I 519 Introduction to Bioinformatics 2012 Genome Comparison

I 519 Introduction to Bioinformatics, 2012 Genome Comparison

Whole genome comparison/alignment § § Build better phylogenies Identify polymorphism Detect gene-level events Compare different assemblies of a single genome

Whole genome comparison § Aligning whole genomes is a fundamentally different problem than aligning short sequences. § Need to consider the presence of large-scale evolutionary events – – Gene duplication & loss Horizontal gene transfer Repetitive sequences (repeats) Gene rearrangement and inversion § Pairwise and multiple genome comparison – Multiple genome alignment provides a basis for research into comparative genomics and the study of evolutionary dynamics.
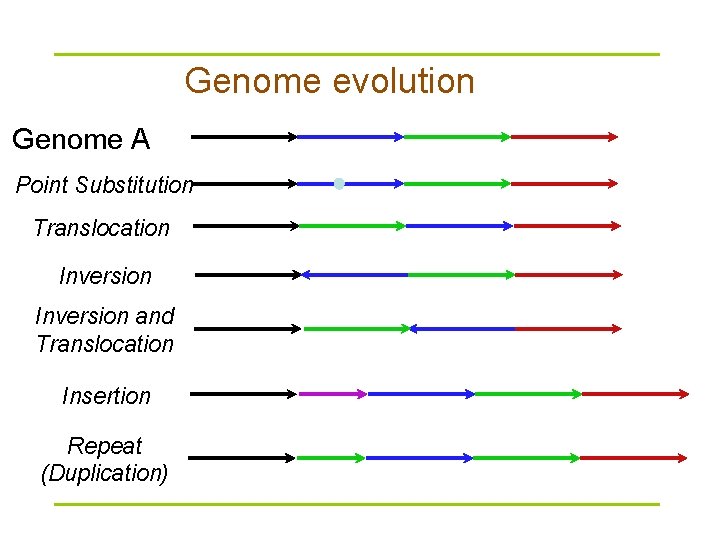
Genome evolution Genome A Point Substitution Translocation Inversion and Translocation Insertion Repeat (Duplication)

Basic algorithms: use anchoring as a heuristic to speed alignment § Assumption: highly similar subsequences can be found quickly and are likely to be part of the correct global alignment. § These local alignments are used to anchor a global alignment (alignment anchor), reducing the number of possible global alignments considered during a subsequent O(n 2) dynamic programming step. § Select a single collinear set of alignment anchors § Many tools have been developed

Rearrangement free or not § Free of rearrangement – Assume the input sequences are free from significant rearrangements of sequence elements, selecting a single collinear set of alignment anchors – Pairwise: MUMmer, GLASS, AVID, and WABA align pairs of long sequences – Multiple alignment: MAVID, MLAGAN, and MGA § Consider rearrangement – Shuffle-LAGAN (2003, first genome comparison method described that explicitly deals with genome rearrangements) – Multi. Pip. Maker (2003) – Mauve (2004, multiple) – Enredo and Pecan (2008) – GR-Aligner (2009, pairwise)

MUMer method § MUMer combines suffix trees, the longest increasing subsequence (LIS) and SW alignment § Maximal Unique Match (MUM) Identification - Identify the longest strings in Genome 1 that have one identical match in Genome 2 – Naïve method: O(N 2) – Using suffix tree: O(N) § Ordered MUM Selection - Identify the longest set of MUMs such that they occur in order in each of the genomes (using a variation of the well-known algorithm to find the LIS of a sequence of integers) § Processing Non-matched Regions - Classify nonmatched regions as either insertions, SNPs or highly polymorphic regions

Suffix tree § Suffix tree is data structure, which allows one to find, extremely efficiently, all distinct subsequences in a given sequence. § There are efficient algorithms to construct suffix trees given by Weiner (1973) and Mc. Creight (1976) (in linear time) § For the task of comparing two DNA sequences, suffix trees allow one to quickly find all subsequences shared by the two inputs. § The genome alignment is then built upon this information.
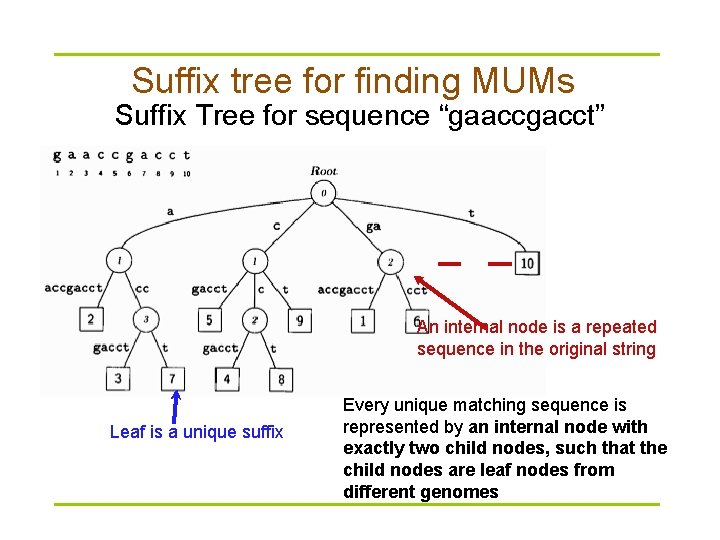
Suffix tree for finding MUMs Suffix Tree for sequence “gaaccgacct” An internal node is a repeated sequence in the original string Leaf is a unique suffix Every unique matching sequence is represented by an internal node with exactly two child nodes, such that the child nodes are leaf nodes from different genomes
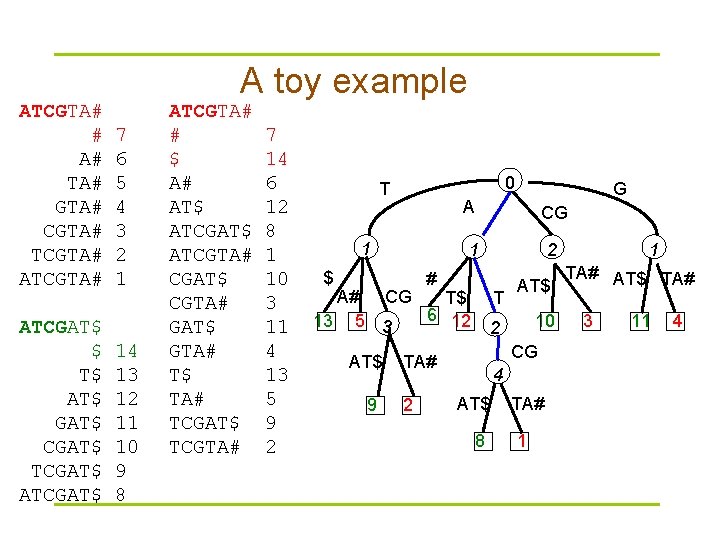
A toy example ATCGTA# # A# TA# GTA# CGTA# TCGTA# ATCGAT$ $ T$ AT$ GAT$ CGAT$ TCGAT$ ATCGAT$ 7 6 5 4 3 2 1 14 13 12 11 10 9 8 ATCGTA# # $ A# AT$ ATCGTA# CGAT$ CGTA# GAT$ GTA# T$ TA# TCGAT$ TCGTA# 7 14 6 12 8 1 10 3 11 4 13 5 9 2 0 T 1 $ A CG 1 2 # A# CG T$ 6 12 13 5 3 AT$ 9 T AT$ 10 2 CG TA# 2 G 4 AT$ 8 TA# 1 1 TA# AT$ TA# 3 11 4

Suffix tree & suffix array for string matching § Preprocess text T, not pattern P – O(m) preprocess time (m: the length of the text) – O(n+k) search time (n: the length of the pattern) • k is number of occurrences of P in T § Match pattern P against tree starting at root until – Case 1, P is completely matched • Every leaf below this match point is the starting location of P in T – Case 2: No match is possible • P does not occur in T
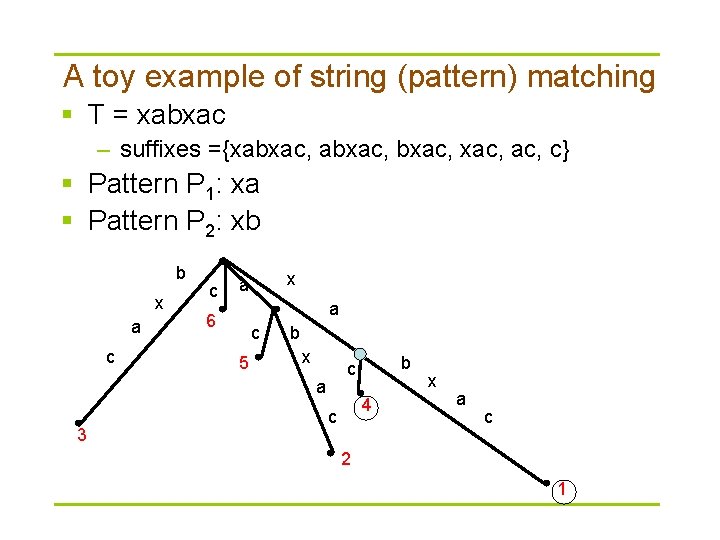
A toy example of string (pattern) matching § T = xabxac – suffixes ={xabxac, ac, c} § Pattern P 1: xa § Pattern P 2: xb b x a c c x a a 6 c 5 b x a 3 b c 4 c x a c 2 1

Suffix array: a sorted list of the suffixes of a given string; the start positions are sorted in lexicographical (alphabetical) order Straightforward implementation: O(m 2 logm), reduced to O(mlogm) (utilizing partial sorts) m: the length of the text Suffix array enables binary search for any substring, e. g. CAD O(nlogm), reduced to O(n + logm) if use LCP (longest common prefix) n: the length of the pattern Suffix array is more compact than a suffix tree ABRACADABRA# 11 10 7 0 3 5 8 1 4 6 9 2 webglimpse. net/pubs/suffix. pdf # A# ABRACADABRA# BRACADABRA# RACADABRA#
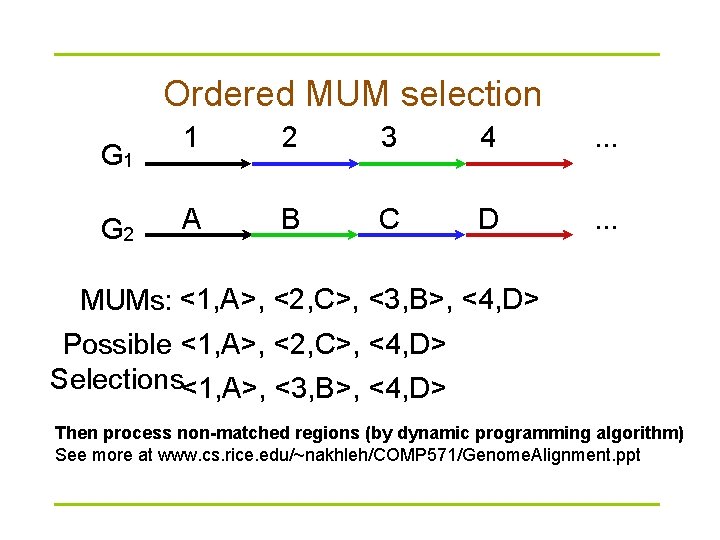
Ordered MUM selection G 1 G 2 1 2 3 4 . . . A B C D . . . MUMs: <1, A>, <2, C>, <3, B>, <4, D> Possible <1, A>, <2, C>, <4, D> Selections<1, A>, <3, B>, <4, D> Then process non-matched regions (by dynamic programming algorithm) See more at www. cs. rice. edu/~nakhleh/COMP 571/Genome. Alignment. ppt
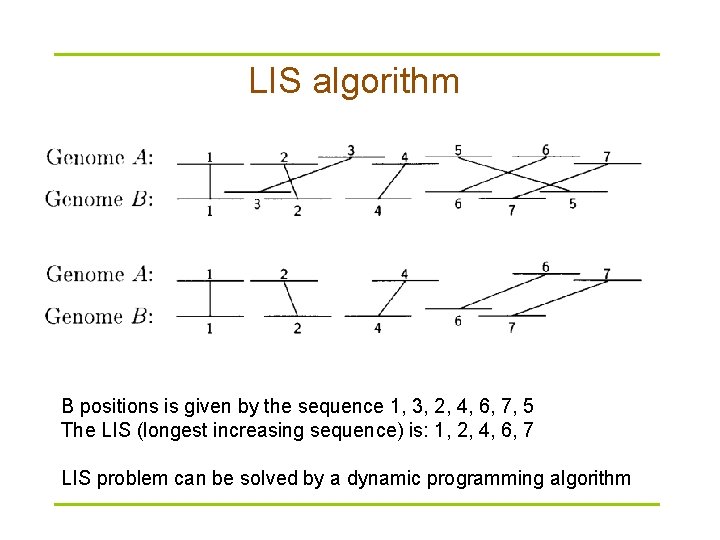
LIS algorithm B positions is given by the sequence 1, 3, 2, 4, 6, 7, 5 The LIS (longest increasing sequence) is: 1, 2, 4, 6, 7 LIS problem can be solved by a dynamic programming algorithm

Mauve § Mauve is a system for efficiently constructing multiple genome alignments in the presence of large-scale evolutionary events § Identifies conserved genomic regions, rearrangements and inversions in conserved regions, and the exact sequence breakpoints of such rearrangements across multiple genomes. § Also performs traditional multiple alignment of conserved regions to identify nucleotide substitutions and indels, using the progressive dynamic programming approach of CLUSTALW

Mauve's anchor selection algorithm § Relax anchor selection method: do not assume that the genomes under study are collinear § Identifie and align regions of local collinearity called locally collinear blocks (LCBs) – Each LCB is a homologous region of sequence shared by two or more of the genomes under study – Does not contain any rearrangements of homologous sequence (within LCB)

Mauve algorithm 1. Find local alignments (multi-MUMs), using seed-and-extend hashing method (time complexity O(G 2 n + Gn log. Gn), G is the number of genomes and n the average genome length) 2. Use the multi-MUMs to calculate a phylogenetic guide tree. 3. Select a subset of the multi-MUMs to use as anchors—these anchors are partitioned into collinear groups called LCBs, using a greedy breakpoint elimination algorithm 4. Perform recursive anchoring to identify additional alignment anchors within and outside each LCB. 5. Perform a progressive alignment of each LCB using the guide tree.

Greedy breakpoint elimination in three genomes Darling A C et al. Genome Res. 2004; 14: 1394 -1403 © 2004 by Cold Spring Harbor Laboratory Press
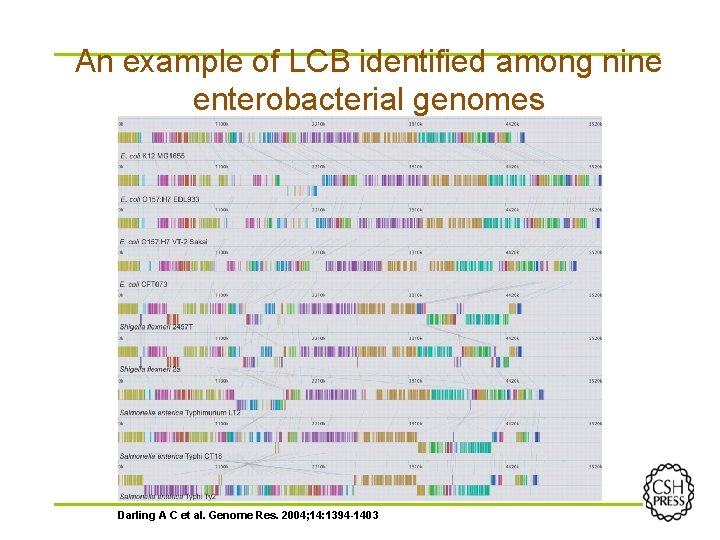
An example of LCB identified among nine enterobacterial genomes Darling A C et al. Genome Res. 2004; 14: 1394 -1403
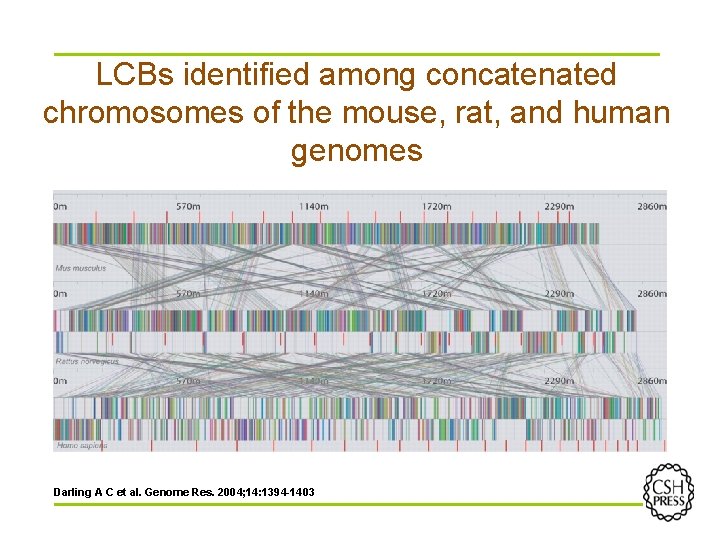
LCBs identified among concatenated chromosomes of the mouse, rat, and human genomes Darling A C et al. Genome Res. 2004; 14: 1394 -1403

Turnip vs cabbage: almost identical mt. DNA gene sequences § In 1980 s Jeffrey Palmer studied evolution of plant organelles by comparing mitochondrial genomes of the cabbage and turnip (using physical mapping) § 99%-99. 9% similarity between genes § These surprisingly identical gene sequences differed in gene order § This study helped pave the way to analyzing genome rearrangements in molecular evolution

Why we care about genome rearrangement § Evolutionary and functional analysis § Examples: – “Dynamics of Genome Rearrangement in Bacterial Populations”, using comparison of eight Yersinia (pathogenic bacteria) genomes. PLo. S Genet 4(7): e 1000128, 2008 – Genome-wide DNA excision (Oxytricha trifallax destroys 95% of its germline genome during development, including the elimination of all transposon DNA, through an exaggerated process of genome rearrangement). Science, Vol. 324. no. 5929, pp. 935 – 938, 2009
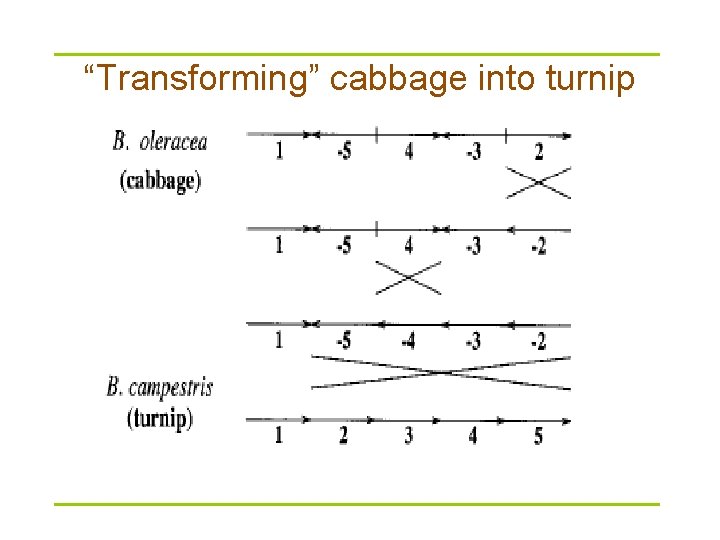
“Transforming” cabbage into turnip
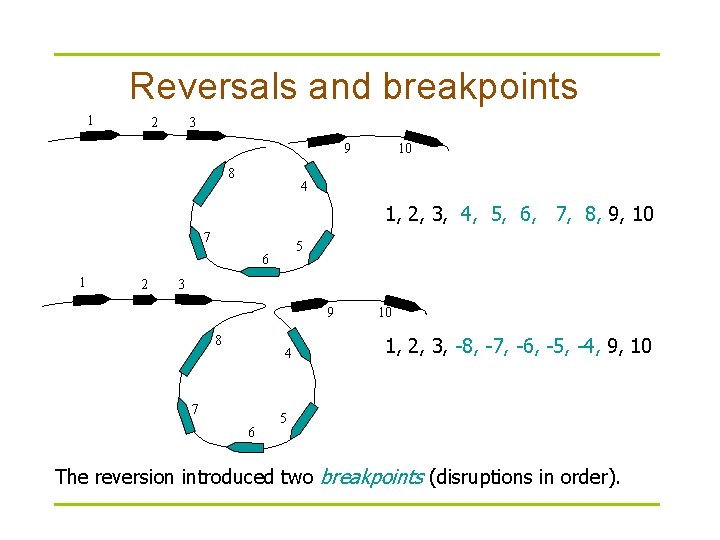
Reversals and breakpoints 1 2 3 9 8 10 4 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 7 5 6 1 2 3 9 8 4 7 6 10 1, 2, 3, -8, -7, -6, -5, -4, 9, 10 5 The reversion introduced two breakpoints (disruptions in order).
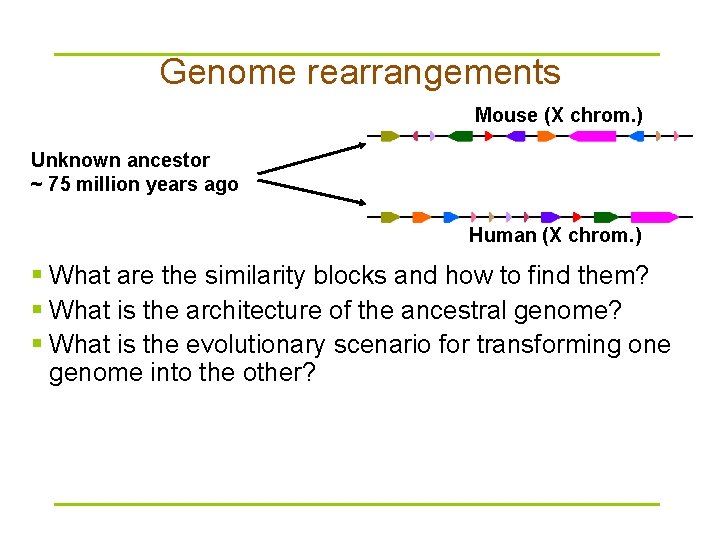
Genome rearrangements Mouse (X chrom. ) Unknown ancestor ~ 75 million years ago Human (X chrom. ) § What are the similarity blocks and how to find them? § What is the architecture of the ancestral genome? § What is the evolutionary scenario for transforming one genome into the other?
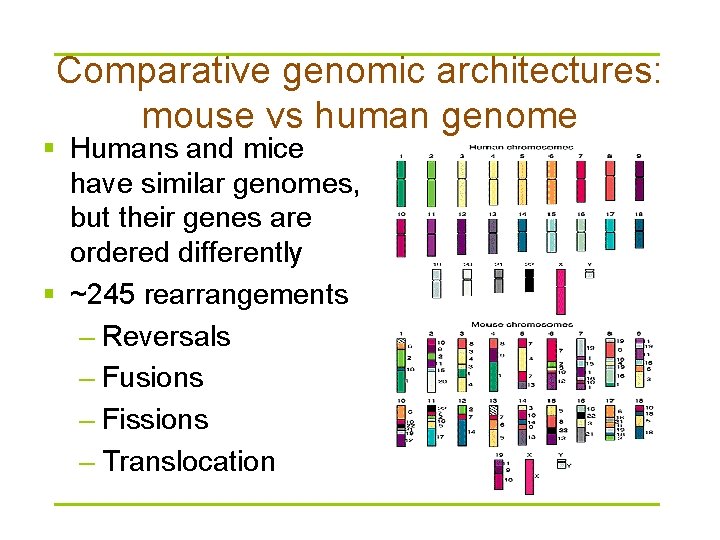
Comparative genomic architectures: mouse vs human genome § Humans and mice have similar genomes, but their genes are ordered differently § ~245 rearrangements – Reversals – Fusions – Fissions – Translocation

History of Chromosome X Rat Consortium, Nature, 2004

GRIMM § Real genome architectures are represented by signed permutations § Efficient algorithms to sort signed permutations have been developed § GRIMM web server computes the reversal distances between signed permutations: http: //nbcr. sdsc. edu/GRIMM/mgr. cgi
- Slides: 29