VLSH Voronoibased Locality Sensitive Hashing Sungeui Yoon Authors

VLSH: Voronoi-based Locality Sensitive Hashing Sung-eui Yoon Authors: Lin Loi, Jae-Pil Heo, Junghwan Lee, and Sung-Eui Yoon KAIST http: //sglab. kaist. ac. kr/VLSH/

Main Goals ● Provide efficient nearest neighbor search for various motion planners, PRM and RRT ● Works with high-dimensional data sets ● Supports a diverse set of distance metrics 2
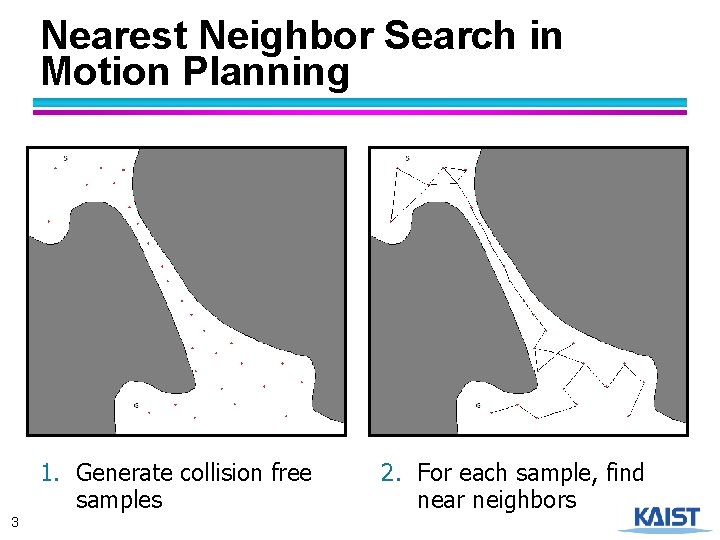
Nearest Neighbor Search in Motion Planning 1. Generate collision free samples 3 2. For each sample, find near neighbors
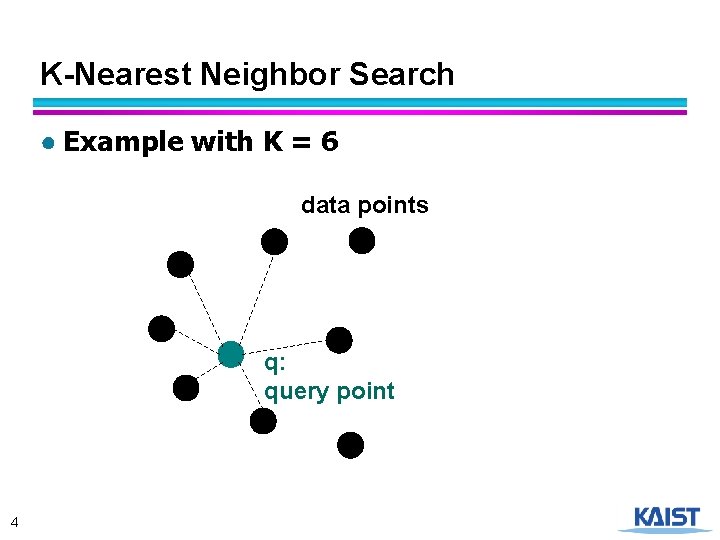
K-Nearest Neighbor Search ● Example with K = 6 data points q: query point 4
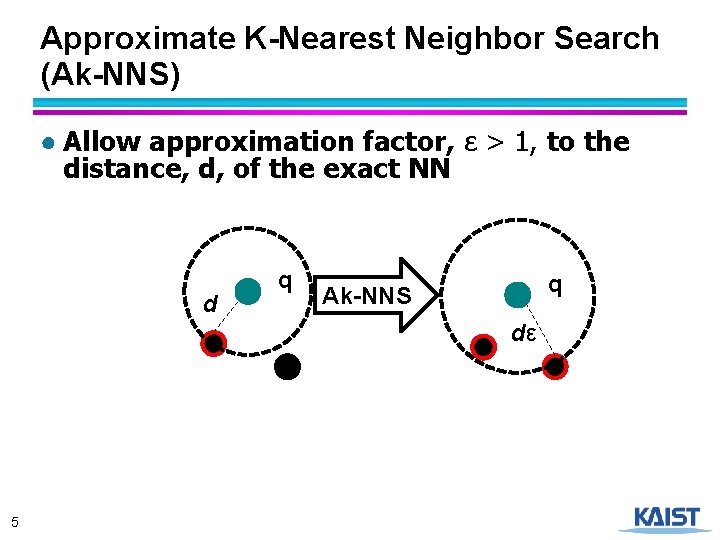
Approximate K-Nearest Neighbor Search (Ak-NNS) ● Allow approximation factor, ε > 1, to the distance, d, of the exact NN d q q Ak-NNS dε 5

Main Contributions ● 6

Previous Work ● Spatial subdivision data structures (e. g. , kd-trees) ● Suffer from the curse of dimensionality [Weber et al. 1998] ● Culling techniques using the triangle inequality [Chavez et al. 01] ● Geometric Near Access Tree (GNAT) [Brin, 95] ● Used widely at OOPSMP and OMPL ● GPU based acceleration [Pan et al. , 10] 7

Previous Work on LSH and Embedding ● Locality Sensitivity Hashing (LSH) ● Fast algorithms for high-dimensional NNS problems [Datar and Indyk 2004] ● Supports for a limited set of motion planning distance metrics (e. g. , Euler angles) [Pan et al. 2010] ● Embedding ● Well studied topic [Indyk et al. , 04] ● Embed motion planning spaces to the Euclidean space [Plaku and Kavraki 2006] 8
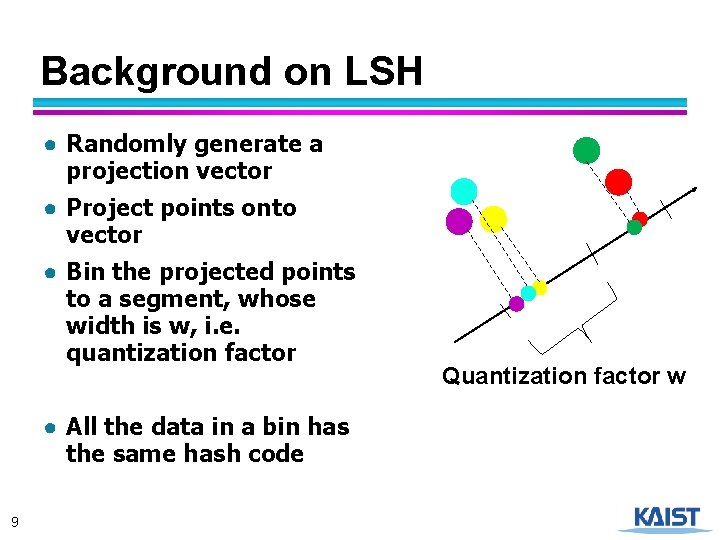
Background on LSH ● Randomly generate a projection vector ● Project points onto vector ● Bin the projected points to a segment, whose width is w, i. e. quantization factor ● All the data in a bin has the same hash code 9 Quantization factor w
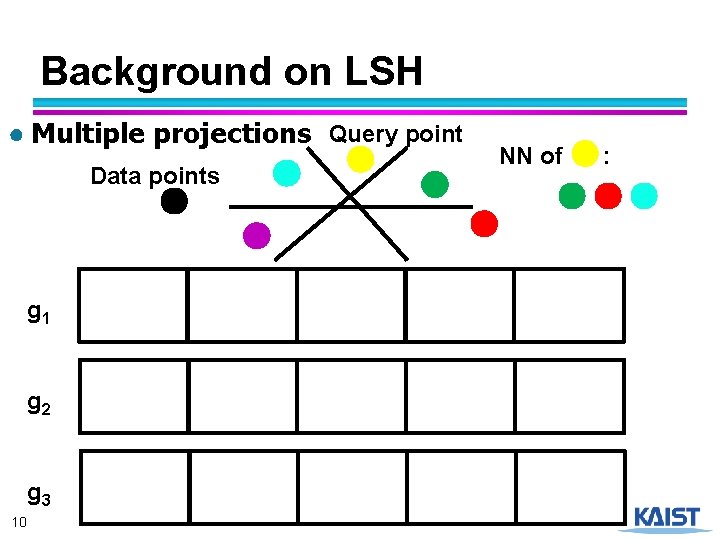
Background on LSH ● Multiple projections Query point Data points g 1 g 2 g 3 10 NN of :

Issues of LSH ● 11
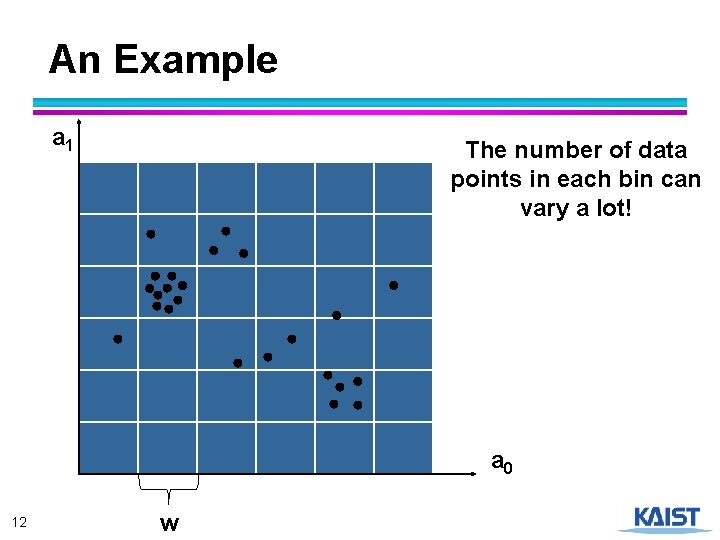
An Example a 1 12 w The number of data points in each bin can vary a lot! a 0
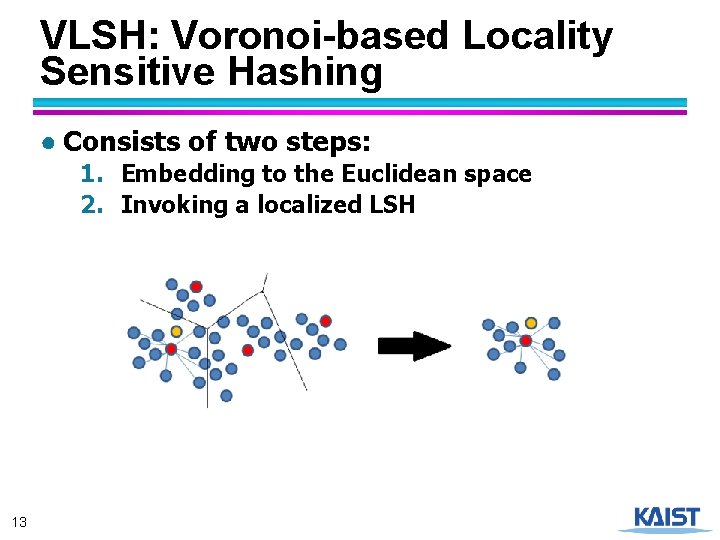
VLSH: Voronoi-based Locality Sensitive Hashing ● Consists of two steps: 1. Embedding to the Euclidean space 2. Invoking a localized LSH 13
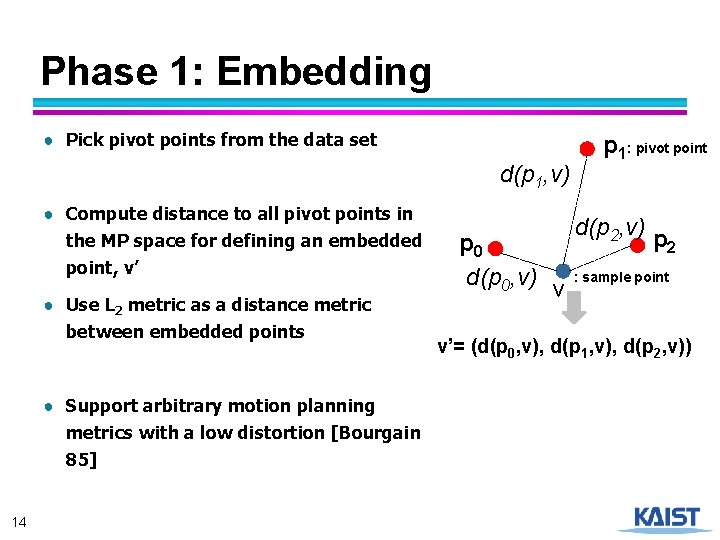
Phase 1: Embedding ● Pick pivot points from the data set d(p 1, v) ● Compute distance to all pivot points in the MP space for defining an embedded point, v’ ● Use L 2 metric as a distance metric between embedded points ● Support arbitrary motion planning metrics with a low distortion [Bourgain 85] 14 p 0 d(p 0, v) p 1: pivot point d(p 2, v) p 2 v : sample point v’= (d(p 0, v), d(p 1, v), d(p 2, v))
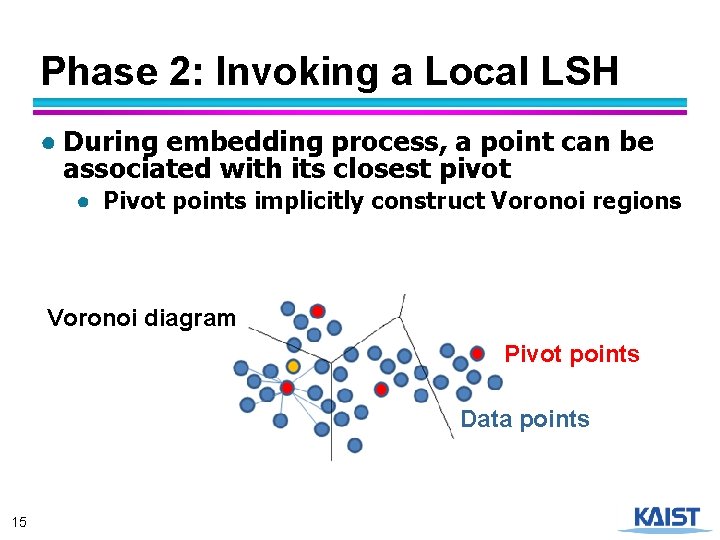
Phase 2: Invoking a Local LSH ● During embedding process, a point can be associated with its closest pivot ● Pivot points implicitly construct Voronoi regions Voronoi diagram Pivot points Data points 15
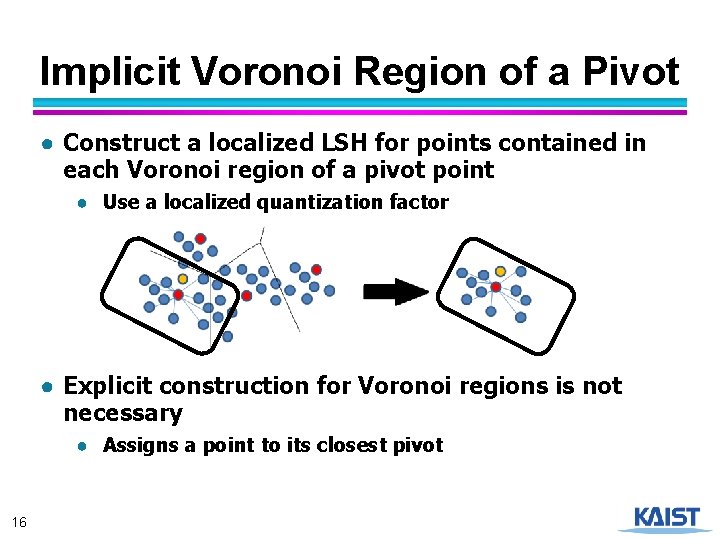
Implicit Voronoi Region of a Pivot ● Construct a localized LSH for points contained in each Voronoi region of a pivot point ● Use a localized quantization factor ● Explicit construction for Voronoi regions is not necessary ● Assigns a point to its closest pivot 16
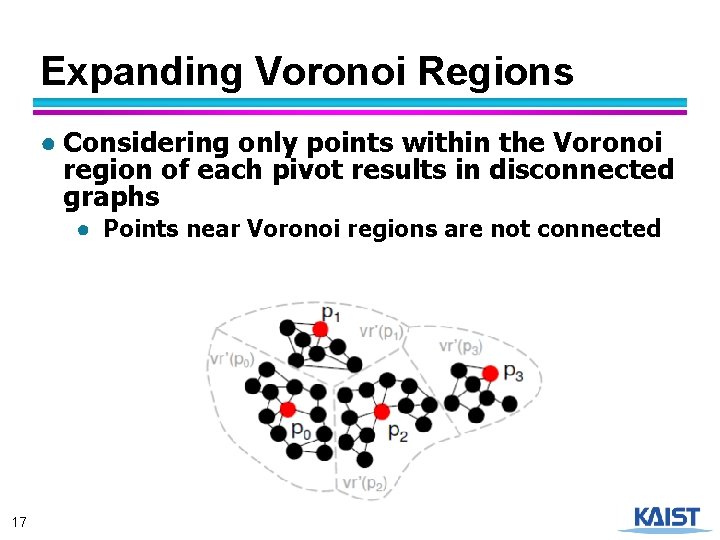
Expanding Voronoi Regions ● Considering only points within the Voronoi region of each pivot results in disconnected graphs ● Points near Voronoi regions are not connected 17
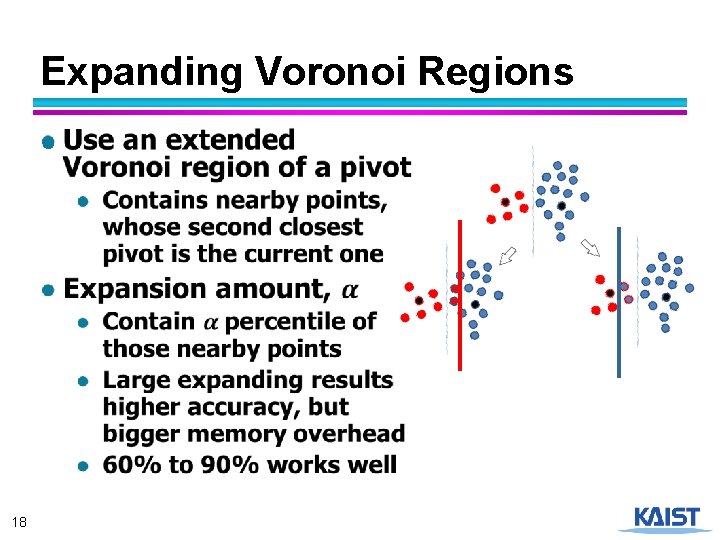
Expanding Voronoi Regions ● 18
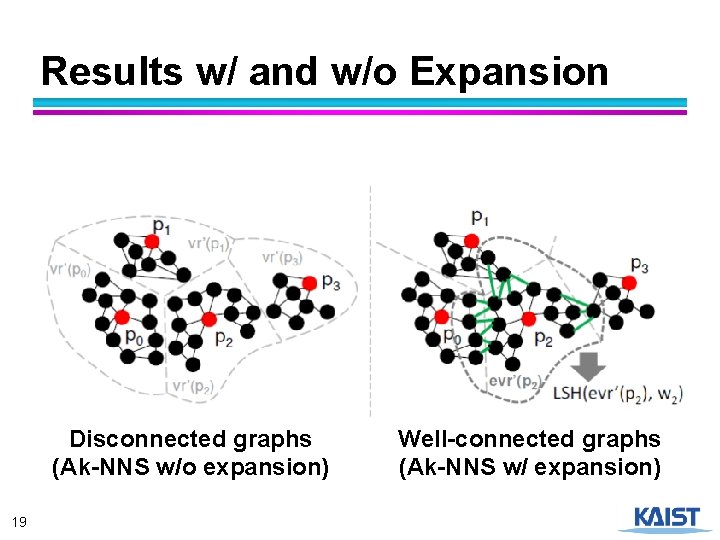
Results w/ and w/o Expansion Disconnected graphs (Ak-NNS w/o expansion) 19 Well-connected graphs (Ak-NNS w/ expansion)

Query-Time Algorithm ● Compute the embedded point from a query point ● Find the closest pivot point and use its localized LSH ● Return candidate nearest neighbors located in hash-buckets of the localized LSH 20

Test Configurations ● Intel i 7 3. 3 GHz CPU with C++ implementation ● Compare our method against LSH and GNAT (OOPSMP) ● Test them with samples from a PRM planner ● 15 nearest neighbor search, i. e. k = 15 ● 10 pivots for our method 21

Benchmarks ● Wiper: 6 dimensions (1 robot for the wiper) ● Bug trap: ● 24 dimensions: 4 rod robots ● 36 dimensions: 6 rod robots 22
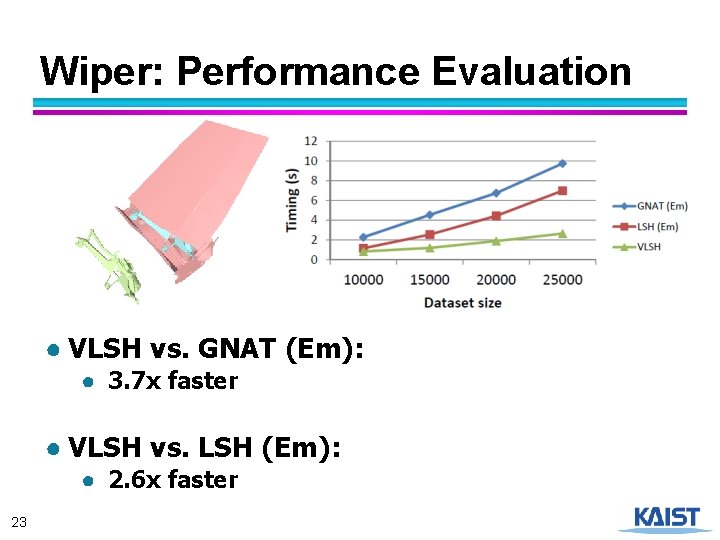
Wiper: Performance Evaluation ● VLSH vs. GNAT (Em): ● 3. 7 x faster ● VLSH vs. LSH (Em): ● 2. 6 x faster 23
![Wiper: Quality Evaluation ● Measure fractional distance error (fde) [Plaku and Kavraki 06] ● Wiper: Quality Evaluation ● Measure fractional distance error (fde) [Plaku and Kavraki 06] ●](http://slidetodoc.com/presentation_image_h/22f6c3006a9dad4e4b3651ab96193ed4/image-24.jpg)
Wiper: Quality Evaluation ● Measure fractional distance error (fde) [Plaku and Kavraki 06] ● Measure difference between computed approximate and ground truth results ● Lower values indicate more accurate results 24
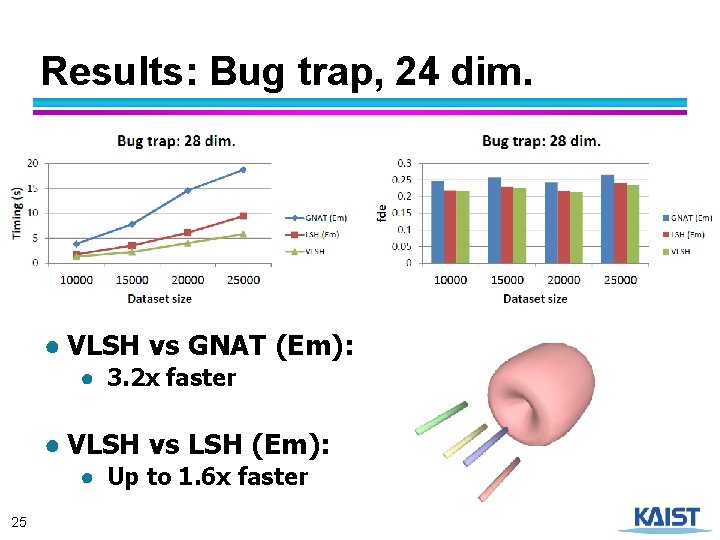
Results: Bug trap, 24 dim. ● VLSH vs GNAT (Em): ● 3. 2 x faster ● VLSH vs LSH (Em): ● Up to 1. 6 x faster 25
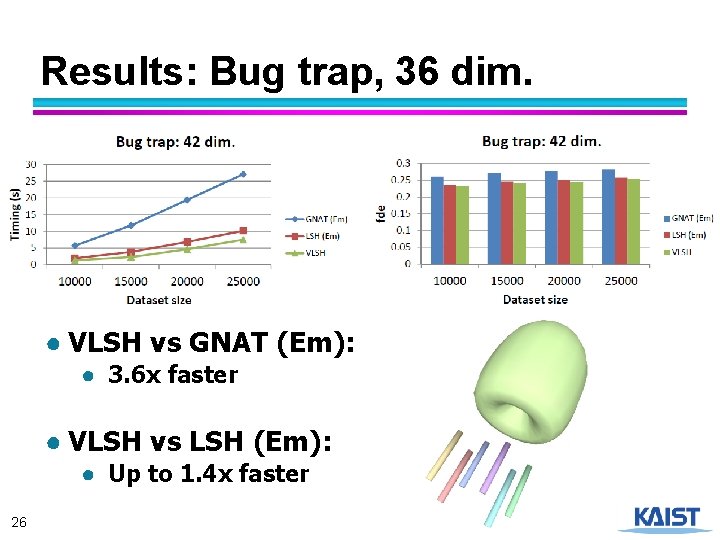
Results: Bug trap, 36 dim. ● VLSH vs GNAT (Em): ● 3. 6 x faster ● VLSH vs LSH (Em): ● Up to 1. 4 x faster 26

Conclusions ● Fast approximate nearest neighbor search algorithm for high-dimensional motion planning problems ● Achieve up to 3. 7 x faster running time over prior approaches ● Supports all distance metrics and consider data distributions for higher accuracy 27

Limitation and Future Work ● Memory overhead ● Duplicate points in our method ● Overall it is not significant, since it takes tens of MB in the testes cases ● Support RRTs that dynamically generate data points ● Supports GPUs for higher performance 28

Acknowledgements ● Anonymous reviewers ● Our funding agency Project webpage: http: //sglab. kaist. ac. kr/VLSH/ 29
- Slides: 29