Fast Sequence Alignments Table of contents FASTA BLAST

Fast Sequence Alignments

Table of contents • FASTA • BLAST

Heuristic search •

FASTA • FASTA is a heuristic method for local alignment of a query string against a text string. • There are several variants of FASTA, and it continues to evolve. • The following description is "generic" and is based on the original FASTA papers • FASTA was developed by David J. Lipman and William R. Pearson in 1985

Visualizing FASTA - dotplot Note: This is only a visualization, not a data structure.

FASTA – Actual data structure The actual data structure is a Hash Table. Keys are ktups (words of length k) Values are their positions in the text.

FASTA – Actual data structure The actual data structure is a Hash Table. Keys are ktups (words of length k) Values are their positions in the text.

FASTA – Actual data structure The actual data structure is a Hash Table. Keys are ktups (words of length k) Values are their positions in the text.

FASTA – Actual data structure The actual data structure is a Hash Table. Keys are ktups (words of length k) Values are their positions in the text.

FASTA – The algorithm • text query
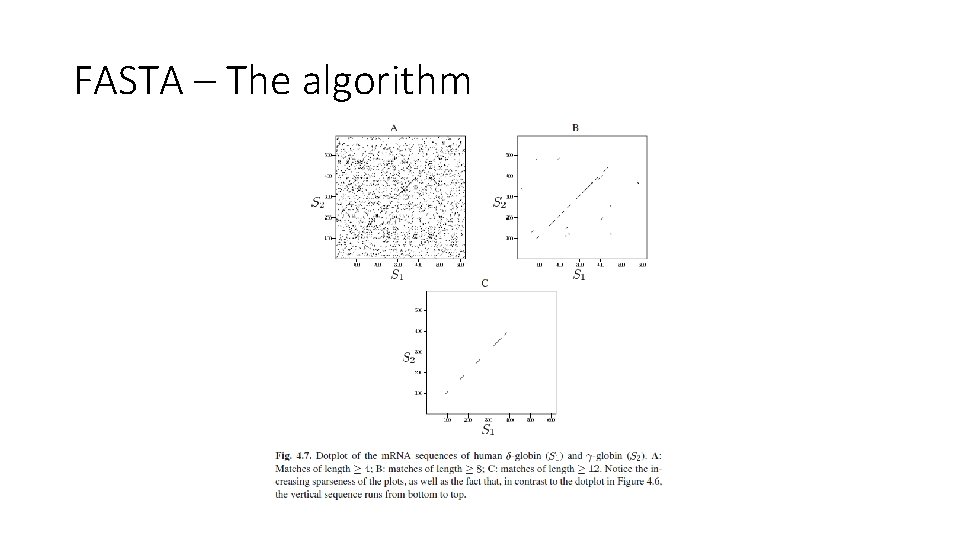
FASTA – The algorithm

FASTA – The algorithm Step 2: • Locate 10 best ‘diagonal runs’ of hot-spots in the table. • A diagonal run is a sequence of consecutive hotspots in a single diagonal. query • Each hot-spot has positive score • The space between consecutive hot-spots has negative score that decreases with increasing distance. • The score of a diagonal run is the sum of the hotspot scores and the inter-hotspot scores. text

FASTA – The algorithm Step 2: • The 10 diagonal runs are re-scored using a scoring matrix. • Originally were scored using PAM, later used BLOSUM 50 • Choose only diagonal runs with score above some threshold

FASTA – The algorithm Step 3: • Chaining diagonal runs – allowing insertions and deletions • The chaining is preformed by finding heaviest path in a graph text query

FASTA – The algorithm Step 4: • Perform local alignment in a narrow band that encompasses the top scoring segments – optimal path may go through different segments text query

BLAST – Basic Local Alignment Search Tool • BLAST was published two years after FASTA and immediately became the dominant search engine for biological sequence database. • The main reasons are: speed, it outputs a number of solutions, and it outputs an estimate of statistical significance. • Do note that both algorithms have changed quite a bit since, and both have multiple versions. • Similar to FASTA, BLAST also excludes unpromising areas of the alignment matrix.

BLAST – Maximal Segment Pair • Maximal Segment Pairs (MSPs) are pairs of equal length substrings (of the query and text sequences) whose scores cannot be improved by extending or shortening each segement. • MSPs are the central notion of the BLAST algorithm.
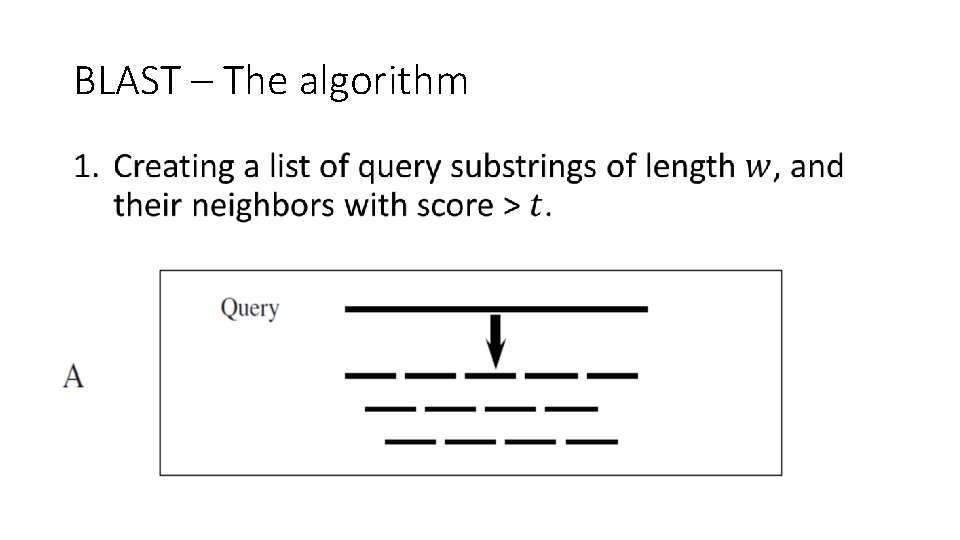
BLAST – The algorithm •
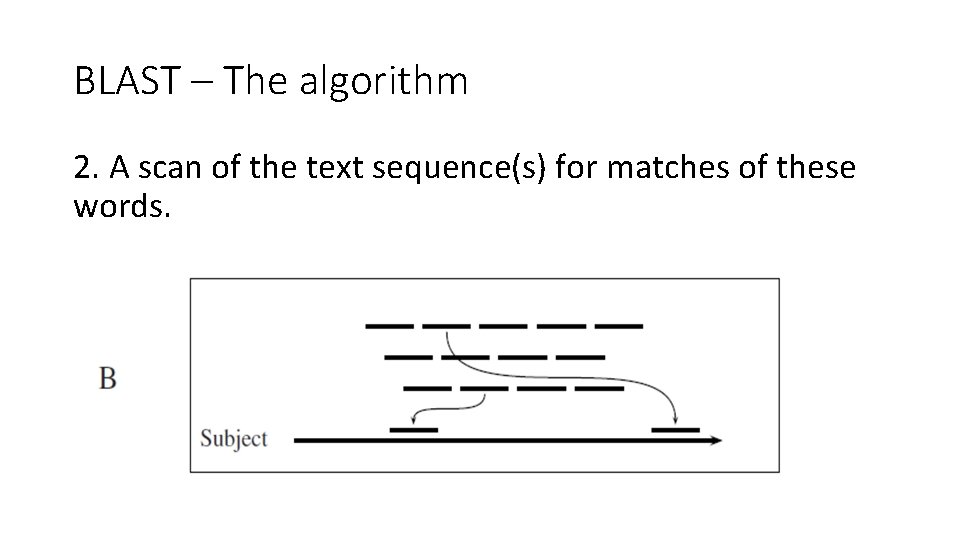
BLAST – The algorithm 2. A scan of the text sequence(s) for matches of these words.
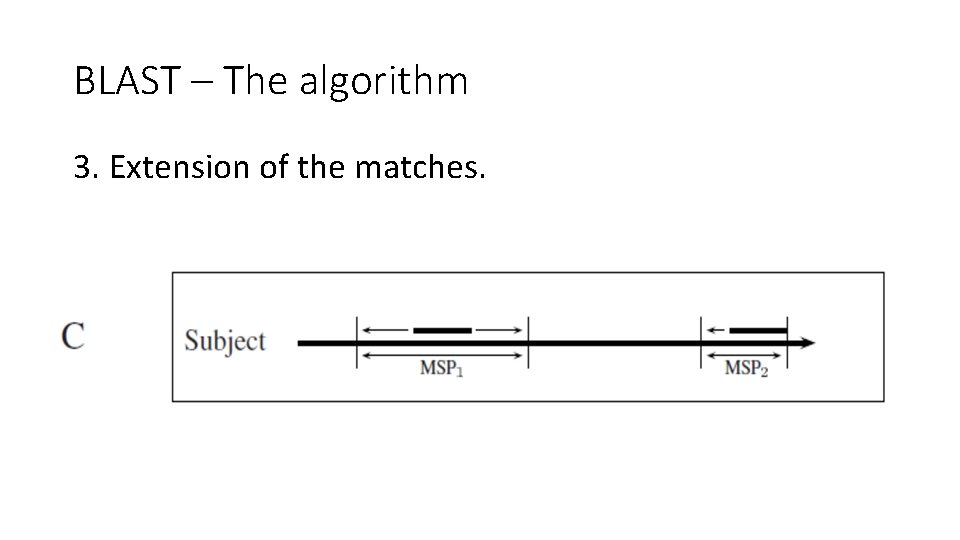
BLAST – The algorithm 3. Extension of the matches.
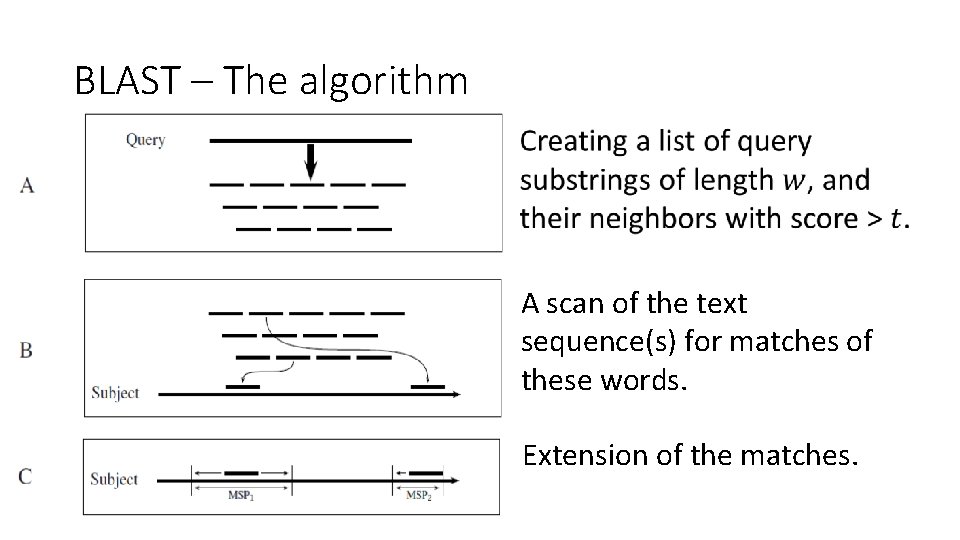
BLAST – The algorithm A scan of the text sequence(s) for matches of these words. Extension of the matches.

BLAST – Stage 1 – query listing •

BLAST – Stage 1 – query listing •
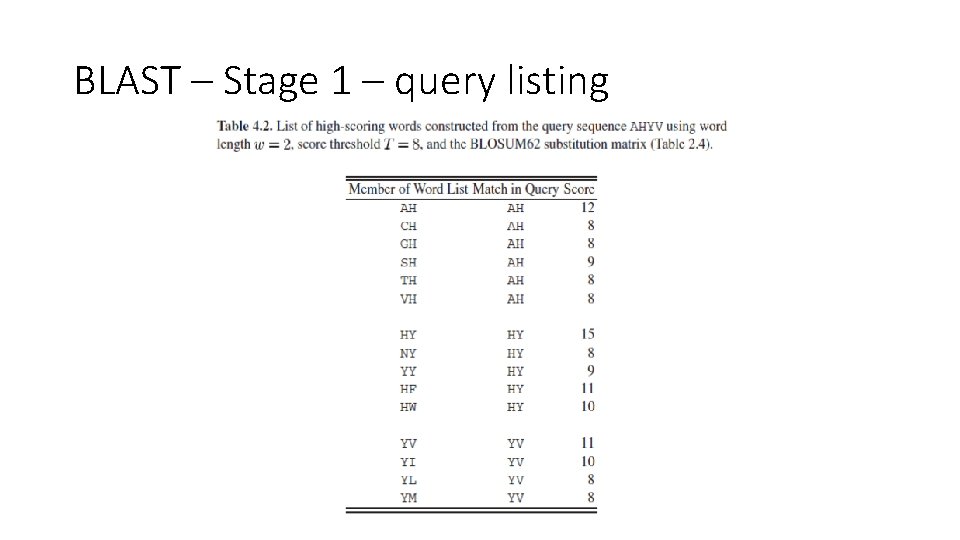
BLAST – Stage 1 – query listing
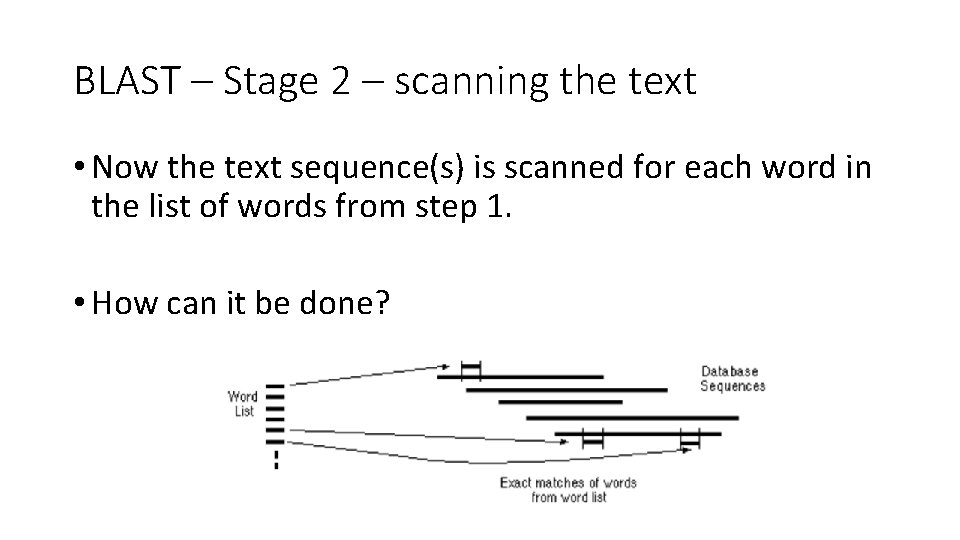
BLAST – Stage 2 – scanning the text • Now the text sequence(s) is scanned for each word in the list of words from step 1. • How can it be done?

BLAST – Stage 3 – extension • The high scoring segment pair found before (HSPs) are now extended on both sides, when only matches and substitutions are allowed. • Extension stops when the extended HSP’s score drops X below the maximal score obtained so far (not the same one from stage 2). • The results are MSPs.

BLAST – Stage 4 – Evaluate significance •

BLAST – Stage 4 – Evaluate significance •

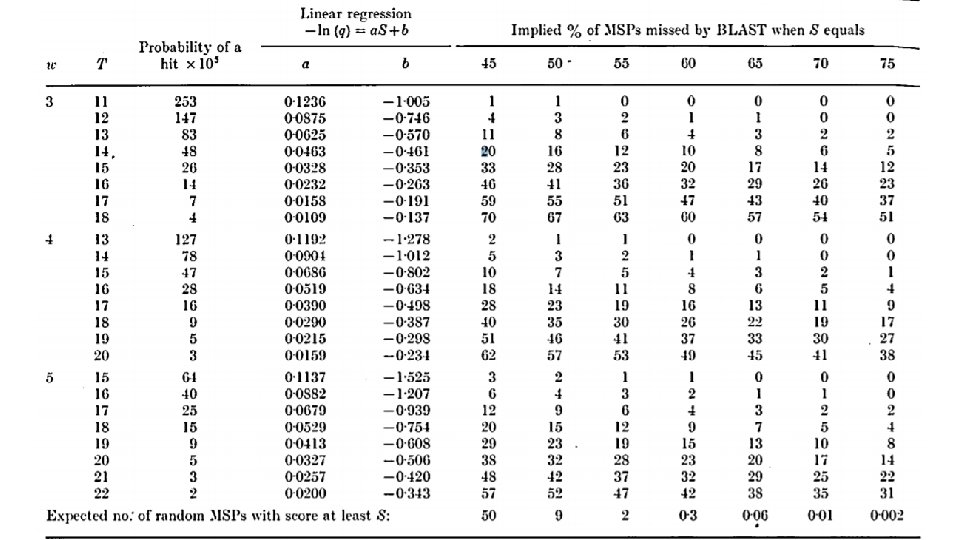
BLAST – Some further runtime discussion Let us introduce two definitions to help us analyze BLAST (and many other similar algorithms): • Sensitivity – The proportion of correct results returned (in our case, homologous) among all correct results in the DB. • Specificity – The proportion of correct results returned among all of the returned results.

BLAST – Some further runtime discussion •

Mini-Project Data • Go to: • https: //www. cs. bgu. ac. il/~negevcb/OGMFinder/input/ • There is a readme file explaining the content and structure of the files
- Slides: 32