Labs Dimension Reduction Multidimensional Scaling SVM Peter Fox

Labs: Dimension Reduction, Multi-dimensional Scaling, SVM Peter Fox and Greg Hughes Data Analytics – 4600/6600 Group 3 Lab 1, March 2, 2017 1

On your own… library(EDR) # effective dimension reduction library(dr) library(clustrd) install. packages("edr. Graphical. Tools") library(edr. Graphical. Tools) demo(edr_ex 1) demo(edr_ex 2) demo(edr_ex 3) demo(edr_ex 4) 2

Some examples – group 3/ • • lab 1_dr 1. R lab 1_dr 2. R lab 1_dr 3. R lab 1_dr 4. R 3

MDS – group 3/ • lab 1_mds 1. R • lab 1_mds 2. R • lab 1_mds 3. R • http: //www. statmethods. net/advstats/mds. htm l • http: //gastonsanchez. com/blog/howto/2013/01/23/MDS-in-R. html 4

R – many ways (of course) library(igraph) g <- graph. full(nrow(dist. au)) V(g)$label <- city. names layout <- layout. mds(g, dist = as. matrix(dist. au)) plot(g, layout = layout, vertex. size = 3) 5

Work through these… • lab 1_svm 1. R –> lab 1_svm 11. R • lab 1_svm_rpart 1. R • Exercise various parts of SVM, parameters, kernels, etc… • Karatzoglou et al. 2006 http: //aquarius. tw. rpi. edu/html/DA/v 15 i 09. pdf 6

Ozone > library(e 1071) > library(rpart) > data(Ozone, package=“mlbench”) ># http: //math. furman. edu/~dcs/courses/math 47/R/library/mlbench/html/Ozone. html # for field codes > ## split data into a train and test set > index <- 1: nrow(Ozone) > testindex <- sample(index, trunc(length(index)/3)) > testset <- na. omit(Ozone[testindex, -3]) > trainset <- na. omit(Ozone[-testindex, -3]) > svm. model <- svm(V 4 ~. , data = trainset, type=“C-classification”, cost = 1000, gamma = 0. 0001) > svm. pred <- predict(svm. model, testset[, -3]) > crossprod(svm. pred - testset[, 3]) / length(testindex) See: http: //cran. r-project. org/web/packages/e 1071/vignettes/svmdoc. pdf 7

Glass library(e 1071) library(rpart) data(Glass, package="mlbench") index <- 1: nrow(Glass) testindex <- sample(index, trunc(length(index)/3)) testset <- Glass[testindex, ] trainset <- Glass[-testindex, ] svm. model <- svm(Type ~. , data = trainset, cost = 100, gamma = 1) # cost = “C” coefficient (Lagrange multiplier) svm. pred <- predict(svm. model, testset[, -10]) 8
![> table(pred = svm. pred, true = testset[, 10]) true pred 1 2 3 > table(pred = svm. pred, true = testset[, 10]) true pred 1 2 3](http://slidetodoc.com/presentation_image/b88c30985ceff6bad809a855eed99d14/image-9.jpg)
> table(pred = svm. pred, true = testset[, 10]) true pred 1 2 3 5 6 7 1 12 9 1 0 0 0 2 6 19 6 5 2 2 3 1 0 2 0 0 0 5 0 0 0 6 0 0 1 0 7 0 1 0 0 0 4 9

Example lab 1_svm 1. R n <- 150 # number of data points p <- 2 # dimension sigma <- 1 # variance of the distribution meanpos <- 0 # centre of the distribution of positive examples meanneg <- 3 # centre of the distribution of negative examples npos <- round(n/2) # number of positive examples nneg <- n-npos # number of negative examples # Generate the positive and negative examples xpos <- matrix(rnorm(npos*p, mean=meanpos, sd=sigma), npos, p) xneg <- matrix(rnorm(nneg*p, mean=meanneg, sd=sigma), npos, p) x <- rbind(xpos, xneg) # Generate the labels y <- matrix(c(rep(1, npos), rep(-1, nneg))) # Visualize the data plot(x, col=ifelse(y>0, 1, 2)) legend("topleft", c('Positive', 'Negative'), col=seq(2), pch=1, text. col=seq(2)) 10
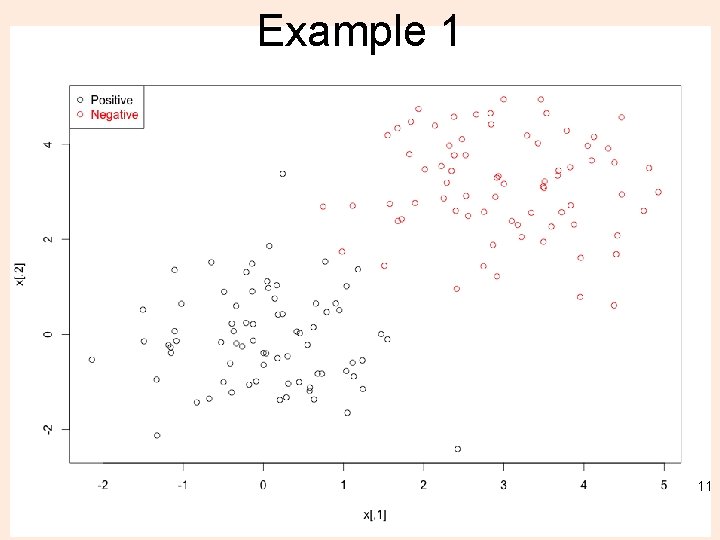
Example 1 11

Train/ test ntrain <- round(n*0. 8) # number of training examples tindex <- sample(n, ntrain) # indices of training samples xtrain <- x[tindex, ] xtest <- x[-tindex, ] ytrain <- y[tindex] ytest <- y[-tindex] istrain=rep(0, n) istrain[tindex]=1 # Visualize plot(x, col=ifelse(y>0, 1, 2), pch=ifelse(istrain==1, 1, 2)) legend("topleft", c('Positive Train', 'Positive Test', 'Negative Train', 'Negative Test'), col=c(1, 1, 2, 2), pch=c(1, 2, 1, 2), text. col=c(1, 1, 2, 2)) 12
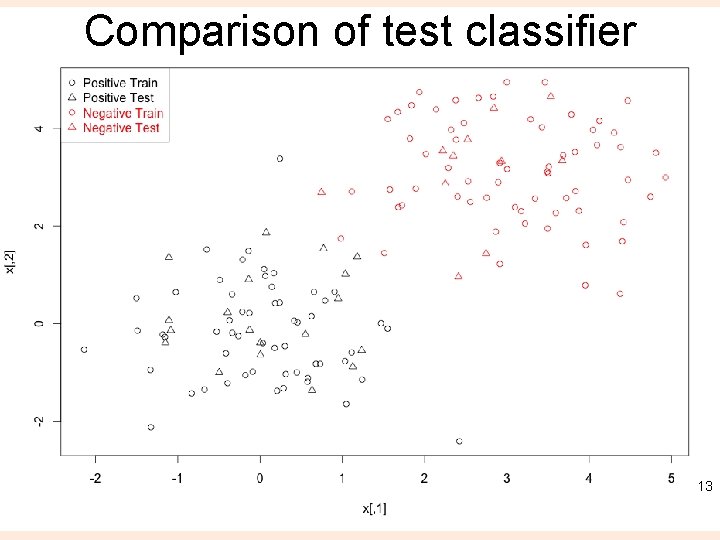
Comparison of test classifier 13

Example ctd svp <- ksvm(xtrain, ytrain, type="C-svc", kernel='vanilladot', C=100, scaled=c()) > # For example, the support vectors # General summary > alpha(svp) svp [[1]] [1] 71. 05875 28. 94125 100. 00000 # Attributes that you can access attributes(svp) # did you look? > alphaindex(svp) # For example, the support vectors [[1]] [1] 10 74 93 alpha(svp) alphaindex(svp) > b(svp) [1] -17. 3651 b(svp) # remember b? # Use the built-in function to pretty-plot the classifier plot(svp, data=xtrain) 14
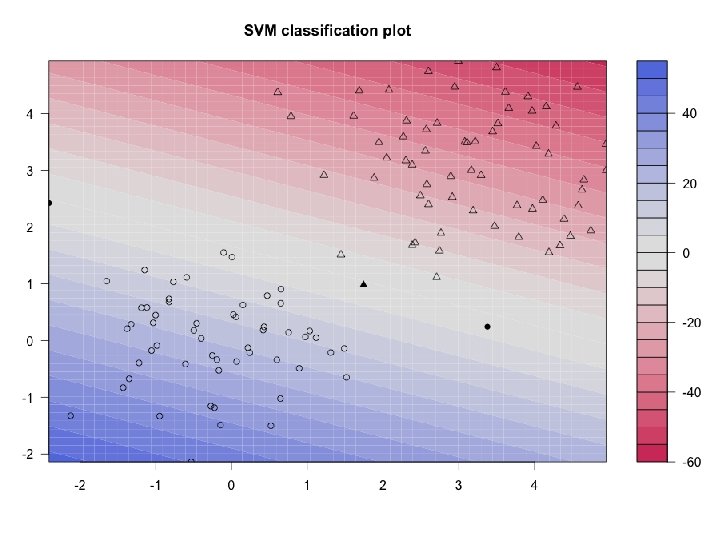
15

Do SVM for iris 16
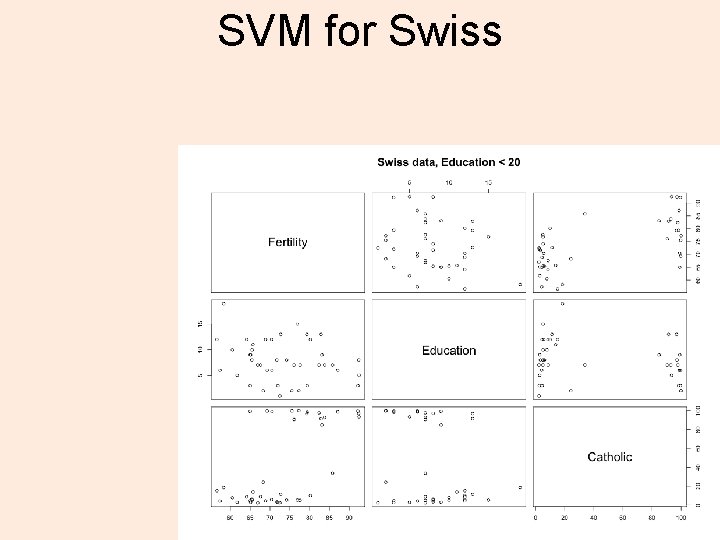
SVM for Swiss 17

e. g. Probabilities… library(kernlab) data(promotergene) ## create test and training set ind <- sample(1: dim(promotergene)[1], 20) genetrain <- promotergene[-ind, ] genetest <- promotergene[ind, ] ## train a support vector machine gene <- ksvm(Class~. , data=genetrain, kernel="rbfdot", kpar=list(sigma=0. 015), C=70, cross=4, prob. model=TRUE) ## predict gene type probabilities on the test set genetype <- predict(gene, genetest, type="probabilities") 18
![Result > genetype + [1, ] 0. 205576217 0. 794423783 [2, ] 0. 150094660 Result > genetype + [1, ] 0. 205576217 0. 794423783 [2, ] 0. 150094660](http://slidetodoc.com/presentation_image/b88c30985ceff6bad809a855eed99d14/image-19.jpg)
Result > genetype + [1, ] 0. 205576217 0. 794423783 [2, ] 0. 150094660 0. 849905340 [3, ] 0. 262062226 0. 737937774 [4, ] 0. 939660586 0. 060339414 [5, ] 0. 003164823 0. 996835177 [6, ] 0. 502406898 0. 497593102 [7, ] 0. 812503448 0. 187496552 [8, ] 0. 996382257 0. 003617743 [9, ] 0. 265187582 0. 734812418 [10, ] 0. 998832291 0. 001167709 [11, ] 0. 576491204 0. 423508796 [12, ] 0. 973798660 0. 026201340 [13, ] 0. 098598411 0. 901401589 [14, ] 0. 900670101 0. 099329899 [15, ] 0. 012571774 0. 987428226 [16, ] 0. 977704079 0. 022295921 [17, ] 0. 137304637 0. 862695363 [18, ] 0. 972861575 0. 027138425 [19, ] 0. 224470227 0. 775529773 [20, ] 0. 004691973 0. 995308027 19

R-SVM • http: //www. stanford. edu/group/wonglab/RSV Mpage/r-svm. tar. gz • http: //www. stanford. edu/group/wonglab/RSV Mpage/R-SVM. html – Read/ skim the paper – Explore this method on a dataset of your choice, e. g. one of the R built-in datasets 20

kernlab • http: //aquarius. tw. rpi. edu/html/DA/svmbasic_ notes. pdf • Some scripts: lab 1_svm 12. R, lab 1_svm 13. R 21

kernlab, svmpath and kla. R • http: //aquarius. tw. rpi. edu/html/DA/v 15 i 09. pdf • Start at page 9 (bottom) 22
- Slides: 22