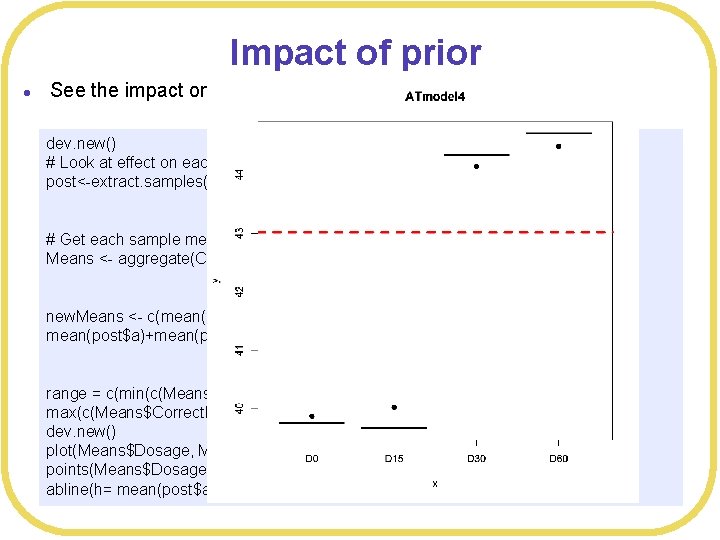
Impact of prior l See the impact on the means dev. new() # Look at effect on each subject post<-extract. samples(ATmodel 4, n= 2000) # Get each sample mean Means <- aggregate(Correct. Responses~Dosage, FUN=mean, data=ATdata) new. Means <- c(mean(post$a), mean(post$a)+mean(post$b 2), mean(post$a)+mean(post$b 3), mean(post$a)+mean(post$b 4)) range = c(min(c(Means$Correct. Responses, new. Means)), max(c(Means$Correct. Responses, new. Means))) dev. new() plot(Means$Dosage, Means$Correct. Responses, main="ATmodel 4", ylim=range) points(Means$Dosage, new. Means, pch=19) abline(h= mean(post$a)+mean(post$Gbmean), col="red", lwd=3, lty=2)

Compare models l The hierarchical models beat the others w Not much difference between them l Usually, it is advantageous to let the hierarchical structure guide prior developememt w Saves you the trouble of justifying your priors! > compare(ATmodel 1, ATmodel 2, ATmodel 3, ATmodel 4) WAIC SE d. WAIC d. SE p. WAIC weight ATmodel 3 644. 7 13. 02 0. 0 NA 22. 4 0. 41 ATmodel 4 645. 3 13. 25 0. 5 1. 06 22. 7 0. 31 ATmodel 1 645. 6 13. 86 0. 9 3. 53 23. 4 0. 26 ATmodel 2 651. 9 13. 97 7. 2 4. 33 22. 2 0. 01 >
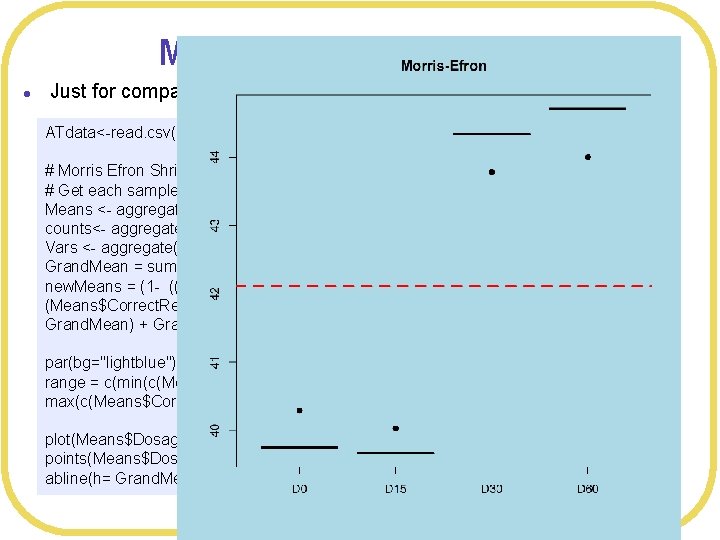
Morris-Efron shrinkage l Just for comparison ATdata<-read. csv(file="ADHDTreatment. csv", header=TRUE, strings. As. Factors=TRUE) # Morris Efron Shrinkage # Get each sample mean Means <- aggregate(Correct. Responses~Dosage, FUN=mean, data=ATdata) counts<- aggregate(Correct. Responses~Dosage, FUN=length, data=ATdata) Vars <- aggregate(Correct. Responses~Dosage, FUN=var, data=ATdata) Grand. Mean = sum(Means$Correct. Responses)/4 new. Means = (1 - ((4 -3))*Vars$Correct. Responses/counts$Correct. Responses /sum( (Means$Correct. Responses - Grand. Mean)^2 )) *(Means$Correct. Responses - Grand. Mean) + Grand. Mean par(bg="lightblue") range = c(min(c(Means$Correct. Responses, new. Means)), max(c(Means$Correct. Responses, new. Means))) plot(Means$Dosage, Means$Correct. Responses, main="Morris-Efron", ylim=range) points(Means$Dosage, new. Means, pch=19) abline(h= Grand. Mean, col="red", lwd=3, lty=2)

Conclusions l Shrinkage l Side effect of prior across a group l Helps prediction l Need “appropriate” prior w Not too tight l Hierarchical models are a pretty good option for this w. Not perfect w. Informative priors are even better, if you have them
- Slides: 36