Bayesian style ANOVA Greg Francis PSY 626 Bayesian

Bayesian style ANOVA Greg Francis PSY 626: Bayesian Statistics for Psychological Science Fall 2020 Purdue University

Smiles and Leniency l La. France & Hect (1995): vignette describes a person committing some minor infraction. A photo shows the person with different smile types (or neutral) l The question is how lenient subjects are in punishing the person in the vignette l Each subject contributes one score
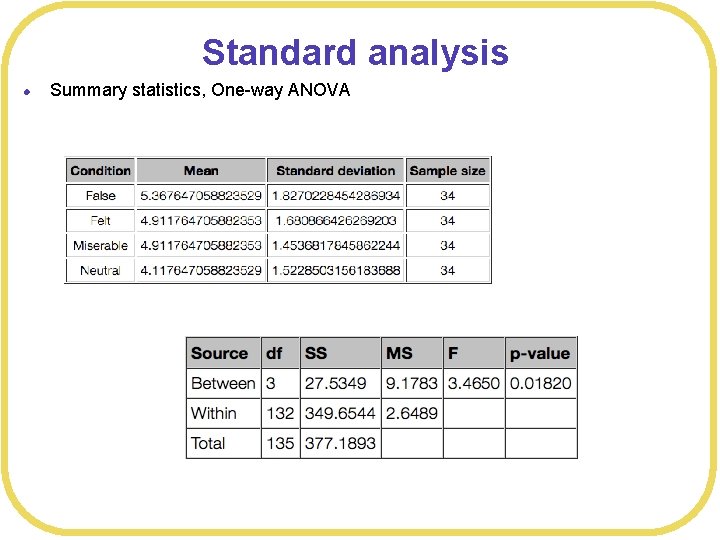
Standard analysis l Summary statistics, One-way ANOVA
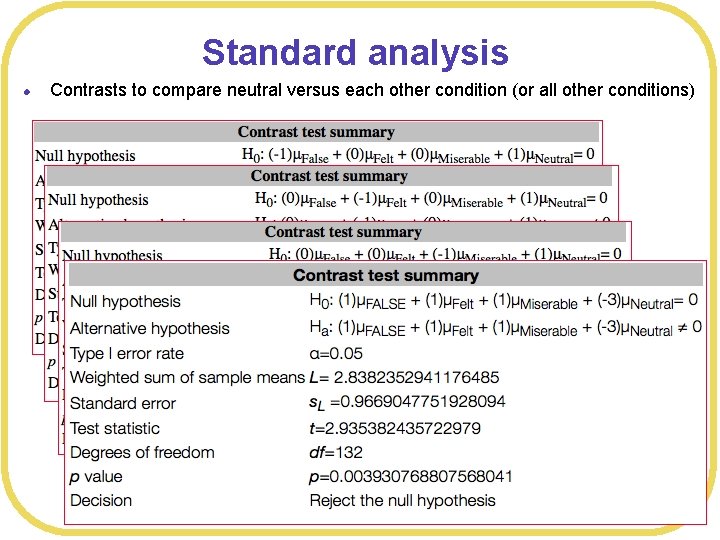
Standard analysis l Contrasts to compare neutral versus each other condition (or all other conditions)

Bayesian variation of ANOVA l Follow along by downloading “Smiles. Leniency. csv” and “Smiles. Leniency 1. R” from the class web site # load the rethinking library(rethinking) # load data file SLdata<-read. csv(file="Smiles. Leniency. csv", header=TRUE, strings. As. Factors=TRUE) > head(SLdata) Subject. ID Smile. Type Leniency 1 Subject 1 FALSE 2. 5 2 Subject 2 FALSE 5. 5 3 Subject 3 FALSE 6. 5 4 Subject 4 FALSE 3. 5 5 Subject 5 FALSE 3. 0 6 Subject 6 FALSE 3. 5

Define the null model l # Null model: one mean for all smile types l SLmodel 0 <- map( l alist( Leniency ~ dnorm(mu, sigma), l mu <- a, l a~ dnorm(5, 5), l sigma ~ dunif(0, 10) l ), data= SLdata ) l cat("Finished SLmodel 0n") l Leniency scores are between 0 and 10, so these priors should cover pretty much the entire range of possible values

Null model output l print(SLmodel 0) Quadratic approximate posterior distribution Formula: Leniency ~ dnorm(mu, sigma) mu <- a a ~ dnorm(5, 5) sigma ~ dunif(0, 10) Posterior means: a sigma 4. 827347 1. 665369 Log-likelihood: -262. 34

Define the alternative model l # Alternative model: different means for different smile types l Make an index for each smile type: l SLdata$False <- ifelse(SLdata$Smile. Type =="FALSE", 1, 0) l SLdata$Felt <- ifelse(SLdata$Smile. Type =="Felt", 1, 0) l SLdata$Miserable <- ifelse(SLdata$Smile. Type =="Miserable", 1, 0) l SLdata$Neutral <- ifelse(SLdata$Smile. Type =="Neutral", 1, 0) > SLdata$Felt [1] 0 0 0 0 0 0 0 0 0 1 1 1 111111111100000 [79] 0 0 0 0 0 0 0 0 0 0 0 0 00000 > SLdata$Neutral [1] 0 0 0 0 0 0 0 0 0 0 0 0 000000000000000 [79] 0 0 0 0 0 0 1 1 1 1 1 1 11111 >

Define the alternative model l # Alternative model 1: different mean for each condition (dumy variable coding) l SLmodel 1 <- map( l alist( Leniency ~ dnorm(mu, sigma), l mu <- a 1*False + a 2*Felt + a 3*Miserable + a 4*Neutral, l a 1 ~ dnorm(5, 5), l a 2 ~ dnorm(5, 5), l a 3 ~ dnorm(5, 5), l a 4 ~ dnorm(5, 5), l sigma ~ dunif(0, 10) l ), data= SLdata ) Quadratic approximate posterior distribution Formula: Leniency ~ dnorm(mu, sigma) mu <- a 1 * False + a 2 * Felt + a 3 * Miserable + a 4 * Neutral a 1 ~ dnorm(5, 5) a 2 ~ dnorm(5, 5) a 3 ~ dnorm(5, 5) a 4 ~ dnorm(5, 5) sigma ~ dunif(0, 10) Posterior means: a 1 a 2 a 3 a 4 sigma 5. 366476 4. 912051 4. 912099 4. 120332 1. 603450 Log-likelihood: -257. 19 >

Comparing models l Which model best predicts future data? > compare(SLmodel 0, SLmodel 1) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 1 524. 4 15. 14 0 NA 4. 9 0. 88 SLmodel 0 528. 4 14. 65 4 6. 61 1. 7 0. 12 l Smaller WAIC for SLmodel 1, which suggests that a model having different means for each smile type will do better than a model using the same mean for different smile types w Kind of like indicating a significant result from an ANOVA l Any further conclusions (similar to contrasts) can be based on the posterior distributions
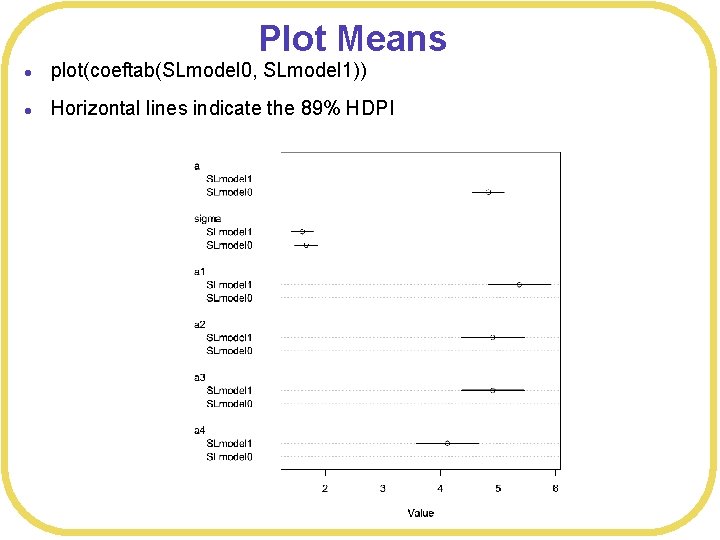
Plot Means l plot(coeftab(SLmodel 0, SLmodel 1)) l Horizontal lines indicate the 89% HDPI
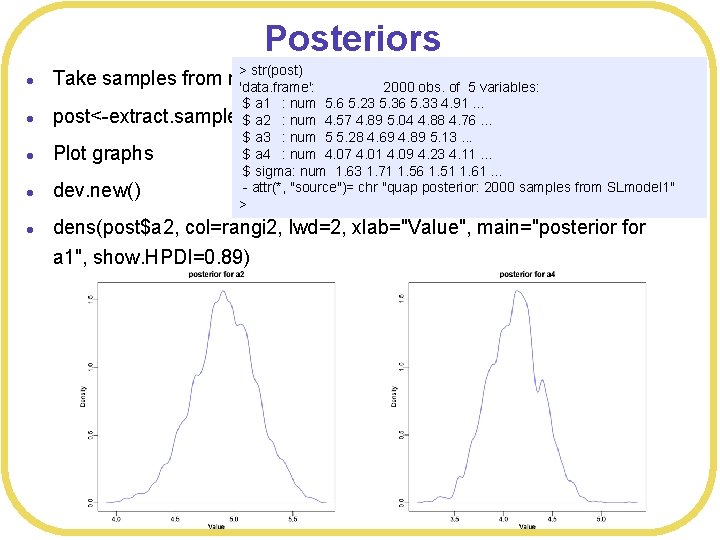
Posteriors l l l > str(post) Take samples from model 'data. frame': 2000 obs. of 5 variables: $ a 1 : num 5. 6 5. 23 5. 36 5. 33 4. 91. . . post<-extract. samples(SLmodel 1, n= 2000) $ a 2 : num 4. 57 4. 89 5. 04 4. 88 4. 76. . . $ a 3 : num 5 5. 28 4. 69 4. 89 5. 13. . . $ a 4 : num 4. 07 4. 01 4. 09 4. 23 4. 11. . . Plot graphs $ sigma: num 1. 63 1. 71 1. 56 1. 51 1. 61. . . - attr(*, "source")= chr "quap posterior: 2000 samples from SLmodel 1" dev. new() > dens(post$a 2, col=rangi 2, lwd=2, xlab="Value", main="posterior for a 1", show. HPDI=0. 89)

Probabilities What is the probability the mean for a neutral smile type (a 4) is smaller than the mean for a felt smile type (a 2)? l > length(post$a 4[post$a 4 <= post$a 2])/length(post$a 4) [1] 0. 9805 What is the probability the mean for a neutral smile type (a 4) is smaller than the mean for a false smile type (a 1)? l > length(post$a 4[post$a 4 <= post$a 1])/length(post$a 4) [1] 1 What is the probability the mean for a neutral smile type (a 4) is smaller than the mean for a miserable smile type (a 3)? l length(post$a 4[post$a 4 <= post$a 3])/length(post$a 4) [1] 0. 9815 l What is the probability the mean for a miserable smile type (a 3) is smaller than the mean for a felt smile type (a 2)? length(post$a 3[post$a 3 <= post$a 2])/length(post$a 3) [1] 0. 5075

Probabilities What is the probability the mean for a felt smile type (a 2) is smaller than the mean for a false smile type (a 1)? l length(post$a 2[post$a 2 <= post$a 1])/length(post$a 2) [1] 0. 8765 > l Is this “almost” significant? l You have to define what you mean by “significant” w There is no control of Type I error here, setting alpha=0. 05 does not make sense l I think the more reasonable thing is to say that the probability is around 0. 88 that the mean leniency rating for a felt smile is lower than the mean for a false smile w If that is even a comparison you are interested in (you don’t have to be, you know) w The estimated means are simply what they are, and that might be good enough for what you want to do

Other models l You could set up a model that is similar to some contrasts l # Contrast model: neutral versus any smile type l SLmodel 2 <- map( l alist( Leniency ~ l mu <- a 1*(1 -Neutral) + a 4*Neutral, > compare(SLmodel 0, SLmodel 2) l a 1 ~ dnorm(5, 5), l a 4 ~ dnorm(5, 5), l sigma ~ dunif(0, 10) l ), data= SLdata ) l cat("Finished SLmodel 2n") l print(compare(SLmodel 0, SLmodel 1, SLmodel 2)) WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 2 522. 3258 14. 60239 0. 000000 NA 2. 883889 0. 78084359 SLmodel 1 525. 1976 15. 18323 2. 871707 3. 046096 5. 259080 0. 18577222 dnorm(mu, sigma), SLmodel 0 528. 6304 14. 75359 6. 304585 5. 691067 1. 835595 0. 03338419 WAIC SE d. WAIC d. SE p. WAIC weight SLmodel 2 522. 2378 14. 48483 0. 000000 NA 2. 813139 0. 95934283 SLmodel 0 528. 5600 14. 68716 6. 322146 5. 738711 1. 801080 0. 04065717

Alternative alternative model l Using dummy variables can get complicated l A different format (but mathematically identical) is to use indices l SLdata$Condition. Index <-0*SLdata$Leniency # just makes an array of the right length, filled with 0's l SLdata$Condition. Index[SLdata$Smile. Type =="FALSE"] =1 l SLdata$Condition. Index[SLdata$Smile. Type =="Felt"] =2 l SLdata$Condition. Index[SLdata$Smile. Type =="Miserable"] =3 l SLdata$Condition. Index[SLdata$Smile. Type =="Neutral"] =4 l New variable, Condition. Index= 1, 2, 3, or 4, depending on the smile type

Alternative alternative model l Model set up: >SLmodel 1 Quadratic approximate posterior distribution SLmodel 3 <- map( alist( Leniency ~ dnorm(mu, Formula: sigma), Leniency ~ dnorm(mu, sigma) mu <- a[Condition. Index], mu <- a 1 * False + a 2 * Felt + a 3 * Miserable + a 4 * Neut a 1 5), ~ dnorm(5, 5) a[Condition. Index] ~ dnorm(5, a 2 ~ dnorm(5, 5) sigma ~ dunif(0, 10) a 3 ~ dnorm(5, 5) a 4 ~ dnorm(5, 5) ), data= SLdata ) Quadratic approximate posterior distribution sigma ~ dunif(0, 10) Formula: Leniency ~ dnorm(mu, sigma) mu <- a[Condition. Index] ~ dnorm(5, 5) sigma ~ dunif(0, 10) Posterior means: a 1 a 2 a 3 a 4 sigma 5. 366541 4. 912032 4. 120306 1. 603431 Log-likelihood: -257. 19 Posterior means: a[1] a[2] a[3] a[4] sigma 5. 366594 4. 912045 4. 912006 4. 120315 1. 603429 Log-likelihood: -257. 19

Posterior l Extracting data from posterior involves a bit different syntax to get the “column” for each parameter value post<-extract. samples(SLmodel 3, n= 2000) > str(post) List of 2 $ sigma: num [1: 2000] 1. 53 1. 62 1. 5 1. 47. . . $ a : num [1: 2000, 1: 4] 5. 52 5. 61 5. 53 4. 89 4. 82. . . - attr(*, "source")= chr "quap posterior: 2000 samples from SLmodel 3" > >cat("Probability neutral mean is smaller than felt mean: n") Probability neutral mean is smaller than felt mean: > print( length(post$a[, 4][post$a[, 4] <= post$a[, 2]])/length(post$a[, 4]) ) [1] 0. 976

ANOVA versus Bayesian l l l Note, we get the same pattern of results either way It looks like the Neutral condition is different from each of the other conditions Don’t we worry about multiple comparisons? w Not really w The Bayesian analysis is not about making decisions, it is about estimating parameter values, given the prior and the data w You might make Type I errors, but you cannot control the Type I error rate, anyhow

Using priors l SLmodel 3 <- map( l alist( Leniency ~ dnorm(mu, sigma), l mu <- a[Condition. Index], l a[Condition. Index] ~ dnorm(5, 5), l sigma ~ dunif(0, 10) l ), data= SLdata ) l Class activity: try to “break” the analysis by using crazy priors

Conclusions l Bayesian ANOVA l Model comparison l Probability statements l No multiple comparisons issues (because no decisions, yet) l And controlling Type I error is not feasible, anyhow l Priors
- Slides: 21