ReninAngiotensin System Genes and their Associations with Renal


















- Slides: 34

Renin-Angiotensin System Genes and their Associations with Renal Function in MESA Preliminary Analyses Catherine Campbell, M. D. Wendy Post, M. D. , M. S. Johns Hopkins University Co-authors: Joe Coresh, Hunter Young, Michael Shlipak, Carmen Peralta, Bruce Psaty, Stephen Rich, Jerome Rotter, Xiuqing Guo, Belle Fang, Holly Kramer

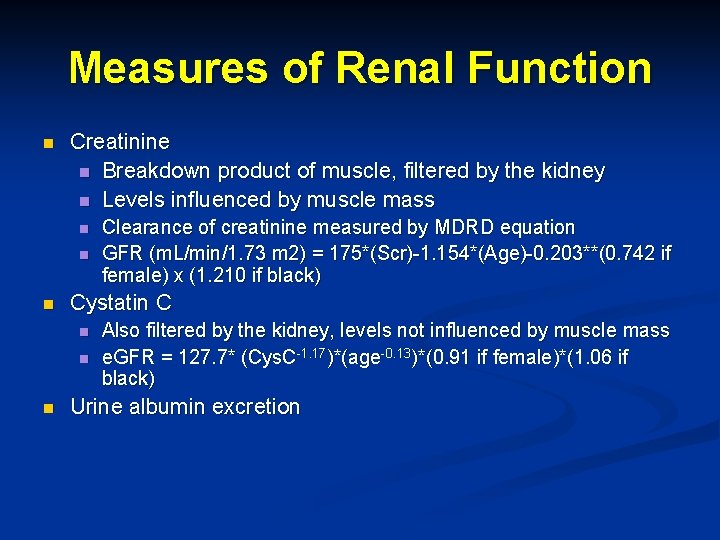
Measures of Renal Function n Creatinine n Breakdown product of muscle, filtered by the kidney n Levels influenced by muscle mass n n n Cystatin C n n n Clearance of creatinine measured by MDRD equation GFR (m. L/min/1. 73 m 2) = 175*(Scr)-1. 154*(Age)-0. 203**(0. 742 if female) x (1. 210 if black) Also filtered by the kidney, levels not influenced by muscle mass e. GFR = 127. 7* (Cys. C-1. 17)*(age-0. 13)*(0. 91 if female)*(1. 06 if black) Urine albumin excretion

Chronic Kidney Disease Progressive decline in renal function, rarely reversible n Relatively common disorder: affects about one in nine adults n Over 50% of late-stage chronic kidney disease is due to hypertension or diabetes n African Americans and Hispanics have higher rates of chronic kidney disease than whites n

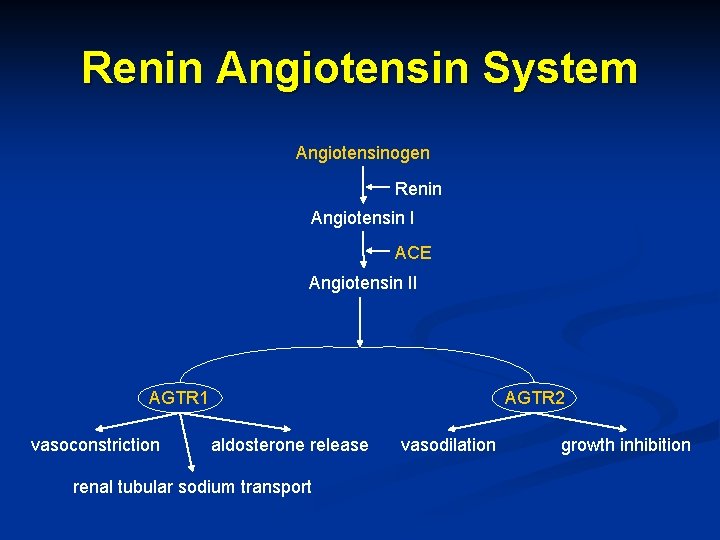
Renin Angiotensin System Angiotensinogen Renin Angiotensin I ACE Angiotensin II AGTR 1 vasoconstriction AGTR 2 aldosterone release renal tubular sodium transport vasodilation growth inhibition

Purpose n Do polymorphisms in renin-angiotensin system genes contribute to variation in renal function seen in the general population? n n angiotensin converting enzyme (ACE) angiotensinogen (AGT) angiotensin II receptor type 1 (AGTR 1) angiotensin II receptor type 2 (AGTR 2)

Methods n ACE, AGTR 1, and AGTR 2 are candidate genes selected for genotyping in 2880 unrelated MESA participants (720 in each ethnic group) using Illumina n SNPs were selected to sample as much of the intrinsic multi-ethnic genetic variation in these genes as possible n MAF, D prime, HWE calculated using automated methods

Methods n n n Associations between each SNP and measures of renal function determined using multivariate regression, adjusting for age and gender, stratified by race/ethnicity and combined Additional models also adjusted for hypertension and diabetes SNP associations n n n Additive model used if the MAF >10% Dominant model used if MAF <10% Haplotype associations n Dominant and recessive models

Methods: Primary outcomes n Estimated GFR (using MDRD based on serum creatinine) n n n continuous variable dichotomous variable (<60 ml/min/m 2) Urine albumin excretion (UAE) albumin/creatinine n n continuous variable (log transformed) dichotomous variable n n > 17 mg/g for men > 25 mg/g for women

Methods: Secondary outcomes n Estimated GFR with cystatin equation n continuous variable n dichotomous variable (<60 ml/min/m 2) n Cystatin C n continuous variable (log-transformed) n dichotomous variable (>1 mg/L) (>1 mg/L

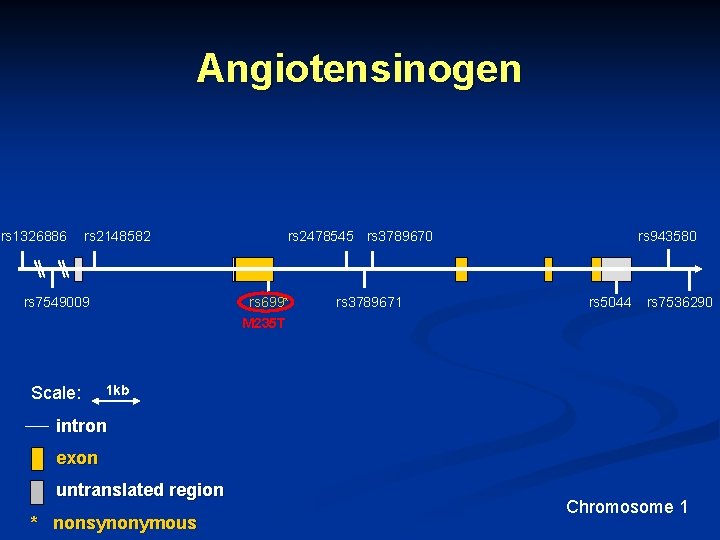
Angiotensinogen rs 1326886 rs 2148582 rs 7549009 Scale: rs 2478545 rs 3789670 rs 699* M 235 T rs 3789671 rs 943580 rs 5044 rs 7536290 1 kb intron exon untranslated region * nonsynonymous Chromosome 1

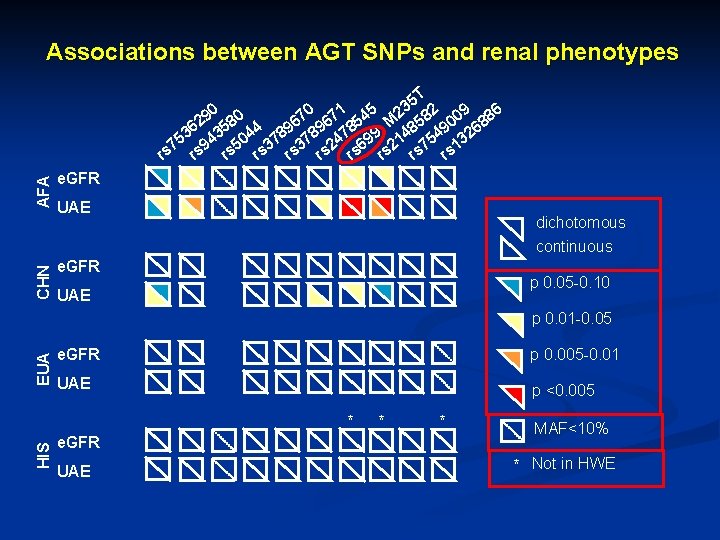
Associations between AGT SNPs and renal phenotypes CHN AFA 5 T 3 0 1 5 9 80 70 67 54 M 2 582 009 886 2 6 36 435 044 789 478 99 148 549 326 5 7 9 3 2 6 5 2 7 1 3 rs rs rs e. GFR UAE dichotomous continuous e. GFR p 0. 05 -0. 10 UAE EUA p 0. 01 -0. 05 e. GFR p 0. 005 -0. 01 UAE p <0. 005 HIS * e. GFR UAE * * MAF<10% * Not in HWE

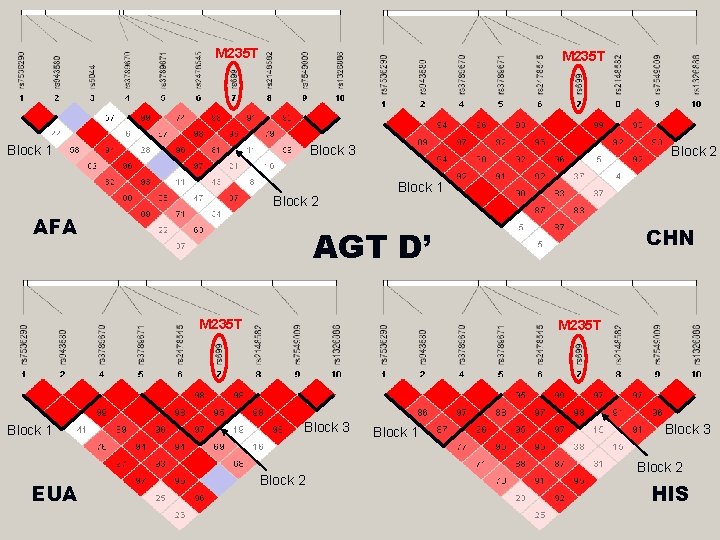
M 235 T Block 1 Block 3 Block 2 AFA Block 2 Block 1 M 235 T Block 1 EUA CHN AGT D’ M 235 T Block 3 Block 2 Block 1 Block 3 Block 2 HIS

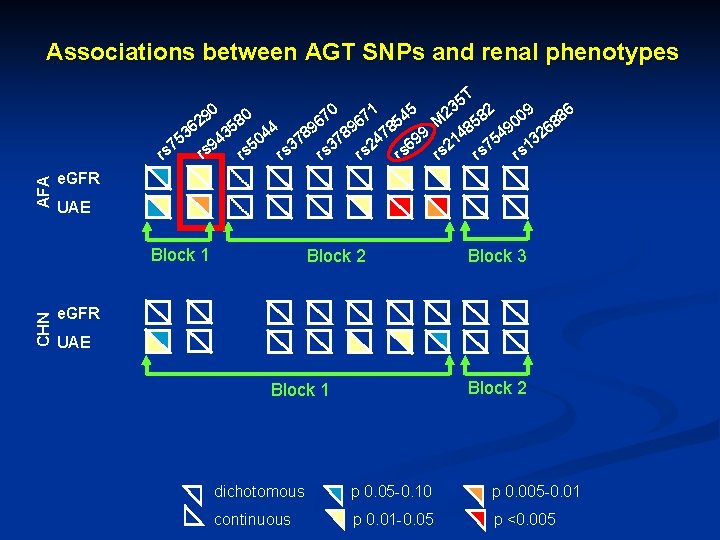
Associations between AGT SNPs and renal phenotypes 5 T 3 1 5 2 0 4 70 67 82 009 886 8 2 5 M 6 5 36 435 044 789 478 99 48 549 326 5 1 9 7 3 2 6 5 3 2 7 1 rs rs rs AFA 90 e. GFR UAE CHN Block 1 Block 2 Block 3 e. GFR UAE Block 2 Block 1 dichotomous p 0. 05 -0. 10 p 0. 005 -0. 01 continuous p 0. 01 -0. 05 p <0. 005

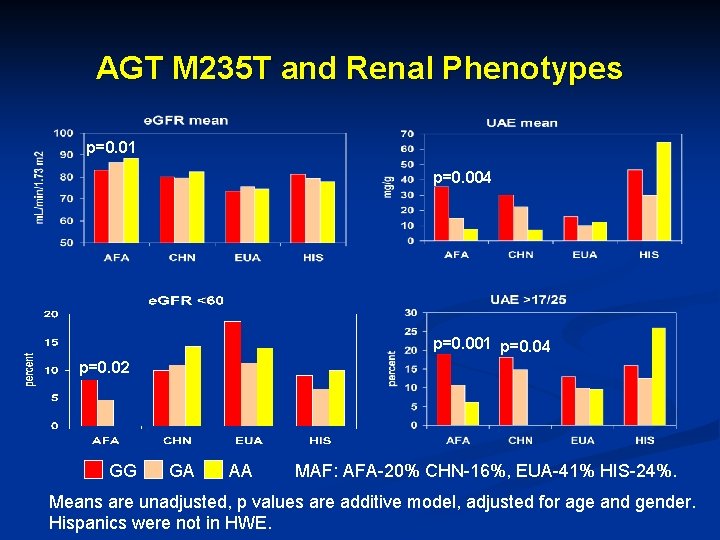
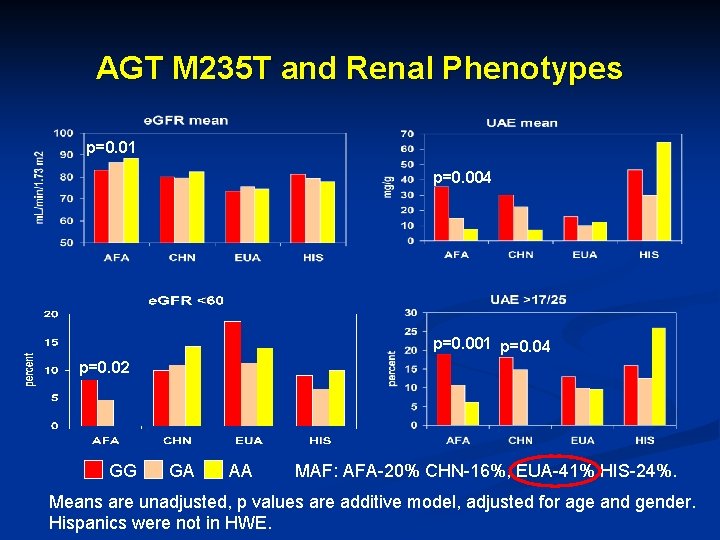
AGT M 235 T and Renal Phenotypes p=0. 01 p=0. 004 p=0. 001 p=0. 04 p=0. 02 GG GA AA MAF: AFA-20% CHN-16%, EUA-41% HIS-24%. Means are unadjusted, p values are additive model, adjusted for age and gender. Hispanics were not in HWE.

Haplotypes 0 29 80 5 3 6 rs AFA Block 2 CHN Block 1 3 75 4 s 9 1 2 3 4 5 A G A A A 7 s 3 r 1 2 3 4 5 6 7 G G G A G 71 6 89 70 6 89 7 s 3 r r 7 45 5 8 r 4 s 2 T 5 23 9 69 s r M 4 82 5 8 21 s r G A G G C A A C C A G G G G A G G A C C G A G G A G

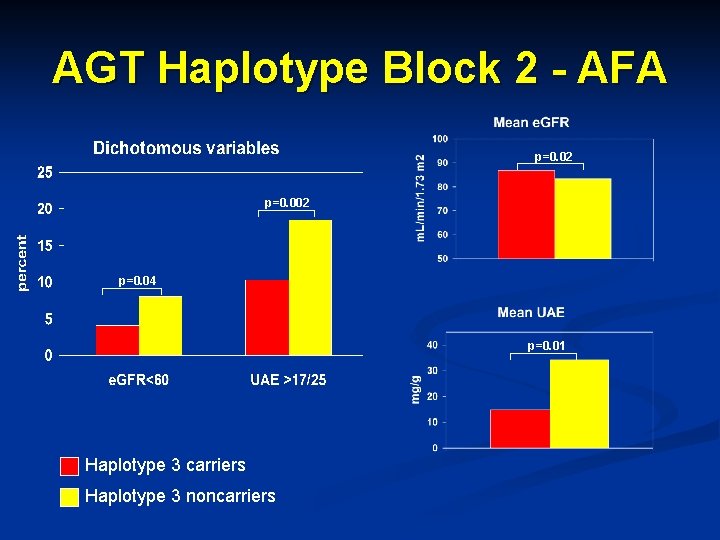
AGT Haplotype Block 2 - AFA p=0. 02 p=0. 04 p=0. 01 Haplotype 3 carriers Haplotype 3 noncarriers

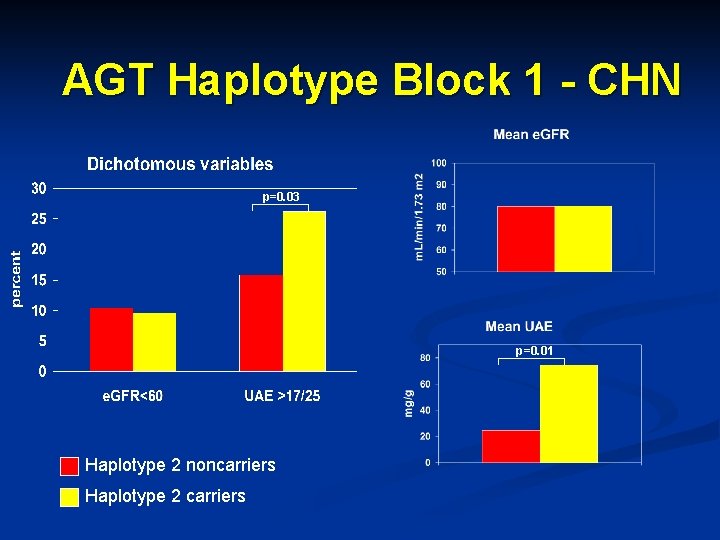
AGT Haplotype Block 1 - CHN p=0. 03 p=0. 01 Haplotype 2 noncarriers Haplotype 2 carriers

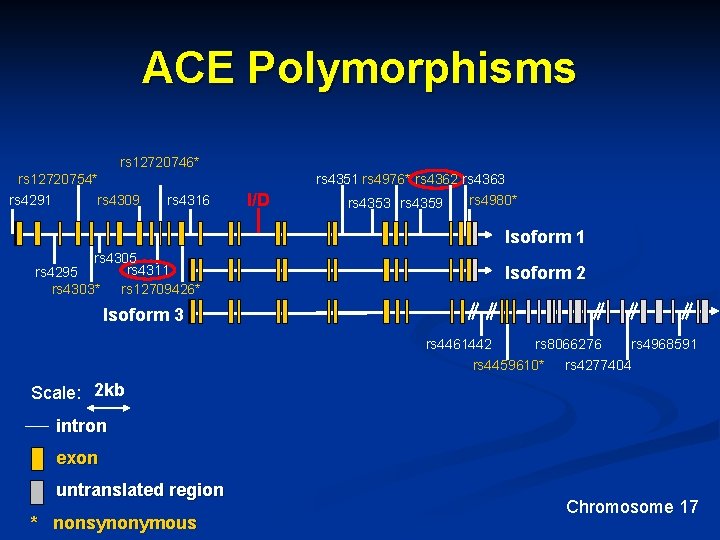
ACE Polymorphisms rs 12720746* rs 12720754* rs 4291 rs 4351 rs 4976* rs 4362 rs 4363 rs 4309 rs 4316 I/D rs 4353 rs 4359 rs 4980* Isoform 1 rs 4305 rs 4311 rs 4295 rs 4303* rs 12709426* Isoform 2 Isoform 3 rs 4461442 rs 8066276 rs 4968591 rs 4459610* rs 4277404 Scale: 2 kb intron exon untranslated region * nonsynonymous Chromosome 17

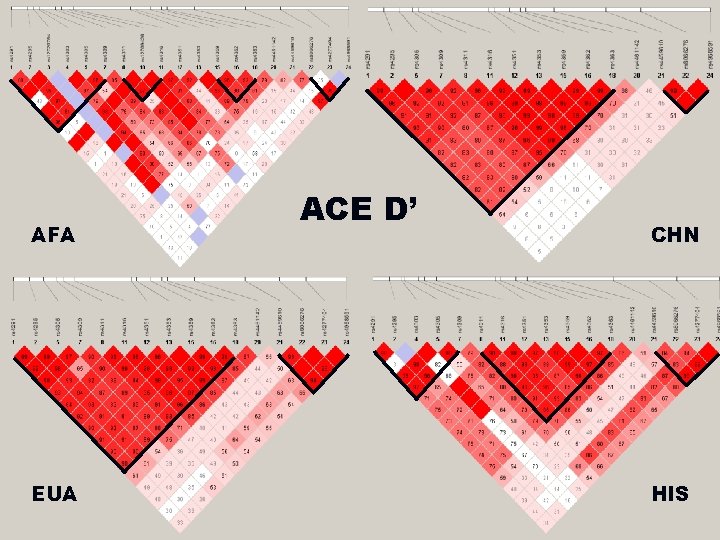
AFA EUA ACE D’ CHN HIS

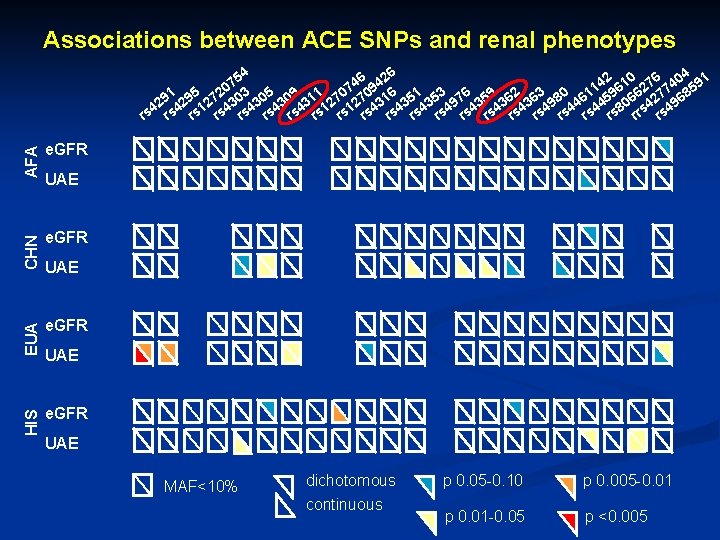
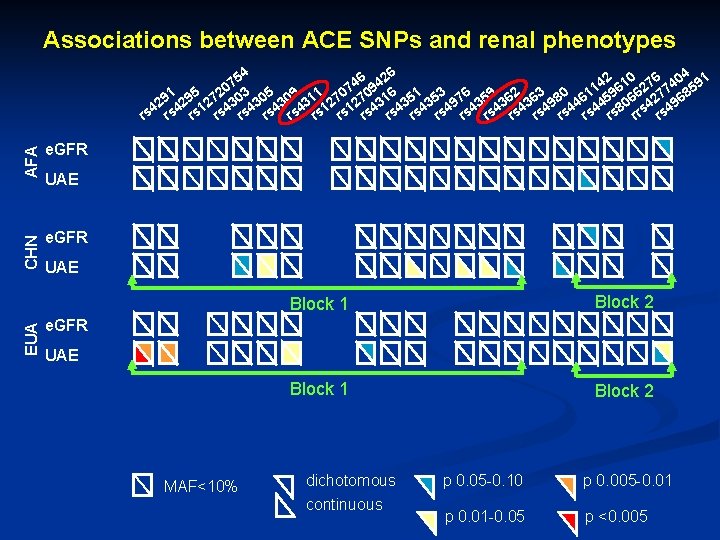
Associations between ACE SNPs and renal phenotypes AFA e. GFR CHN e. GFR EUA e. GFR HIS 6 426 54 2 10 76 404 91 4 7 9 0 1 6 2 7 5 91 295 272 303 305 309 311 270 316 351 353 976 359 362 363 980 461 459 066 427 968 2 4 4 1 1 4 4 4 4 4 8 s 4 rs rs rs rs rs rr rs e. GFR UAE UAE MAF<10% dichotomous continuous p 0. 05 -0. 10 p 0. 005 -0. 01 p 0. 01 -0. 05 p <0. 005

Associations between ACE SNPs and renal phenotypes AFA e. GFR EUA e. GFR CHN 6 426 54 2 10 76 404 91 4 7 9 0 1 6 2 7 5 91 295 272 303 305 309 311 270 316 351 353 976 359 362 363 980 461 459 066 427 968 2 4 4 1 1 4 4 4 4 4 8 s 4 rs rs rs rs rs rr rs UAE Block 1 Block 2 e. GFR UAE MAF<10% dichotomous continuous p 0. 05 -0. 10 p 0. 005 -0. 01 p 0. 01 -0. 05 p <0. 005

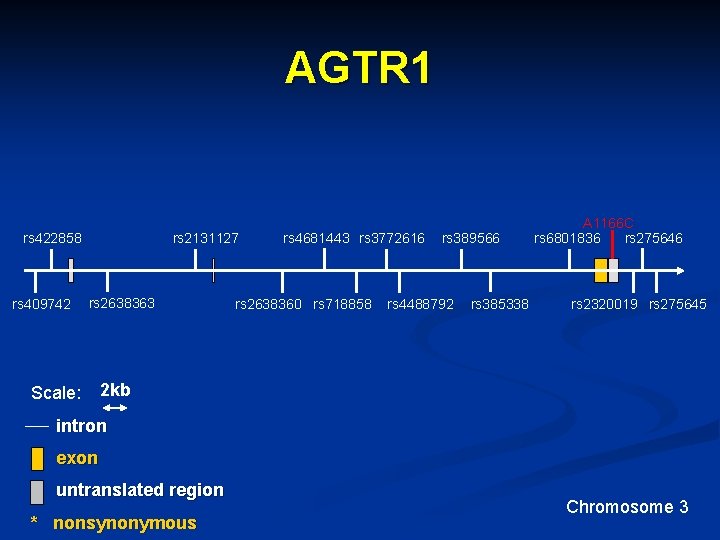
AGTR 1 rs 2131127 rs 422858 rs 409742 rs 2638363 Scale: rs 4681443 rs 3772616 rs 2638360 rs 718858 rs 389566 rs 4488792 rs 385338 A 1166 C rs 6801836 rs 275646 rs 2320019 rs 275645 2 kb intron exon untranslated region * nonsynonymous Chromosome 3

Associations between AGTR 1 SNPs and Renal Phenotypes AFA e. GFR CHN e. GFR EUA 2 58 363 127 360 443 58 616 792 56 38 836 019 46 45 4 97 228 638 131 638 681 188 772 488 895 853 801 320 756 0 2 4 7 3 4 4 4 3 3 6 2 2 2 rs rs rs rs e. GFR UAE UAE HIS * e. GFR UAE MAF<10% * Not in HWE dichotomous p 0. 05 -0. 10 p 0. 005 -0. 01 continuous p 0. 01 -0. 05 p <0. 005

AGTR 2 Polymorphisms rs 5950584 rs 1403543 rs 5950586 Scale: rs 5191* rs 5190 rs 6608590 1 kb intron exon untranslated region * nonsynonymous X Chromosome

Associations between AGTR 2 SNPs and Renal Phenotypes AFA 90 84 586 543 5 5 50 950 403 190 191 608 9 1 5 5 s 6 5 5 r rs rs rs e. GFR dichotomous continuous UAE CHN p 0. 05 -0. 10 e. GFR p 0. 01 -0. 05 UAE HIS EUA p 0. 005 -0. 01 e. GFR p <0. 005 UAE MAF<10% e. GFR UAE * * Not in HWE

Conclusions Many SNPs in the renin-angiotensin system show signficant associations with renal phenotypes n AGT M 235 T is associated with e. GFR and UAE in African Americans and with UAE as a dichotomous variable in Chinese n The haplotype blocks that include this polymorphism also show significant associations with renal phenotypes. n

Conclusions ACE: Many SNPs in ACE Haplotype Block 1 in Caucasians and Chinese show significant associations n AGTR 1: Significant associations seen in African Americans and Hispanics n AGTR 2: Many SNPs are not polymorphic or have low MAF n

To Do Permutation testing n Haplotype analyses n Interaction tests with diabetes and hypertension n AIMs n

Topics for Discussion HWE in Hispanics n Appropriate genetic model n Interpretation of haplotype blocks n

AGT M 235 T and Renal Phenotypes p=0. 01 p=0. 004 p=0. 001 p=0. 04 p=0. 02 GG GA AA MAF: AFA-20% CHN-16%, EUA-41% HIS-24%. Means are unadjusted, p values are additive model, adjusted for age and gender. Hispanics were not in HWE.

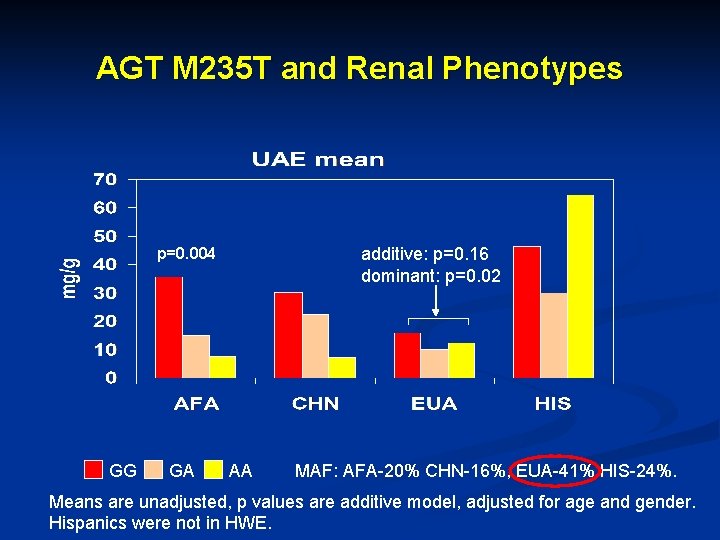
AGT M 235 T and Renal Phenotypes additive: p=0. 16 dominant: p=0. 02 p=0. 004 GG GA AA MAF: AFA-20% CHN-16%, EUA-41% HIS-24%. Means are unadjusted, p values are additive model, adjusted for age and gender. Hispanics were not in HWE.

Topics for Discussion HWE in Hispanics n Appropriate genetic model n Interpretation of haplotype blocks n

Haplotypes 0 29 80 5 3 6 rs AFA Block 2 CHN Block 1 3 75 4 s 9 1 2 3 4 5 A G A A A 7 s 3 r 1 2 3 4 5 6 7 G G G A G 71 6 89 70 6 89 7 s 3 r r 7 45 5 8 r 4 s 2 T 5 23 9 69 s r M 4 82 5 8 21 s r G A G G C A A C C A G G G G A G G A C C G A G G A G

Acknowledgements n Wendy Post, MD, MS Johns Hopkins University n Xiuqing Guo, Ph. D Belle Fang, Ph. D Cedars-Sinai Medical Center Joe Coresh, MD, Ph. D Hunter Young, MD, MHS Michael Shlipak, MD, MPH Carmen Peralta, MD Bruce Psaty, MD, Ph. D Stephen Rich, Ph. D Jerome Rotter, MD Holly Kramer, MD Johns Hopkins University UCSF University of Washington University of Virginia Cedars-Sinai Medical Center Loyola University n n n n n