RNA Secondary Structure Prediction Dynamic Programming Approaches Sarah










- Slides: 22

RNA Secondary Structure Prediction Dynamic Programming Approaches Sarah Aerni http: //www. tbi. univie. ac. at/

Outline l l l RNA folding Dynamic programming for RNA secondary structure prediction Covariance model for RNA structure prediction

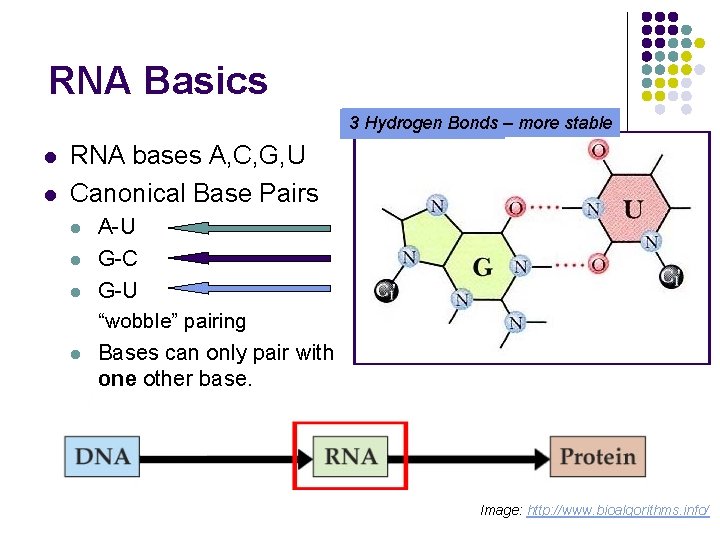
RNA Basics 23 Hydrogen Bonds – more stable l l RNA bases A, C, G, U Canonical Base Pairs l l A-U G-C G-U “wobble” pairing Bases can only pair with one other base. Image: http: //www. bioalgorithms. info/

RNA Basics l l l transfer RNA (t. RNA) messenger RNA (m. RNA) ribosomal RNA (r. RNA) small interfering RNA (si. RNA) micro RNA (mi. RNA) small nucleolar RNA (sno. RNA) http: //www. genetics. wustl. edu/eddy/t. RNAscan-SE/

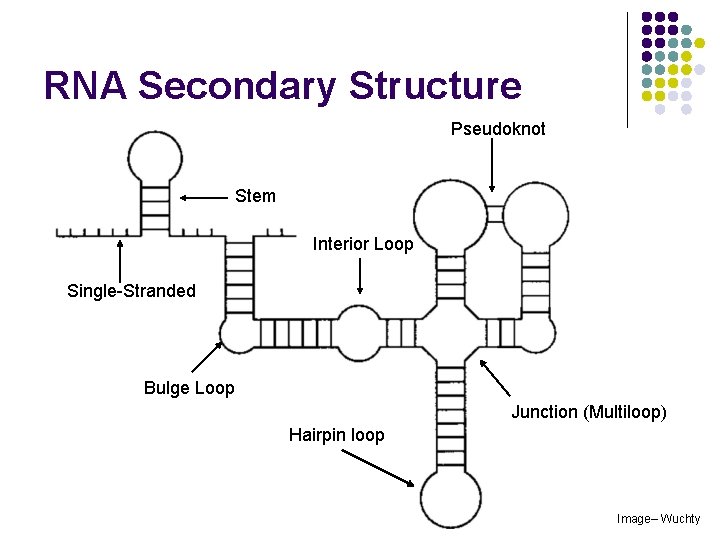
RNA Secondary Structure Pseudoknot Stem Interior Loop Single-Stranded Bulge Loop Junction (Multiloop) Hairpin loop Image– Wuchty

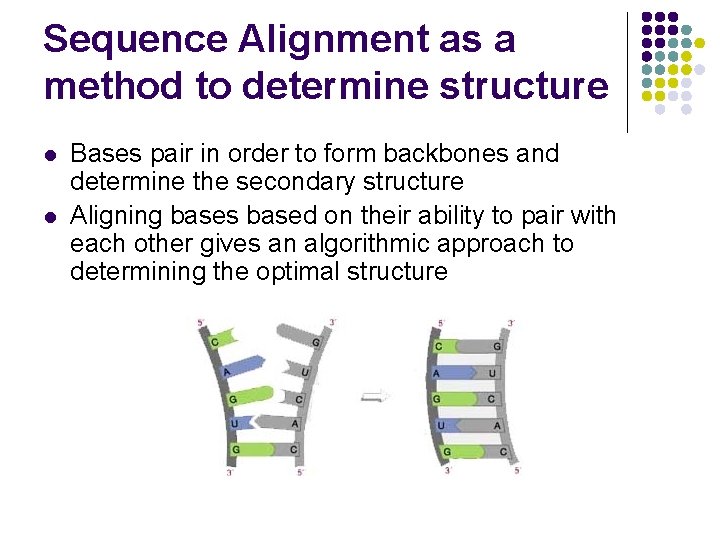
Sequence Alignment as a method to determine structure l l Bases pair in order to form backbones and determine the secondary structure Aligning bases based on their ability to pair with each other gives an algorithmic approach to determining the optimal structure

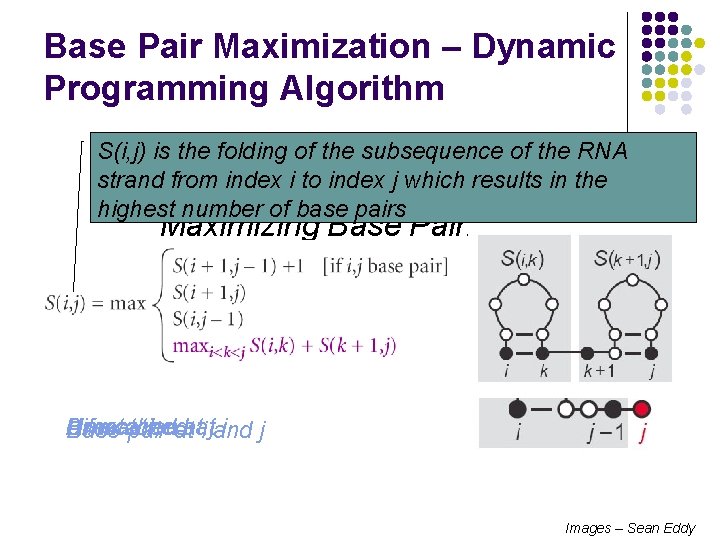
Base Pair Maximization – Dynamic Programming Algorithm S(i, j) is the folding of the subsequence of the RNA strand from. Simple index i to. Example: index j which results in the highest number of base pairs Maximizing Base Pairing Bifurcation Unmatched Umatched i j Base pair atatat i jand Images – Sean Eddy

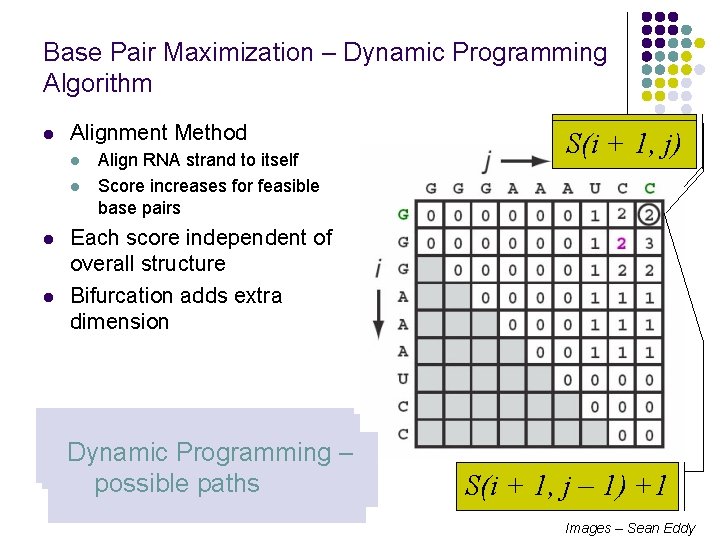
Base Pair Maximization – Dynamic Programming Algorithm l Alignment Method l l Align RNA strand to itself Score increases for feasible base pairs S(i, +j – 1, 1) S(i j) Each score independent of overall structure Bifurcation adds extra dimension Initialize first two diagonal Fill in squares sweeping Bases cannot pair, similar Bases can pair, similar Dynamic Programming – arrays to 0 diagonally to alignment to unmatched alignment possible paths S(i + 1, j – 1) +1 Images – Sean Eddy

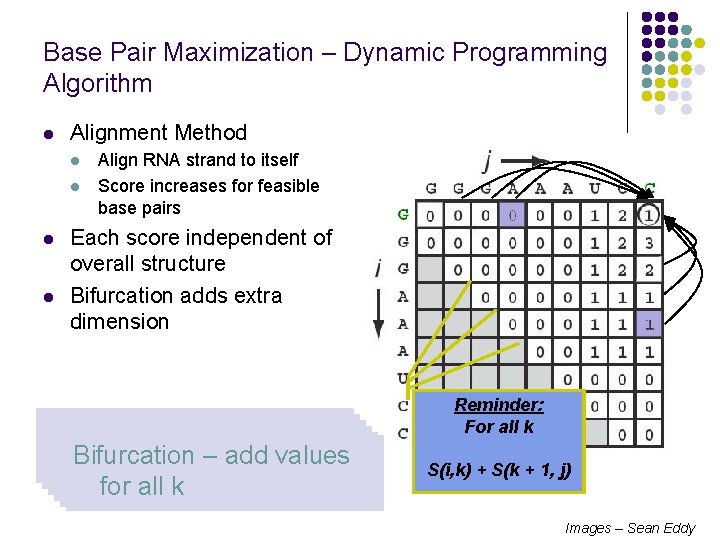
Base Pair Maximization – Dynamic Programming Algorithm l Alignment Method l l Align RNA strand to itself Score increases for feasible base pairs Each score independent of overall structure Bifurcation adds extra dimension Initialize first two diagonal Fill in squares sweeping Bases cannot pair, similar Bases can pair, similar Dynamic Bifurcation arrays to Programming 0 – add values– diagonally to matched alignment possible for all k paths k=0 Reminder: : Bifurcation max. For in this all kcase S(i, k) + S(k + 1, j) Images – Sean Eddy

Base Pair Maximization Drawbacks l Base pair maximization will not necessarily lead to the most stable structure l l May create structure with many interior loops or hairpins which are energetically unfavorable Comparable to aligning sequences with scattered matches – not biologically reasonable

Energy Minimization l Thermodynamic Stability l l l Estimated using experimental techniques Theory : Most Stable is the Most likely No Pseudknots due to algorithm limitations Uses Dynamic Programming alignment technique Attempts to maximize the score taking into account thermodynamics MFOLD and Vienna. RNA

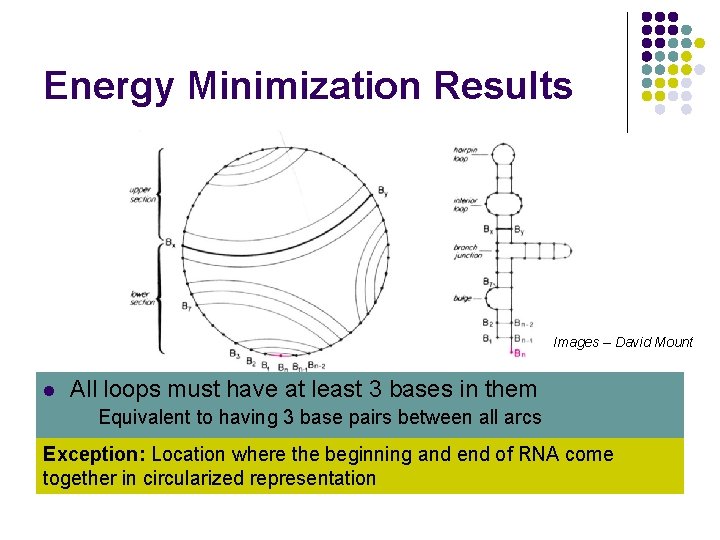
Energy Minimization Results Images – David Mount l l Linear RNA strand folded back 3 on itself to All loops must have at least bases in create them secondary structure l Equivalent to having 3 base pairs between all arcs Circularized representation uses this requirement Exception: Location where the beginning and end of RNA come l Arcs represent base pairing together in circularized representation

Trouble with Pseudoknots Images – David Mount l l Pseudoknots cause a breakdown in the Dynamic Programming Algorithm. In order to form a pseudoknot, checks must be made to ensure base is not already paired – this breaks down the recurrence relations

Energy Minimization Drawbacks l l Compute only one optimal structure Usual drawbacks of purely mathematical approaches l Similar difficulties in other algorithms l l Protein structure Exon finding

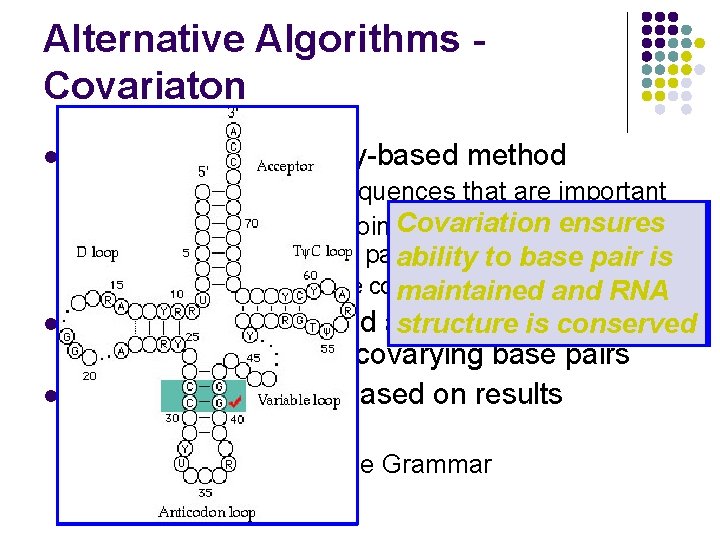
Alternative Algorithms Covariaton l Incorporates Similarity-based method l l Evolution maintains sequences that are important Base Mutation Covariation Expect areas in one ensures creates of base Change in sequence coincides topairing maintain structure through base pairs (Covariance) same yields ability pairing stable pairing toinbase t. RNA pair to be is l l l Cross-species structure conservation example –breaks t. RNA structure impossible maintained covarying inbetween and organisms RNA Manual and automated approaches down structure various structure species is have conserved been used to identify covarying base pairs Models for structure based on results l l Ordered Tree Model Stochastic Context Free Grammar

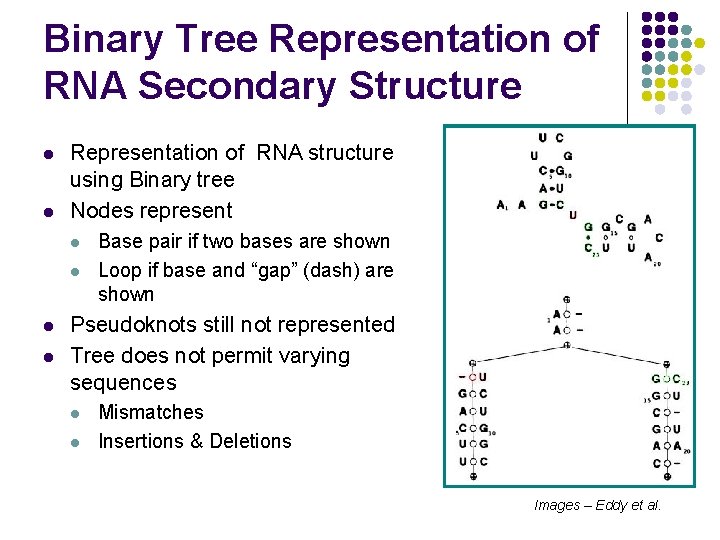
Binary Tree Representation of RNA Secondary Structure l l Representation of RNA structure using Binary tree Nodes represent l l Base pair if two bases are shown Loop if base and “gap” (dash) are shown Pseudoknots still not represented Tree does not permit varying sequences l l Mismatches Insertions & Deletions Images – Eddy et al.

Covariance Model l l HMM which permits flexible alignment to an RNA structure – l emission and transition probabilities Model trees based on finite number of states l Match states – sequence conforms to the model: l l l MATP – State in which bases are paired in the model and sequence MATL & MATR – State in which either right or left bulges in the sequence and the model Deletion – State in which there is deletion in the sequence when compared to the model Insertion – State in which there is an insertion relative to model Transitions have probabilities l Varying probability – Enter insertion, remain in current state, etc l Bifurcation – no probability, describes path

Covariance Model (CM) Training Algorithm l S(i, j) = Score at indices i and j in RNA when aligned to the Covariance Model Frequency of seeing the symbols Independent frequency of seeing the (A, C, G, T) together in locations i and j symbols (A, C, G, T) in locations i or j depending on symbol. l Frequencies obtained by aligning model to “training data” – consists of sample sequences l Reflect values which optimize alignment of sequences to model

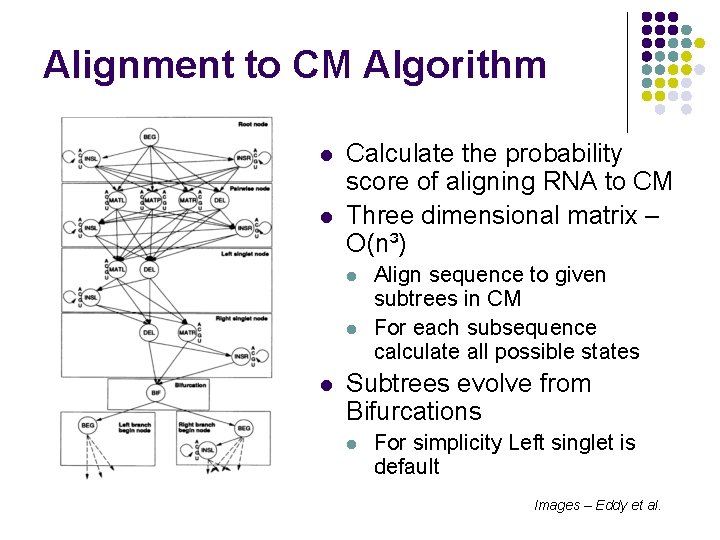
Alignment to CM Algorithm l l Calculate the probability score of aligning RNA to CM Three dimensional matrix – O(n³) l l l Align sequence to given subtrees in CM For each subsequence calculate all possible states Subtrees evolve from Bifurcations l For simplicity Left singlet is default Images – Eddy et al.

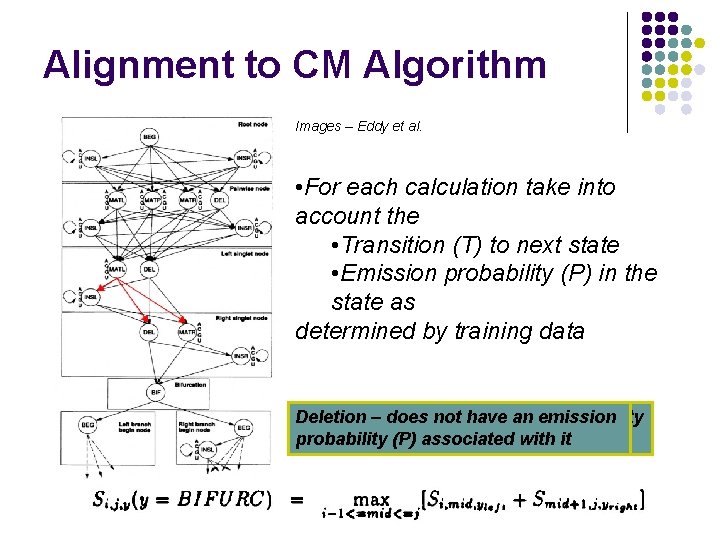
Alignment to CM Algorithm Images – Eddy et al. • For each calculation take into account the • Transition (T) to next state • Emission probability (P) in the state as determined by training data Deletion – does Bifurcation – does notnot have an aemission probability associated (P) probability withassociated the state with it

Covariance Model Drawbacks l l Needs to be well trained Not suitable for searches of large RNA l l l Structural complexity of large RNA cannot be modeled Runtime Memory requirements

References l How Do RNA Folding Algorithms Work? . S. R. Eddy. Nature Biotechnology, 22: 1457 -1458, 2004.