RNA Secondary Structure Prediction 16 s r RNA









- Slides: 16

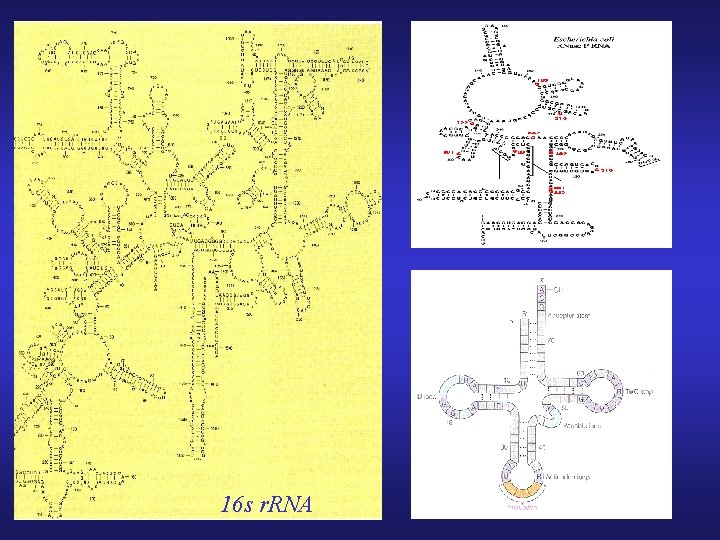
RNA Secondary Structure Prediction


16 s r. RNA

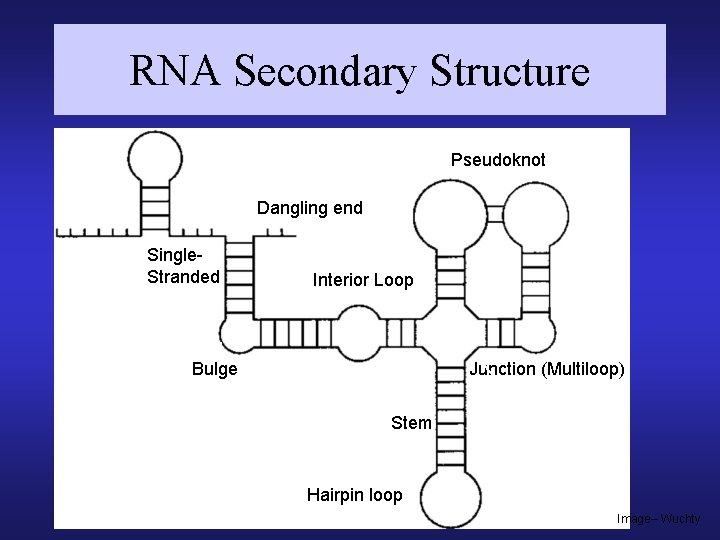
RNA Secondary Structure Pseudoknot Dangling end Single. Stranded Interior Loop Bulge Junction (Multiloop) Stem Hairpin loop Image– Wuchty

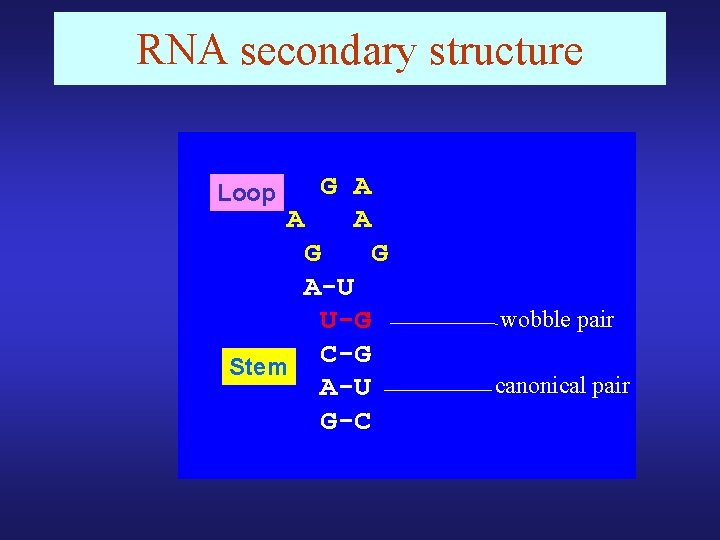
RNA secondary structure G A A A G G A-U U-G C-G Stem A-U G-C Loop wobble pair canonical pair

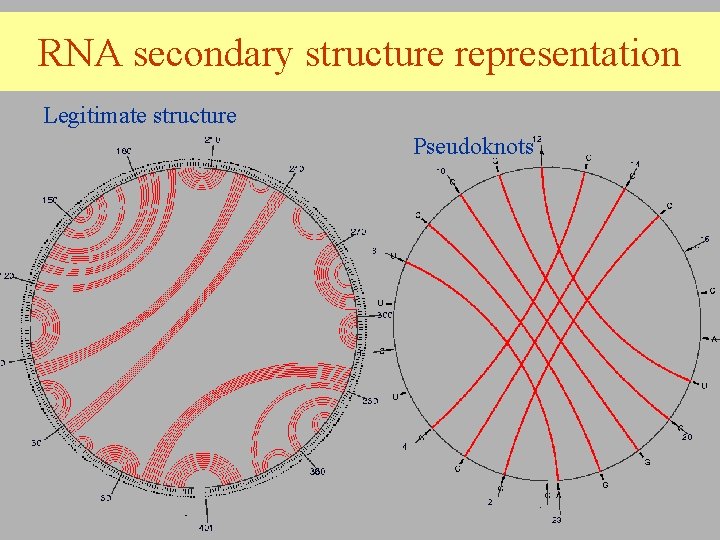
RNA secondary structure representation Legitimate structure Pseudoknots

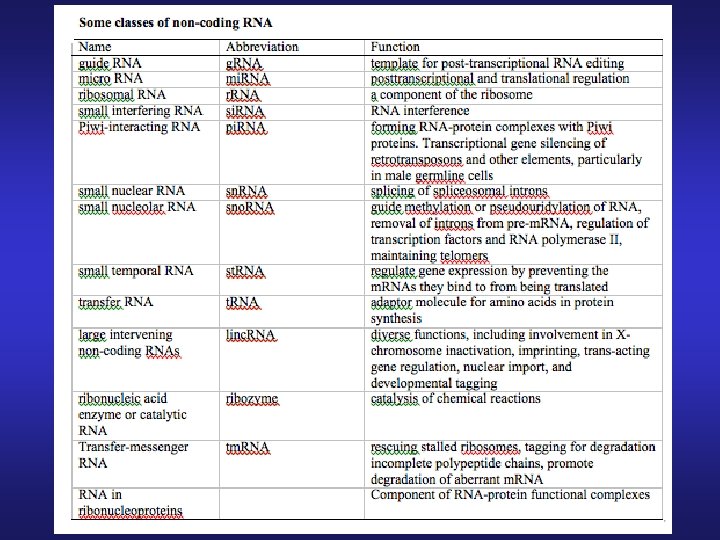
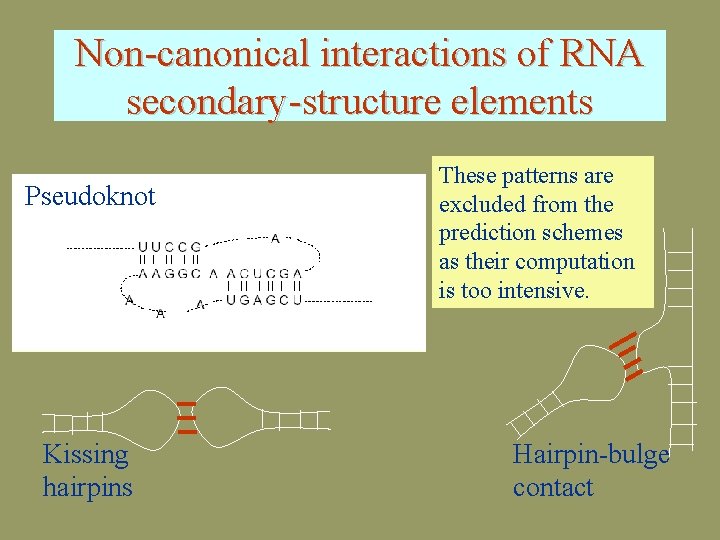
Non-canonical interactions of RNA secondary-structure elements Pseudoknot Kissing hairpins These patterns are excluded from the prediction schemes as their computation is too intensive. Hairpin-bulge contact

“Rules for 2 D RNA prediction” • Base Pairs in stems: GOOD • Additional possible assumptions: e. g. , G: C better than A: T • Bulges, Loops: BAD • Canonical Interactions (base pairs, stems, bulges, loops): OK • Non canonical interactions (pseudoknots, kissing hairpins): Forbidden • The more interactions: The better

Predicting RNA secondary Structure • Allowed base pairing rules (Watson-Crick A: U, G: C, and Wobble pair G: U) • Sequences may form different structures • An free energy value is associated with each possible structure • Predict the structure with the minimal free energy (MFE)

Simplifying Assumptions for Structure Prediction • RNA folds into one minimum free-energy structure. • There are no non-canonical interactions. • The energy of a particular base pair in a double stranded regions is sequence independent – Neighbors have no influence. Was solved by dynamic programming Zucker and Steigler 1981

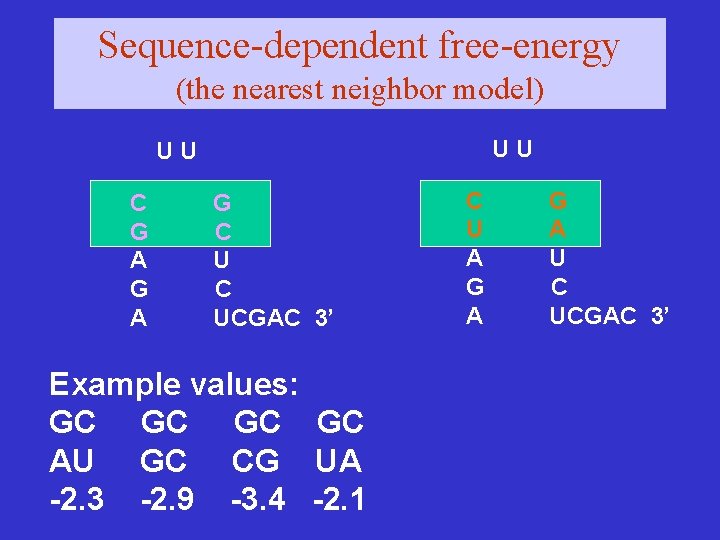
Sequence-dependent free-energy (the nearest neighbor model) UU UU C G A G C UCGAC 3’ Example values: GC GC AU GC CG UA -2. 3 -2. 9 -3. 4 -2. 1 C U A G A U C UCGAC 3’

Free energy computation +5. 9 (4 nt loop) U U A G C +3. 3 (1 nt bulge) G U A C A A A 5’ -1. 1 mismatch of hairpin -2. 9 stacking A 5’ dangling -0. 3 A C A U G U -1. 8 stacking -0. 9 stacking -1. 8 stacking -2. 1 stacking 3’ G= -4. 6 KCAL/MOL

Prediction Programs • Mfold http: //www. bioinfo. rpi. edu/applications/mfold/rna/form 1. cgi • Vienna RNA Secondary Structure Prediction http: //rna. tbi. univie. ac. at/cgi-bin/RNAfold. cgi

Mfold - Suboptimal Folding • For any sequence of N nucleotides, the expected number of structures is greater than 1. 8 N • A sequence of 100 nucleotides has ~3 1025 possible folds. If a computer can calculate 1000 folds/second, it would take 1015 years (age of universe = ~1010 years)! • Mfold generates suboptimal folds whose free energy fall within a certain range of values. Many of these structures are different in trivial ways. These suboptimal folds can still be useful for designing experiments.

Example:

Output: