I 519 Introduction to Bioinformatics Fall 2012 Sequence

I 519 Introduction to Bioinformatics, Fall 2012 Sequence Comparison

Why we compare sequences § Find sequential similarity between protein/DNA sequences – To infer functional similarity – To infer evolutionary history § Find important residues that are important for a protein’s function – Functional sites of a protein – DNA elements (e. g. , transcription factor binding sites)

Comparison of sequences at various levels § We may look at sequences differently – – Whole genome comparison (will be covered later) Whole DNA/protein sequence Protein domains Motifs (protein motifs & motifs at DNA level) § For protein-coding genes, comparison at amino acid level instead of nucleotides to achieve higher sensitivity & specificity (20 letters versus 4 letters)
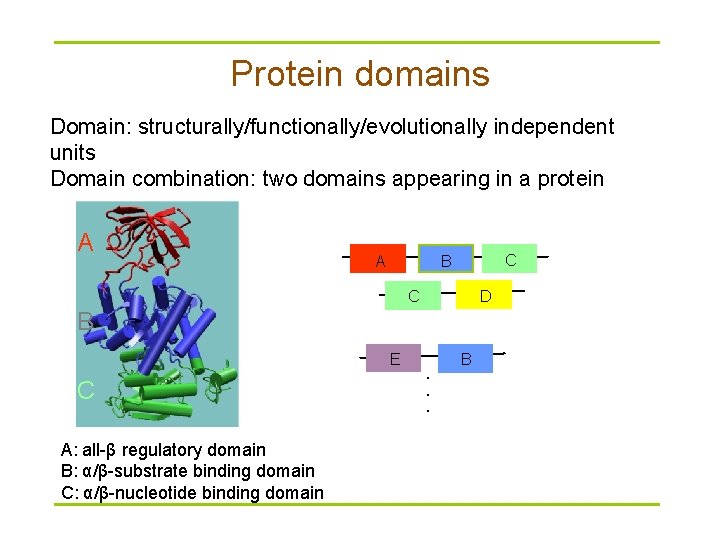
Protein domains Domain: structurally/functionally/evolutionally independent units Domain combination: two domains appearing in a protein A A C B D C B E C A: all-β regulatory domain B: α/β-substrate binding domain C: α/β-nucleotide binding domain . . . B

PROSITE patterns § Described by regular grammars § Programs that allow to search databases for PROSITE patterns (e. g. , Scan. Prosite) § We have seen the ATP-binding motif, [AG]-x{4}-G-K[ST]); another example [EDQH]-x-K-x-[DN]-G-x-R[GACV] § Rules: – – – Each position is separated by a hyphen One character denotes residuum at a given position […] denoted a set of allowed residues (n) denotes repeat of n (n, m) denoted repeat between n and m inclusive Ex. ATP/GTP binding motive [SG]=X(4)-G-K-[DT]

Three principle methods of pairwise sequence alignment § Dot matrix analysis § The dynamic programming algorithm § Word or k-tuple methods, such as used by FASTA and BLAST.
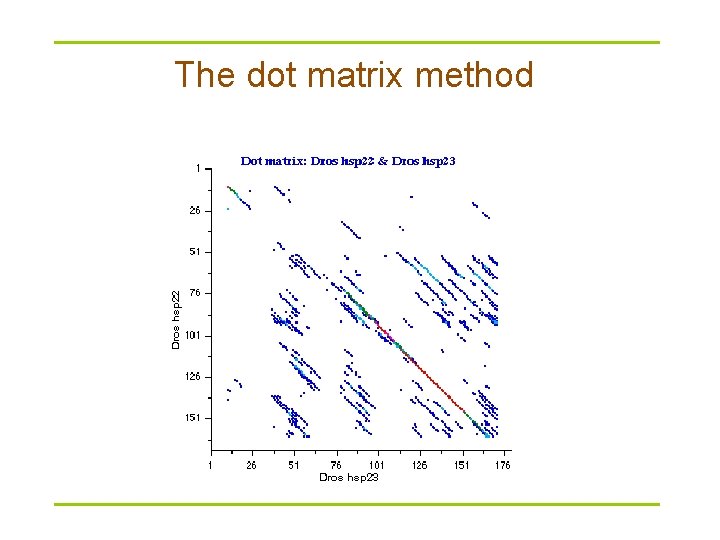
The dot matrix method
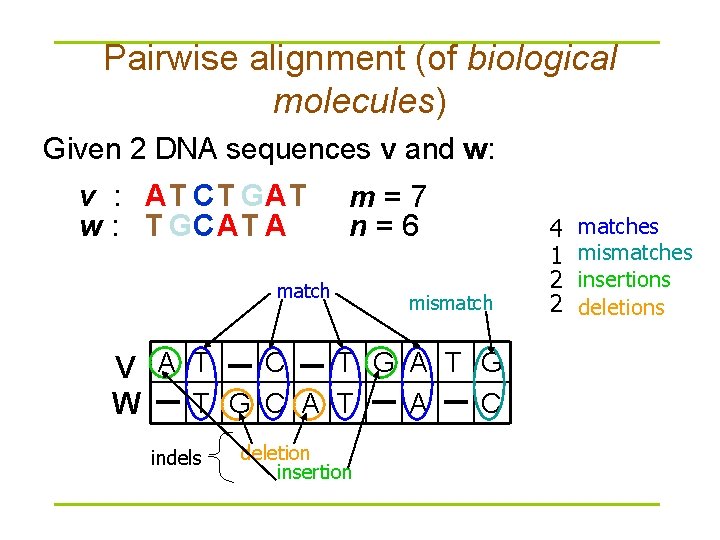
Pairwise alignment (of biological molecules) Given 2 DNA sequences v and w: v : AT CT GAT w : T GCAT A m=7 n=6 match mismatch C T G A T G V A T T G C A T A C W indels deletion insertion 4 1 2 2 matches mismatches insertions deletions

A simple string comparison solution: Hamming distance § The most often used distance on strings in computer science: the number of positions at which the corresponding symbols are different § Hamming distance always compares i-th letter of v with i-th letter of w V = ATAT W = TATA Hamming distance: d(v, w)=8 Computing Hamming distance is a trivial task

Hamming distance is easy to compute, but… • This makes some sense on comparing DNA sequences in some cases. But there are other mutations • • • Substitution ACAGT ACGGT Insertion/deletion (indel) ACAGT ACGT Inversion ACA……GT AG……ACT Translocation AC……AG…TAA AG…TC……AAA Duplication • We only consider the first two mutations for now. • There algorithms for the other mutations…

Comparing two strings: Edit distance Levenshtein (1966) introduced edit distance between two strings as the minimum number of elementary operations (insertions, deletions, and substitutions) to transform one string into the other d(v, w) = MIN number of elementary operations to transform v w

Edit distance & Hamming distance always compares i-th letter of v with i-th letter of w V = ATAT W = TATA Just one shift Make it all line up Hamming distance: d(v, w)=8 Computing Hamming distance is a trivial task Edit distance may compare i-th letter of v with j-th letter of w V = - ATAT W = TATA Edit distance: d(v, w)=2 Computing edit distance is a non-trivial task How to find what j goes with what i ? ? ?

Edit Distance: Example TGCATAT ATCCGAT in 5 steps TGCATAT TGCATA TGCAT ATCCAT ATCCGAT (delete last T) (delete last A) (insert A at front) (substitute C for 3 rd G) (insert G before last A) (Done) Edit distance = 5 But how? Dynamic programming
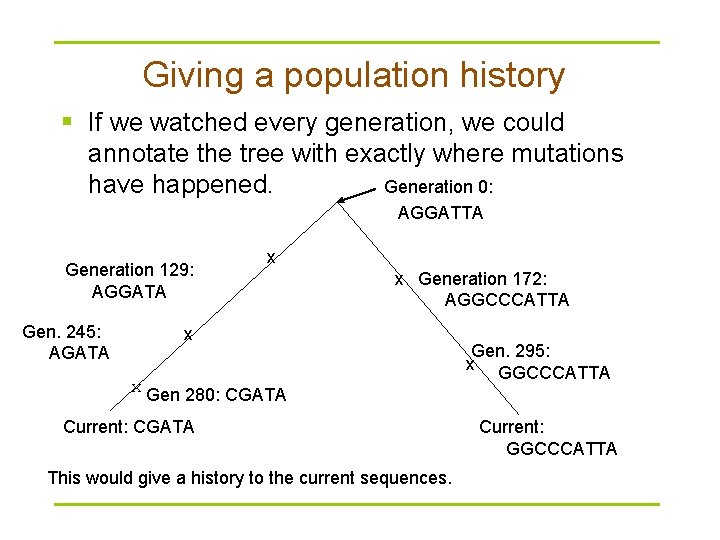
Giving a population history § If we watched every generation, we could annotate the tree with exactly where mutations have happened. Generation 0: AGGATTA Generation 129: AGGATA Gen. 245: AGATA x x Generation 172: AGGCCCATTA x x Gen 280: CGATA Current: CGATA This would give a history to the current sequences. Gen. 295: x GGCCCATTA Current: GGCCCATTA

Edit distance v. s. ancestral reconstruction • Edit distance simpler than ancestral reconstruction • Orders of the edit operations do not matter. • If two events overlap or even cancel each other in the evolution, they cannot be seen at edit distance. • It is a distance metric. • Identity: d(x, y)=0 iff x=y • Symmetry: d(x, y) = d(y, x) • Triangular Inequality: d(x, z) <= d(x, y) + d(y, z)

Alignment • Hard to visually show the edit distance: • E. g. C T@4, insert C@6, delete@9 • Alignment is much nicer: • ATGCA-TTTA ||| | ATGTACTT-A match =0, mismatch = -1, indel = -1. Score = the total score of each position of the alignment. • Then computing edit distance is equivalent to finding the optimal (maximum scoring) alignment.

“Optimal” alignment • The word “optimal” alignment is somewhat misleading. Ideally we want to find the “real” alignment of the sequences according to the real evolution instead. • Here we try to find the “optimal” alignment. • “optimal” solution is not necessary the correct solution. It all depends on how good the score function is. • The identity scoring scheme is not a very accurate one. • For example, transitions and transversions have the same score. • Along this alignment topic, we will refine the score functions.

Scoring sequence alignment § § • • • § How to score an alignment? Simplest scoring scheme: 0 = match -1 = mismatch -1 = indel This is called “linear gap penalty” because the cost of a gap (consecutive indels) is proportional to its length. (We could have each gap position cost g, for some negative constant g. ) § Let’s see some examples

Given alignment, it is trivial to compute alignment score AATGCGA-TTTT || | ||| G-TG--ACTTTC 6 matches: 0 2 mismatches: -2 4 indels: -4 edit distance (alignment score) = -6 AATG-CGATTTT || | || G-TGAC-TTTC 5 matches: 0 3 mismatches: -3 4 indels: -4 edit distance (alignment score) = -7

Alignment with DP § The question is how alignment can be computed with a computer? § Dynamic Programming – Requires the subsolution of an optimal solution is also optimal.
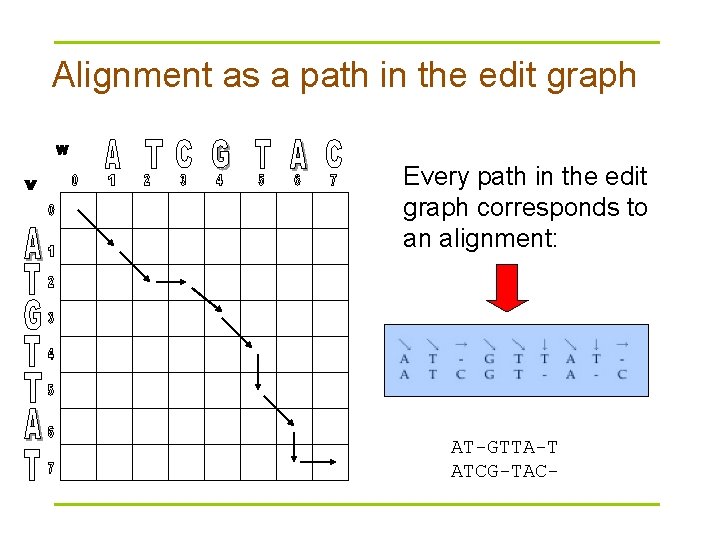
Alignment as a path in the edit graph Every path in the edit graph corresponds to an alignment: AT-GTTA-T ATCG-TAC-

Recursive definition
![Dynamic programming algorithm S[0, 0] = 0 S[i, 0] = S[i-1, 0] + g Dynamic programming algorithm S[0, 0] = 0 S[i, 0] = S[i-1, 0] + g](http://slidetodoc.com/presentation_image_h/2f378a1faf2ae16bfcfe892a7aa55985/image-23.jpg)
Dynamic programming algorithm S[0, 0] = 0 S[i, 0] = S[i-1, 0] + g S[0, j] = S[0, j-1] + g for i from 1 to M for j from 1 to N S[i, j] = max{S[i-1, j-1]+s(x[i], y[j]), S[i-1, j]+g, S[i, j-1]+g} Output S[M, N]
![Fill up the dynamic programming matrix seq[1]=PELICAN seq[2]=CWELACANTH DP Matrix # C W E Fill up the dynamic programming matrix seq[1]=PELICAN seq[2]=CWELACANTH DP Matrix # C W E](http://slidetodoc.com/presentation_image_h/2f378a1faf2ae16bfcfe892a7aa55985/image-24.jpg)
Fill up the dynamic programming matrix seq[1]=PELICAN seq[2]=CWELACANTH DP Matrix # C W E L A # 0 -1 -2 -3 -4 -5 P -1 -1 -2 -3 -4 -5 E -2 -2 -3 -4 L -3 -3 -2 -3 I -4 -4 -3 -3 C -5 -4 -5 -5 -4 -4 A -6 -5 -5 -6 -5 -4 N -7 -6 -6 -5 Scoring function: missmatch = -1 indel = -1 C -6 -6 -5 -4 -4 -3 -4 -5 A -7 -7 -6 -5 -5 -4 -3 -4 N -8 -8 -7 -6 -6 -5 -4 -3 T H -9 -10 -8 -9 -7 -8 -6 -7 -5 -6 -4 -5 A bottom-up calculation to get the optimal score (only!)

Traceback to get the actual alignment # # P E L I C A N 0 -1 -2 -3 -4 -5 -6 -7 C -1 -1 -2 -3 -4 -4 -5 -6 W -2 -2 -2 -3 -4 -5 -5 -6 E -3 -3 -2 -3 -4 -5 -6 -6 L -4 -4 -3 -2 -3 -4 -5 -6 A -5 -5 -4 -3 -3 -4 -4 -5 C -6 -6 -5 -4 -4 -3 -4 -5 A -7 -7 -6 -5 -5 -4 -3 -4 N -8 -8 -7 -6 -6 -5 -4 -3 No need to physically record the green arrows Instead, we will trace back: following the red arrows! T H -9 -10 -8 -9 -7 -8 -6 -7 -5 -6 -4 -5 CWELACANTH -PELICAN--

More formal backtracking Idea: We go from upper left to lower right. Backtrack the optimal path! Start in lower right: let i = m, j = n Until i = 0, j = 0: Figure out which of the three terms gave rise to M[i, j] by picking the largest. M[i-1, j]+indel, M[i, j-1]+indel, M[i-1, j-1]+f(s[i], t[j]) Move to the right place (reduce i, reduce j, or reduce both), and write down the configuration of the current column.

How similar biology and informatics are I B I O L O G Y N F O R M A T I C S

Space, time requirements • The algorithm runs in O(nm) time: Each step requires only 3 checks to other points in the matrix. • We also need O(nm) space, to store the matrix. • If we only want to know the score of the optimal alignment, we can do that in O(min(m, n)) space. • Reconstructing the alignment also requires only O(m+n) space.

Alignments are scored • Need to score alignments. • The alignment that has highest score may not be the one that actually matches evolutionary history. • So you should never trust that an alignment must be right. It just optimizes the score. • When we move to multiple alignments, things get worse: no guarantee of the optimal score, even.
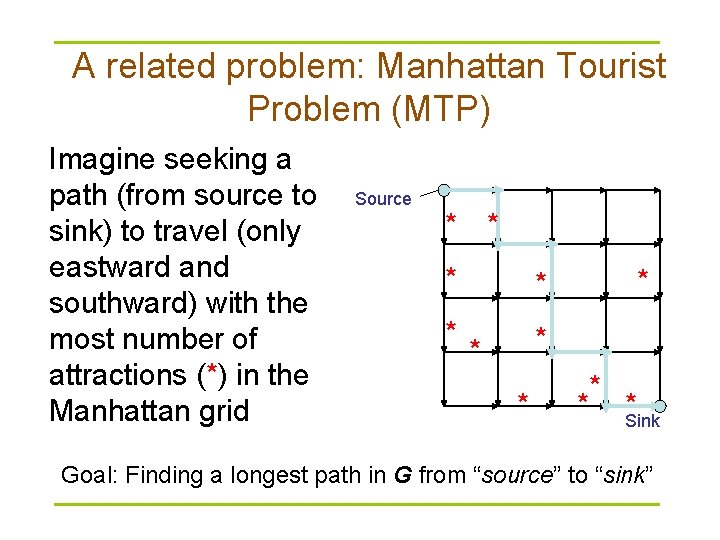
A related problem: Manhattan Tourist Problem (MTP) Imagine seeking a path (from source to sink) to travel (only eastward and southward) with the most number of attractions (*) in the Manhattan grid Source * * *Sink Goal: Finding a longest path in G from “source” to “sink”

Longest Path in DAG Problem • Goal: Find a longest path between two vertices in a weighted DAG • Input: A weighted DAG G with source and sink vertices • Output: A longest path in G from source to sink

“Edit distance problem” Runtime § It takes O(nm) time to fill in the dynamic programming matrix. § Why O(nm)? The pseudocode consists of a nested “for” loop inside of another “for” loop to set up the dynamic programming matrix.

§ Reading: – Chapter 4 (Producing and Analyzing Sequence Alignments) § Next time we will talk about global and local pairwise sequence alignment, focus on – How alignment of biological sequences is different from comparison of two strings (scoring matrix + indel penalties) – Global versus local
- Slides: 33