DNA Technologies Introduction Cloning PCR Gel Electrophoresis DNA










- Slides: 26

DNA Technologies (Introduction) Cloning PCR Gel Electrophoresis DNA Sequencing DNA fingerprinting GMO : Genetically Modified Organism ……

Biotechnology § Biotechnology • Any technique applied to biological systems to manipulate processes § DNA technologies isolate purify, analyze and manipulate DNA sequences • DNA fingerprinting used in forensics § Genetic engineering uses DNA technologies to alter genes for practical purposes

DNA Cloning § Bacterial enzymes (restriction endonucleases) form the basis of DNA cloning § Bacterial plasmids illustrate the use of restriction enzymes in cloning § DNA libraries contain collections of cloned DNA fragments § Polymerase chain reaction (PCR) amplifies DNA in vitro

Recombinant DNA § DNA cloning provides many copies of a gene • Used for research or manipulation § Recombinant DNA contains DNA from multiple sources joined together • Recombinant plasmids containing gene of interest can be cloned in E. coli

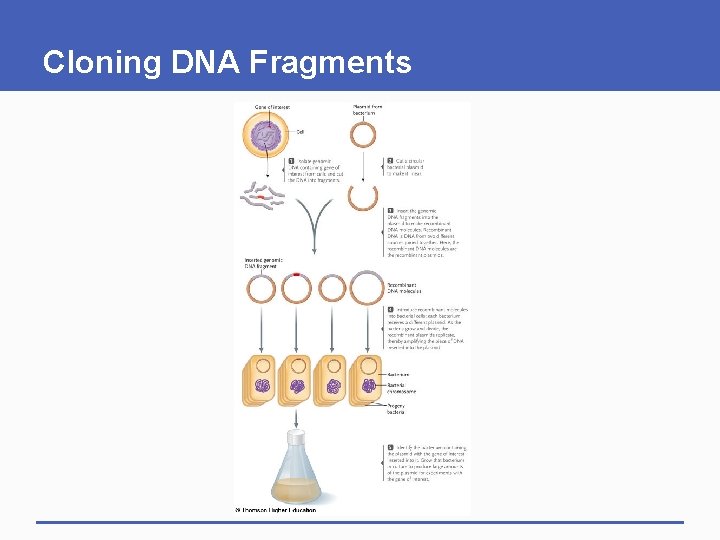
Cloning DNA Fragments

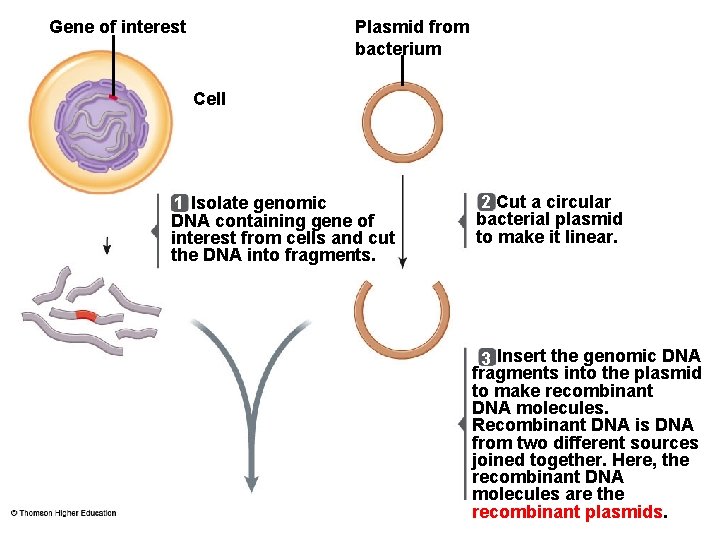
Gene of interest Plasmid from bacterium Cell 1 Isolate genomic DNA containing gene of interest from cells and cut the DNA into fragments. 2 Cut a circular bacterial plasmid to make it linear. 3 Insert the genomic DNA fragments into the plasmid to make recombinant DNA molecules. Recombinant DNA is DNA from two different sources joined together. Here, the recombinant DNA molecules are the recombinant plasmids.

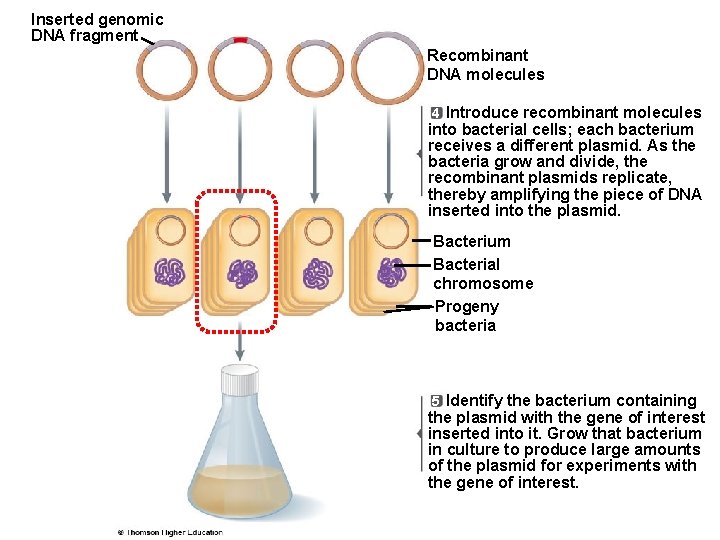
Inserted genomic DNA fragment Recombinant DNA molecules 4 Introduce recombinant molecules into bacterial cells; each bacterium receives a different plasmid. As the bacteria grow and divide, the recombinant plasmids replicate, thereby amplifying the piece of DNA inserted into the plasmid. Bacterium Bacterial chromosome Progeny bacteria 5 Identify the bacterium containing the plasmid with the gene of interest inserted into it. Grow that bacterium in culture to produce large amounts of the plasmid for experiments with the gene of interest.

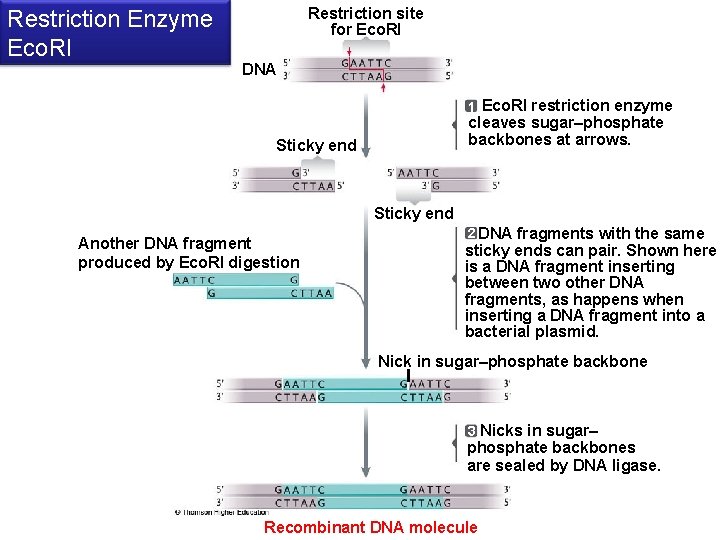
Restriction site for Eco. RI Restriction Enzyme Eco. RI DNA 1 Eco. RI restriction enzyme cleaves sugar–phosphate backbones at arrows. Sticky end Another DNA fragment produced by Eco. RI digestion 2 DNA fragments with the same sticky ends can pair. Shown here is a DNA fragment inserting between two other DNA fragments, as happens when inserting a DNA fragment into a bacterial plasmid. Nick in sugar–phosphate backbone 3 Nicks in sugar– phosphate backbones are sealed by DNA ligase. Recombinant DNA molecule

Endonucleases (Restriction Enzymes) § Restriction enzymes (endunucleases) cut DNA at specific sequences in restriction sites • Restriction fragments result • Sticky ends have unpaired bases at cuts which will hydrogen bond • Ligase stitches together paired sticky ends

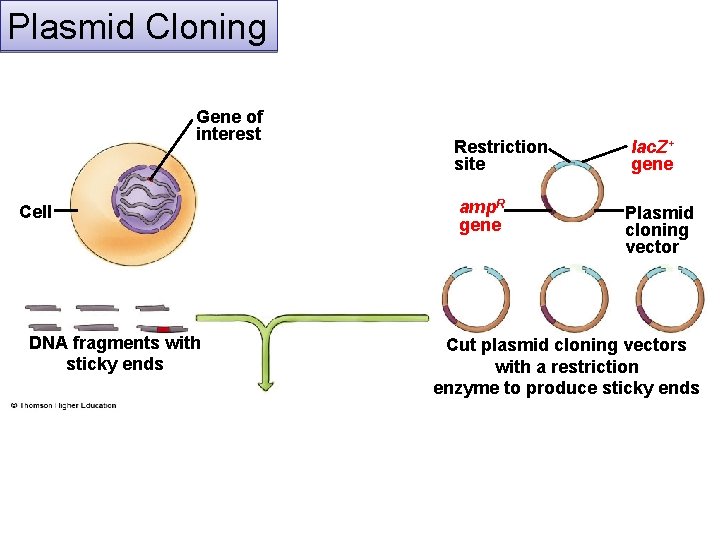
Plasmid Cloning Gene of interest Cell DNA fragments with sticky ends Restriction site amp. R gene lac. Z+ gene Plasmid cloning vector Cut plasmid cloning vectors with a restriction enzyme to produce sticky ends

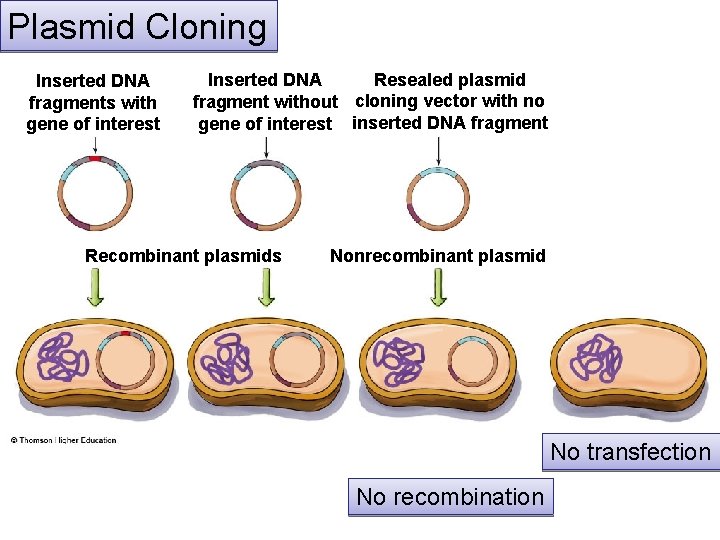
Plasmid Cloning Inserted DNA fragments with gene of interest Resealed plasmid Inserted DNA fragment without cloning vector with no gene of interest inserted DNA fragment Recombinant plasmids Nonrecombinant plasmid No transfection No recombination

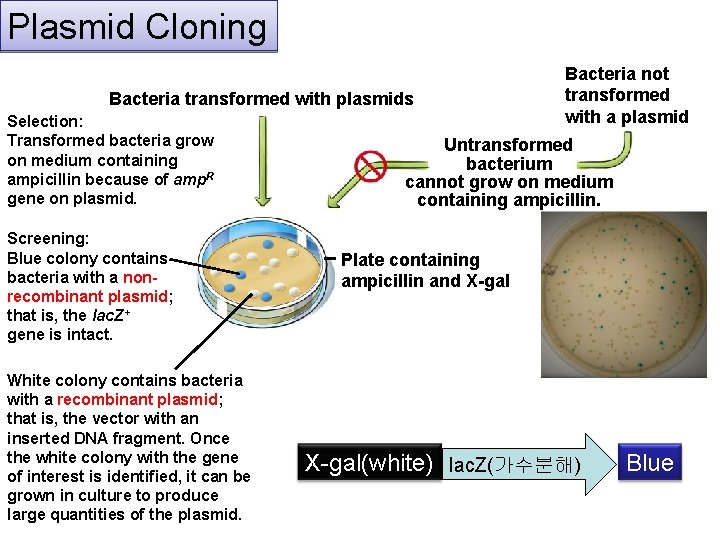
Plasmid Cloning Bacteria transformed with plasmids Selection: Transformed bacteria grow on medium containing ampicillin because of amp. R gene on plasmid. Screening: Blue colony contains bacteria with a nonrecombinant plasmid; that is, the lac. Z+ gene is intact. White colony contains bacteria with a recombinant plasmid; that is, the vector with an inserted DNA fragment. Once the white colony with the gene of interest is identified, it can be grown in culture to produce large quantities of the plasmid. Bacteria not transformed with a plasmid Untransformed bacterium cannot grow on medium containing ampicillin. Plate containing ampicillin and X-gal(white) lac. Z(가수분해) Blue

Plasmid Cloning Vectors § Engineered to contain gene of interest and sorting genes • Sorting genes identify E. coli with cloned plasmid E. coli with appropriate plasmid are 1. ampicillin resistant (transformation check by survival) 2. and blue-white screened on X-gal (recombination check by color change) then, DNA hybridization (probe) finds the gene of interest

DNA Hybridization § Uses nucleic acid probe to identify gene of interest in set of clones • Probe has tag (표지) for detection • Identified colony produces large quantities of cloned gene Tag: radioactive or fluroscent

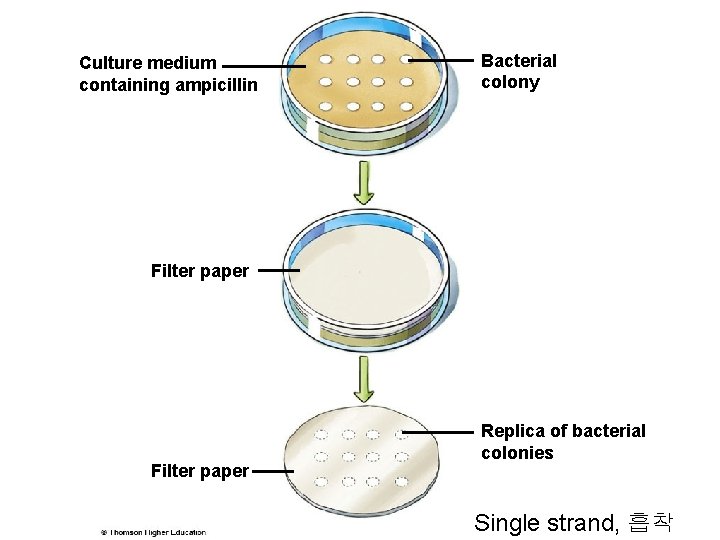
DNA Hybridization

Culture medium containing ampicillin Bacterial colony Filter paper Replica of bacterial colonies Single strand, 흡착

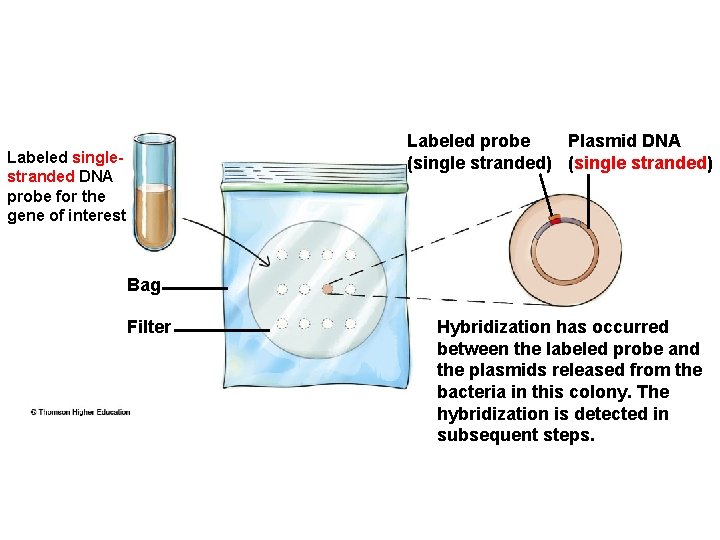
Labeled probe Plasmid DNA (single stranded) Labeled singlestranded DNA probe for the gene of interest Bag Filter Hybridization has occurred between the labeled probe and the plasmids released from the bacteria in this colony. The hybridization is detected in subsequent steps.

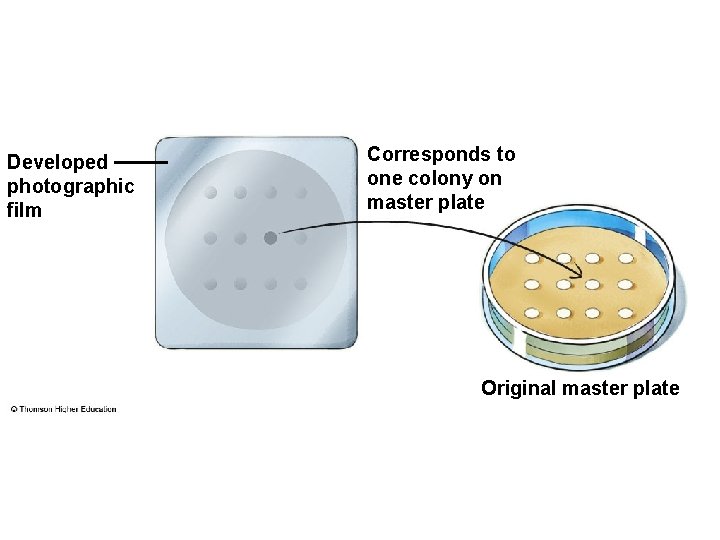
Developed photographic film Corresponds to one colony on master plate Original master plate

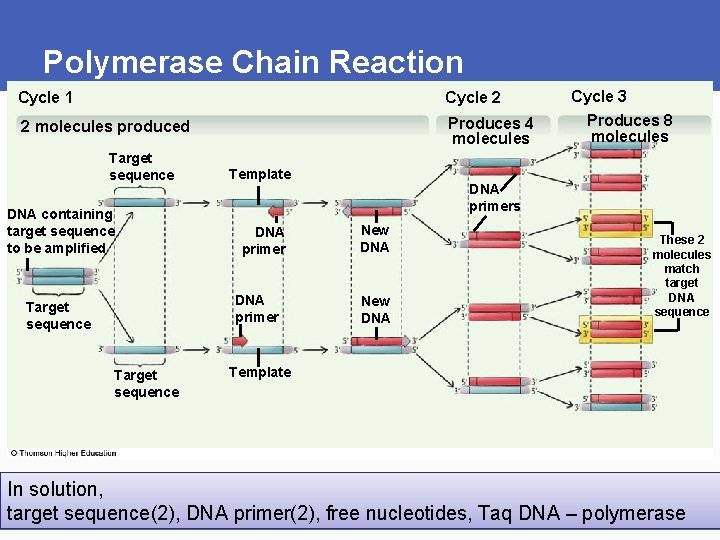
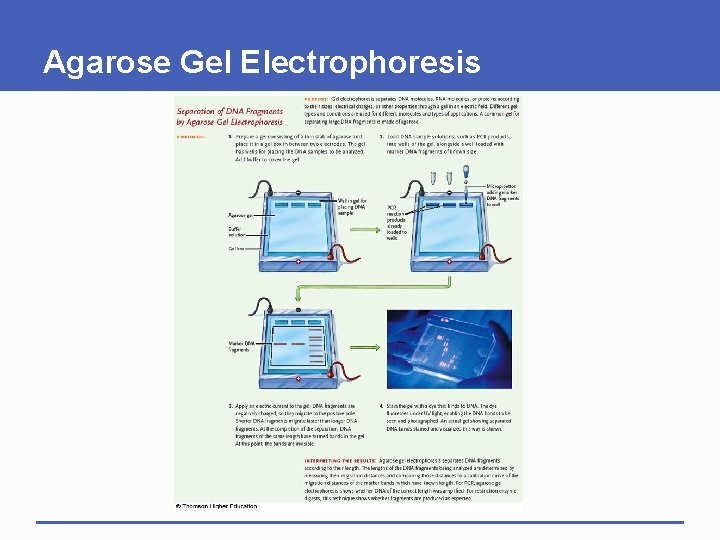
Polymerase Chain Reaction § Polymerase chain reaction (PCR) • Produces many sequence copies without host cloning • Amplifies known DNA sequences for analysis • Only copies sequence of interest • Primers bracket sequence § Agarose gel electrophoresis • Separates fragments by size and charge • Gel molecular sieve

Polymerase Chain Reaction Cycle 1 Cycle 2 2 molecules produced Produces 4 molecules Target sequence DNA containing target sequence to be amplified Template DNA primer Target sequence Cycle 3 Produces 8 molecules DNA primers New DNA These 2 molecules match target DNA sequence Template In solution, target sequence(2), DNA primer(2), free nucleotides, Taq DNA – polymerase

Agarose Gel Electrophoresis

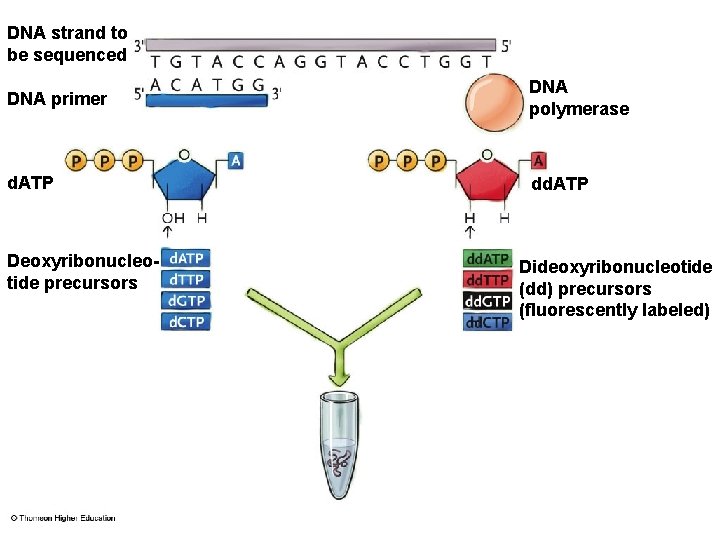
DNA Sequencing § Used for small DNA sequences to genomes § Dideoxy (Sanger) method of sequencing • Dideoxyribonucleotides have –H bound to 3’ C instead of –OH • DNA polymerases place dideoxyribonucleotides in DNA, stops replication • Polyacrylamide gel separates strands varying by one nucleotide

Dideoxy (Sanger) Method

DNA strand to be sequenced DNA primer DNA polymerase d. ATP dd. ATP Deoxyribonucleotide precursors Dideoxyribonucleotide (dd) precursors (fluorescently labeled)

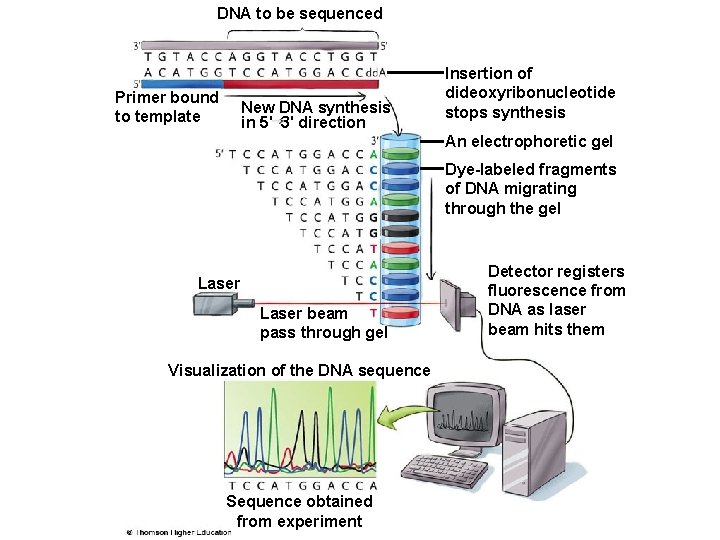
DNA to be sequenced Primer bound to template New DNA synthesis in 5' 3' direction Insertion of dideoxyribonucleotide stops synthesis An electrophoretic gel Dye-labeled fragments of DNA migrating through the gel Laser beam pass through gel Visualization of the DNA sequence Sequence obtained from experiment Detector registers fluorescence from DNA as laser beam hits them

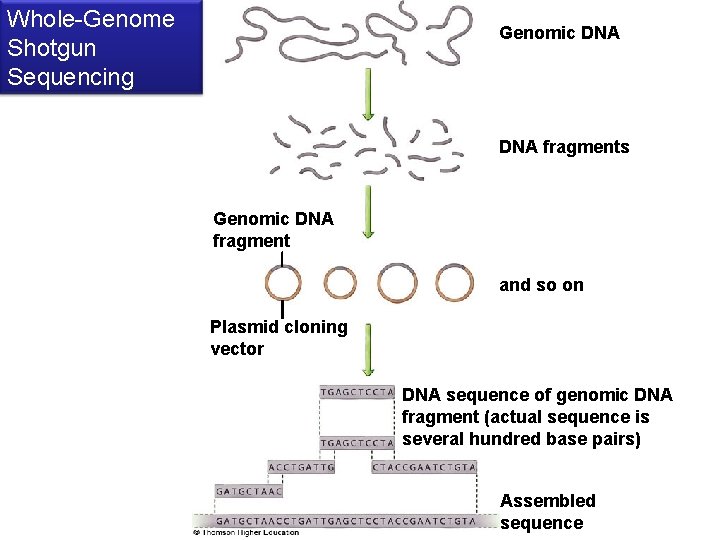
Whole-Genome Shotgun Sequencing Genomic DNA fragments Genomic DNA fragment and so on Plasmid cloning vector DNA sequence of genomic DNA fragment (actual sequence is several hundred base pairs) Assembled sequence