RNA Structure Prediction RNA Structure Basics The RNA

RNA Structure Prediction • RNA Structure Basics • The RNA ‘Rules’ • Programs and Predictions Assigned reading: Ch. 6 from Bioinformatics: A Practical Guide to the Analysis of Genes and Proteins, 3 rd Ed. by Baxevanis and Ouellette. BIO 520 Bioinformatics Jim Lund

RNA classes • m. RNA - messenger RNA. • t. RNA - transfer RNA, small (~80 bases) sequences which bring amino acids to the ribosome. • r. RNA - ribosomal RNA, RNA + proteins = ribosome. • viral RNA (ss. RNA, ds. RNA virii) • mi. RNA: translational/transcriptional gene silencing. • sno. RNA, sn. RNA: splicing, RNA bp modification • Transfer-Messenger RNAs (tm. RNA), Small cytosolic RNAs (sc. RNA), Guide RNAs (g. RNA) • and more…

RNA structures • 1° – Sequence (and modifications) • 2° – Base pairing • 3° – Overall Structure, non Watson-Crick pairs – Experimental structures: t. RNA, ribosome
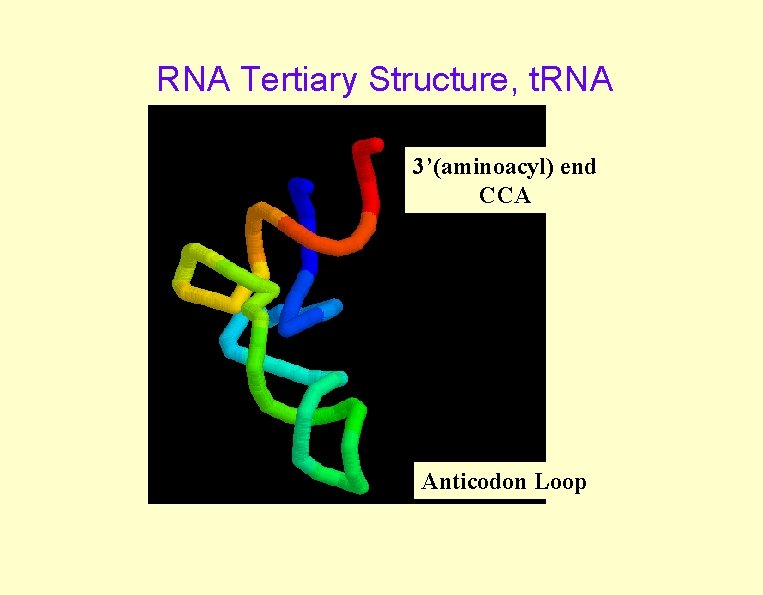
RNA Tertiary Structure, t. RNA 3’(aminoacyl) end CCA Anticodon Loop
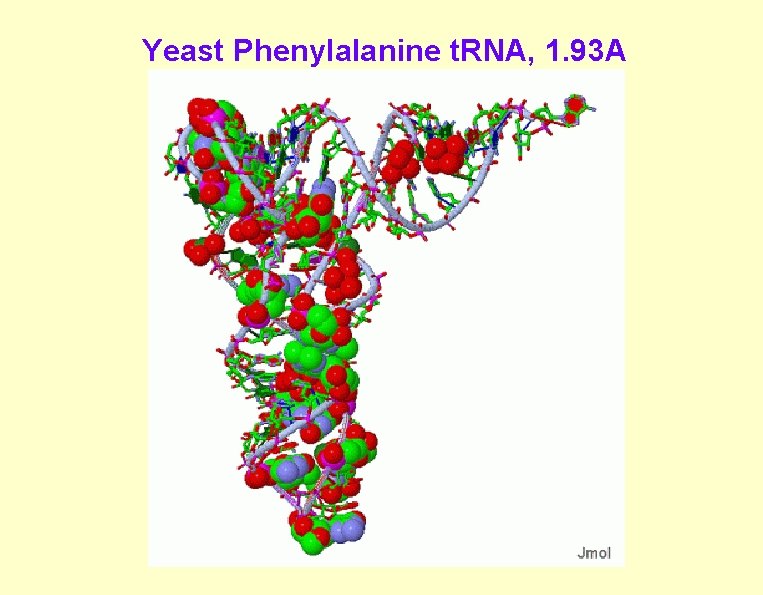
Yeast Phenylalanine t. RNA, 1. 93 A
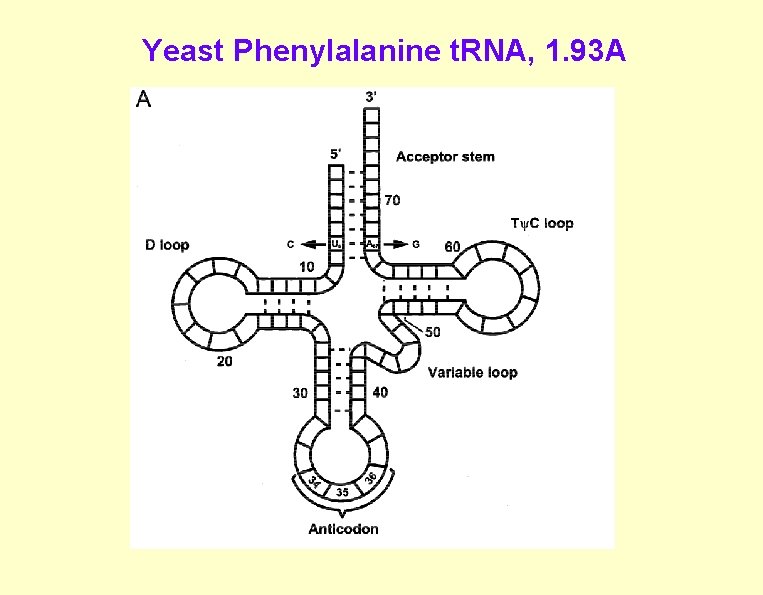
Yeast Phenylalanine t. RNA, 1. 93 A
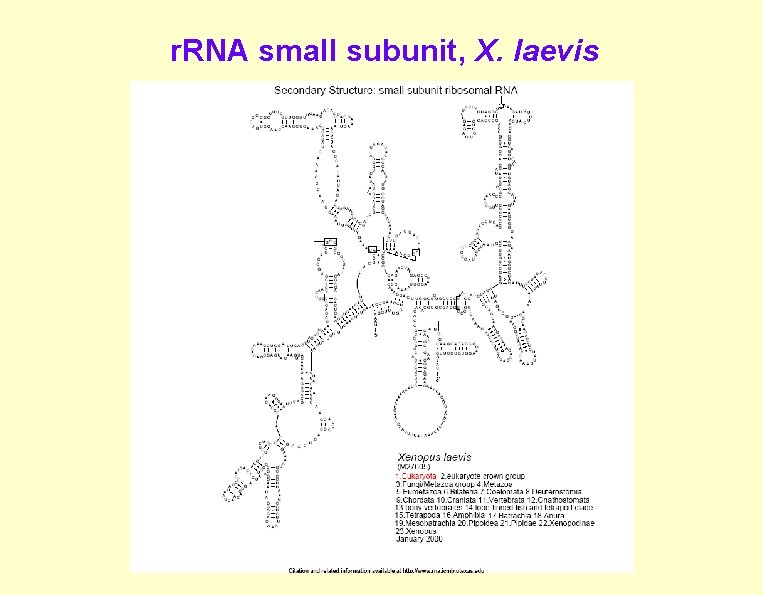
r. RNA small subunit, X. laevis

2° RNA structures • Watson-Crick pairing -> helices • Loop regions – Hairpin loops – Internal loops – Bulge loops – Multibranch loops

RNA Modifications Covalent Modifications-especially t. RNA – r. U r. T, r , r. D r. S 4 U – r. C 3 -CH 3 -C, 5 -CH 3 -C – r. A I, 6 -CH 3 -A, 6 -isopentenyl-A – r. G 7 -CH 3 -G, Q, Y Nucleosides Nucleotides 1999 Jun-Jul; 18(6 -7): 1579 -81

RNA Base pairing • G-C triple hydrogen bond • A -U double hydrogen bond • G-U single hydrogen bond

RNA structure energetics • The number of GC versus AU and GU base pairs. – Higher energy bonds form more stable structures. • Number of base pairs in a stem region. – Longer stems result in more bonds. • Number of base pairs in a hairpin loop region. – Formation of loops with more than 10 or less than 5 bases requires more energy. • Number of unpaired bases (interior loops or bulges). – Unpaired bases decrease the stability of the structure.
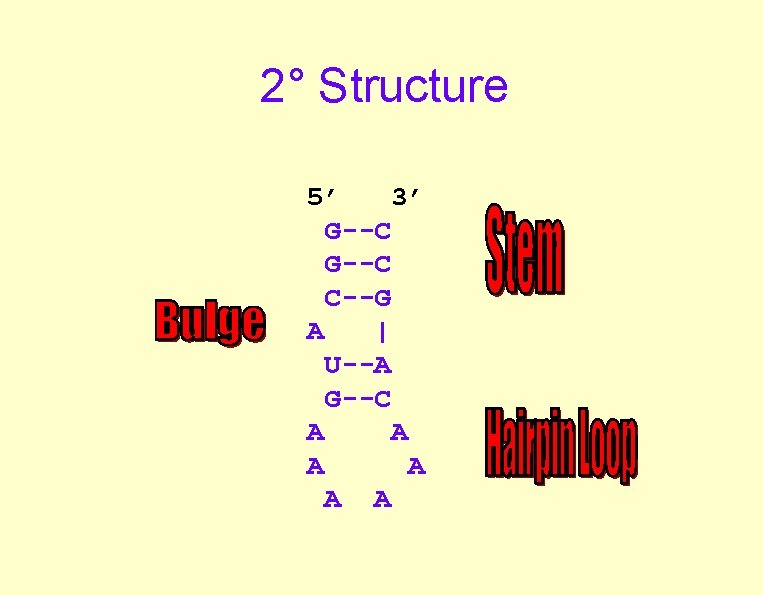
2° Structure 5’ 3’ G--C C--G A | U--A G--C A A A

“The Rules” • Base Pairs -- Good – G: C better than A: T -- And local sequence matters! • Bulges, Loops -- Bad • Many small interactions---Stable Structure • Only predict “Canonical Interactions”
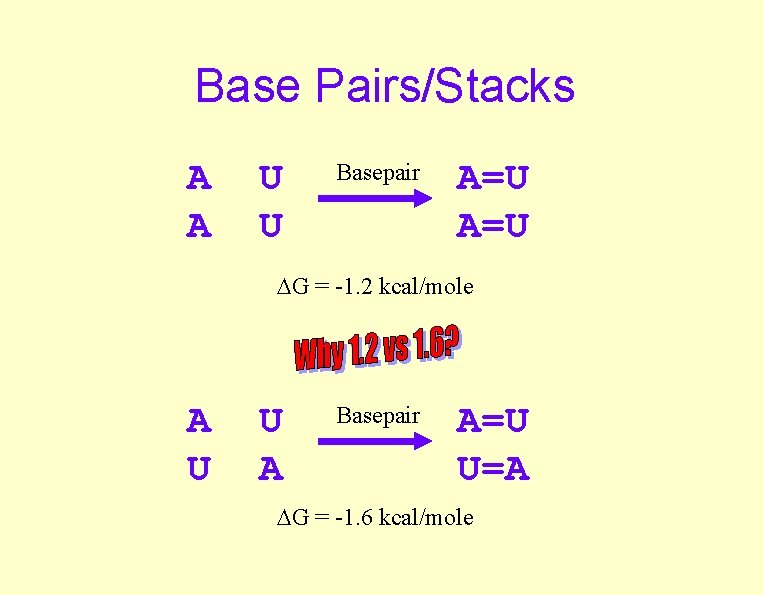
Base Pairs/Stacks A A U U Basepair A=U G = -1. 2 kcal/mole A U U A Basepair A=U U=A G = -1. 6 kcal/mole

Base Pairing/Stacking Bloomfield, Crothers, Tinoco, Physical Chemistry of Nucleic Acids

Hairpin Loops (GC closure) • Tertiary Interactions!

Internal Loops 5’ 3’ G--C C--G A G G A A C T--A G--C

Single-Strand Bulges 5’ 3’ G--C C--G A | G | A | T--A G--C

Prediction Programs • Mfold (M. Zuker) – 2° structure • RNAstructure/Oligo. Walk – 2° structure, oligo/RNA target interactions • alifold – 2° structure constrained by muliple alignment. • Pfold – 2° structure guided by rules derived from known t. RNA/r. RNA structures

Prediction Programs • Mfold (GCG) – M. Zuker • Mfold input to Plotfold – Non-graphic output -G option – Graphics outputs • • • SQUIGGLES mountains circles domes energy plots
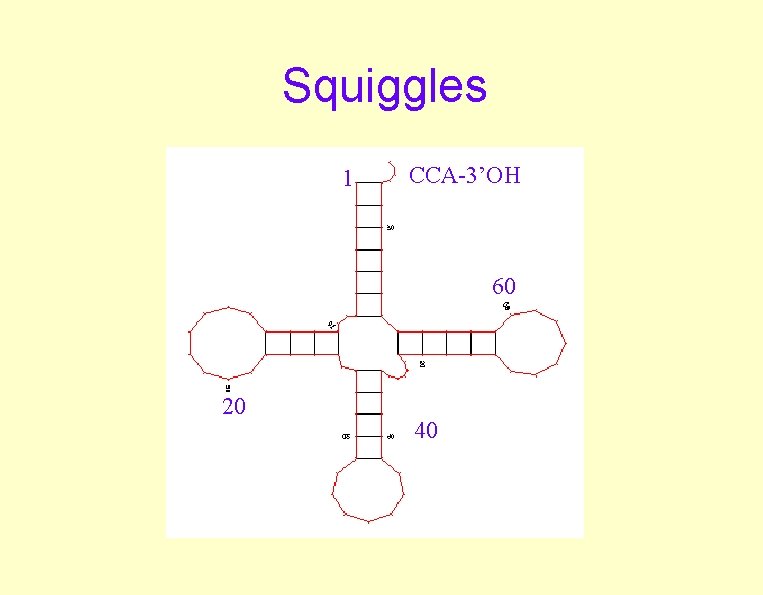
Squiggles 1 CCA-3’OH 60 20 40
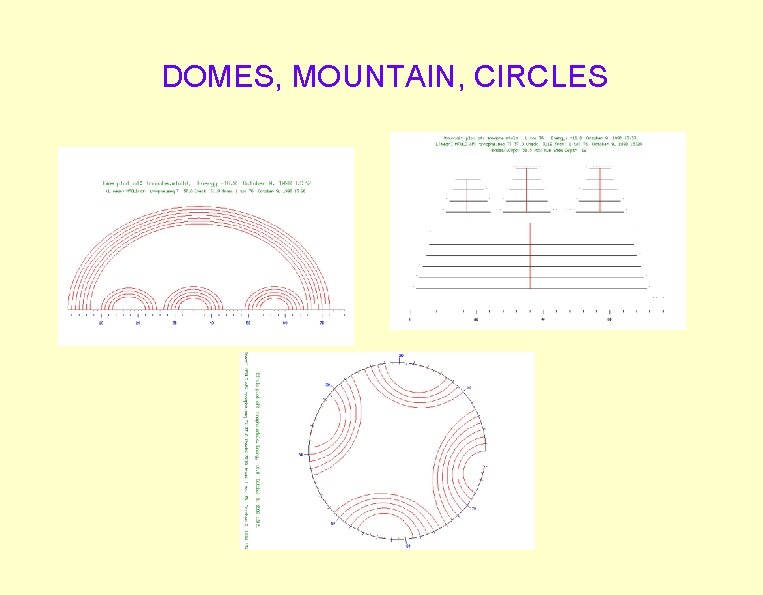
DOMES, MOUNTAIN, CIRCLES

MFOLD Structure Family • Optimal & Suboptimal structures – Can ask for multiple structures • Energy increment and “window size” increment. • View individually. • How variable are the structures? – Energy Plots

ENERGY PLOT
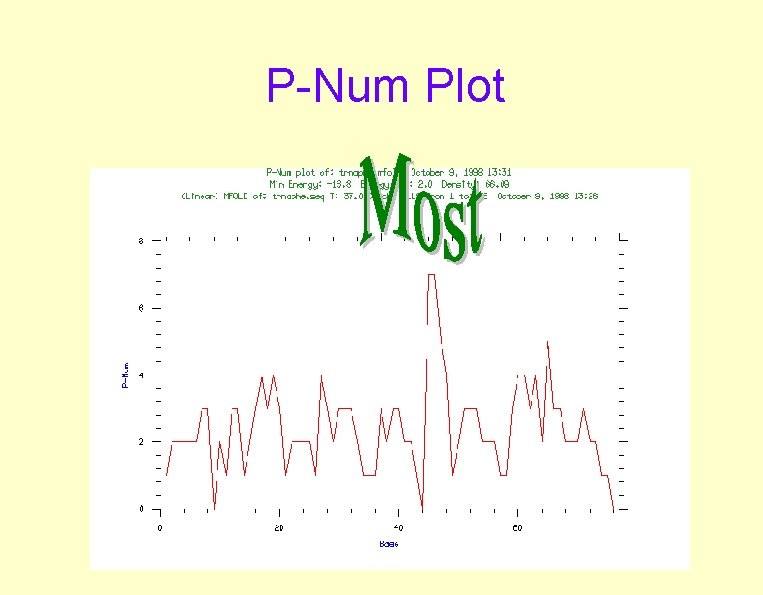
P-Num Plot
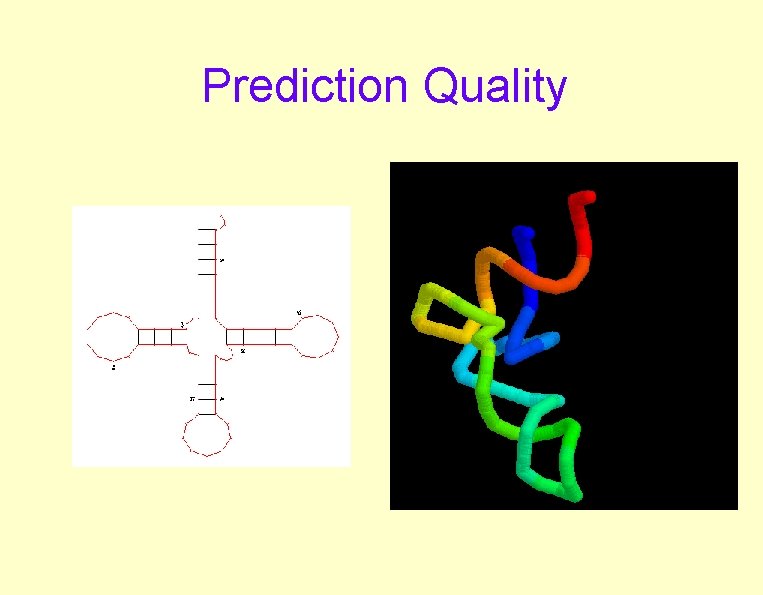
Prediction Quality

Forces in RNA folds • Complementary molecular surfaces • Bridging cations • Pseudoknotting • “kinetic traps” in folding – NOT always 2 first! Annu Rev Biophys Biomol Struct 1999; 28: 57 -73 Proc Natl Acad Sci U S A 1998 Sep 29; 95(20): 11555 -60
RNA Structure Probing • Physical methods – X-ray diffraction, NMR • Enzymatic methods – S 1, Rnases (find ss and ds regions). • Chemical modification – DMS… • Mutagenesis – G: C=>C: G

Ribozymes • Naturally occurring – – RNAase. P Group I introns Group II introns sn. RNA in the splicosome • Artifical – Engineered/evolved in the lab from natural ribozymes to have new substrate RNA. – Cleave m. RNA, drug-like action • mi. RNA/si. RNA – Translational/transcriptional gene silencing
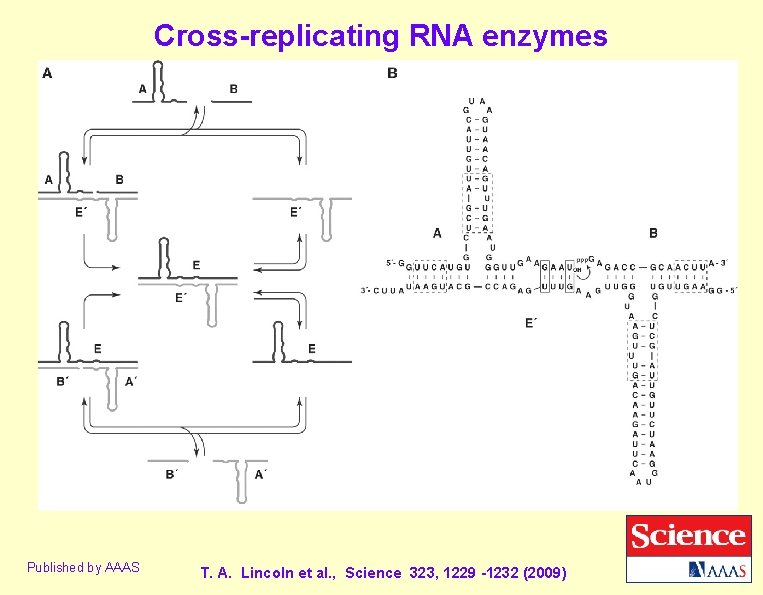
Cross-replicating RNA enzymes Published by AAAS T. A. Lincoln et al. , Science 323, 1229 -1232 (2009)
- Slides: 30