Database Searches Guoqing Lu Office E 115 Beadle

Database Searches Guoqing Lu Office: E 115 Beadle Center Tel: (402) 472 -4982 Email: glu 3@unl. edu Website: http: //biocore. unl. edu

Motivation • Find which sequences in the database are related to your sequence • X is like A • A has function M X is likely to have function M Applications include: • identifying orthologs and paralogs • discovering new genes or proteins • discovering variants of genes or proteins • investigating expressed sequence tags (ESTs) • exploring protein structure and function

Concerns in Database Searches • Sensitivity – The ability of a search method to find most of the members of the protein family represented by the query sequence • Selectivity – The ability of a search method to locate a protein family without making a false-positive classification of members of other families. • Speed – Heuristic, not the optimal but practical

A Naïve Approach • For each target sequence: – Align query to target sequence using e. g. , dynamic programming – Report alignment if above cutoff • Repeat for the next sequence

FASTA • Developed by Lipman & Pearson (1985) – Search for matching sequence patterns or words, or k-tuples • Local alignment – Tries to find paths of regional similarity, rather than trying to find the best alignment between 2 sequences • Heuristic – Not guaranteed to find the best alignment between 2 sequences; it may miss matches – Uses a strategy which is expected to find most matches, but sacrifices complete sensitivity in order to gain speed • Has gone through a series of updates and enhancements leading to version 3, denoted FASTA 3 FTP: ftp. virginia. edu/pub/

FASTA Algorithm – Step 1 • Identify regions shared by two sequences with highest density of identities – 4 -6 for nucleotide searches – 2 for protein – Merge along the diagonals

FASTA Algorithm – Step 2 • Re-calculate INIT 1 using scoring matrix, e. g. , PAM 250 • Keep up top 10 scoring segments • Each segment is a partial alignment without gaps

FASTA Algorithm – Step 3 • Merge INIT 1 regions that pass a threshold by allowing gaps between them • INITN score is sum of INIT 1 scores minus gaps

FASTA Algorithm – Step 4 • Using dynamic programming to optimize the alignment in a narrow band that encompasses the top scoring segments • OPT score

FASTA – Statistics • FASTA calculates a z-score for the sequence pair by multiplying the alignment score by ln[(length(query)/length(db_sequence)] • Using the distribution of the z-score, the program can estimate the number of sequences that would be expected to produce, purely by chance, a z-score greater than or equal to the z-score obtained in the search. This is reported as the E value
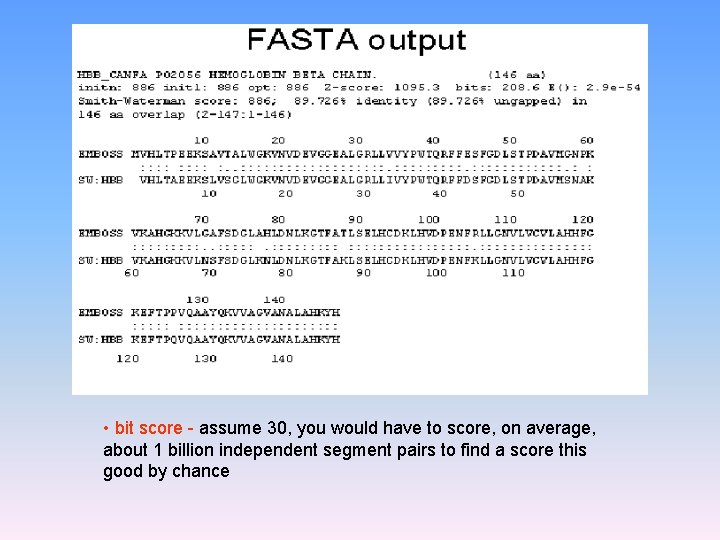
• bit score - assume 30, you would have to score, on average, about 1 billion independent segment pairs to find a score this good by chance

TFASTA • Used to search a DNA database using a protein query sequence • Find any DNA sequences that may code for a protein of interest TFASTA is very slow !!! http: //www. ebi. ac. uk/services/index. html

BLAST • Basic Local Alignment Search Tool • Developed as a way to perform a sequence similarity search by an algorithm that is faster than FASTA while being as sensitive (Altschul et al 1990, 1994, 1997)


BLAST P-P: 7 Q-Q: 5 G-G: 6 … Build NWL & Search Database for NWH
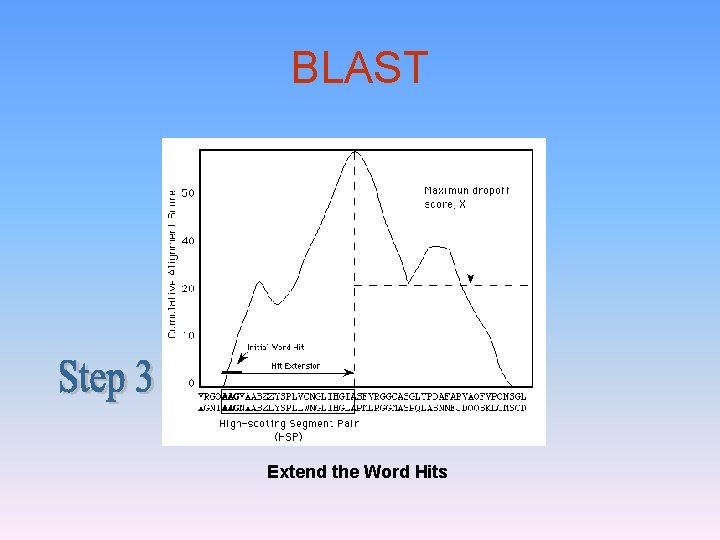
BLAST Extend the Word Hits

How to Interpret a BLAST Search: Expect Value The expect value E is the number of alignments with scores greater than or equal to score S that are expected to occur by chance in a database search. The key equation describing an E value is: E = Kmn e-l. S

E = Kmn e-l. S This equation is derived from a description of the extreme value distribution S = the score E = the expect value = the number of HSPs expected to occur with a score of at least S m, n = the length of two sequences l, K = Karlin Altschul statistics

From Raw Scores to Bit Scores • There are two kinds of scores: – raw scores (calculated from a substitution matrix) and bit scores (normalized scores) • Bit scores are comparable between different searches because they are normalized to account for the use of different scoring matrices and different database sizes S’ = bit score = (l. S - ln. K) / ln 2 The E value corresponding to a given bit score is: E = mn 2 -S’ Bit scores allow you to compare results between different database searches, even using different scoring matrices.

How to Interpret BLAST: E values and p values A p value is a different way of representing the significance of an alignment. p = 1 - e -E E 10 5 2 1 0. 05 0. 001 0. 0001 p 0. 99995460 0. 99326205 0. 86466472 0. 63212056 0. 09516258 (about 0. 1) 0. 04877058 (about 0. 05) 0. 00099950 (about 0. 001) 0. 0001000

• Blastp – Compares an amino acid query sequence against a protein sequence database. • Blastn – Compares a nucleotide query sequence against a nucleotide sequence database. • Blastx – Compares a nucleotide query sequence translated in all reading frames against a protein sequence database. • Tblastn - Compares a protein query • sequence against a nucleotide sequence database dynamically translated in all reading frames. Tblastx - Compares the six-frame translations of a nucleotide query sequence against the six -frame translations of a nucleotide sequence database.
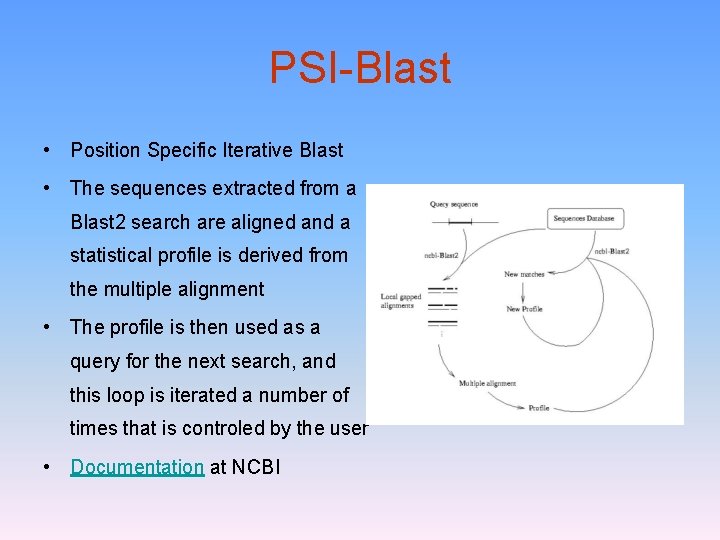
PSI-Blast • Position Specific Iterative Blast • The sequences extracted from a Blast 2 search are aligned and a statistical profile is derived from the multiple alignment • The profile is then used as a query for the next search, and this loop is iterated a number of times that is controled by the user • Documentation at NCBI

PHI-Blast • Pattern-Hit Initiated Blast • The search space is restricted to the database sequences that match a motif, which is specified by the user and has also to be contained in the query sequence. • The resulting alignment is anchored to this motif.

Comparison of BLAST and FASTA Algorithms • Both: heuristic • BLAST: several alignments per database entry FASTA: one alignment per database entry • Word length: BLAST: 11, FASTA: 6 Therefore, FASTA may be better for DNA seqs • BLAST treats automatically low complexity sequences • Both provide: Ranking of alignments Alignment scores Statistical significance of alignments Alignments

Scoring VLSPADKTNVKAAWGKVGA ||| | | || VLSEGEWQLVLHVWAKVEA • The alignment score = Matches - penalty for gap extension Do matches (V, V), (L, L), (S, S) have the same value? ? ?

PAM Matrices • Point Accepted Mutation Matrices (Dayhoff 1978) – List the likelihood of change from one base or amino acid to another in homologous sequences during evolution • • AA PAM matrices are derived from families of closely related sequences. • Likelihood (odds) ratio for residues a and b: Evolutionary distance of 1 PAM = probability of 1 point mutation per 100 residues – Probability a-b is a mutation / probability a-b is chance • PAM matrices contain log-odds figures – Val>0: likely mutation – Val=0: random mutation – Val<0: unlikely mutation • • • 250 PAM: similarity scores equivalent to 20% identity Low PAM – good for finding short, strong local similarities High PAM - long weak similarity


BLOSUM Matrices • Blocks Substitution Matrices (Henikoff & Henikoff 1992) • Aligned, ungapped conserved region of a protein family • Calculate the frequency with which any amino acid can appear at each position • Compute the probability that any amino acid can substitute for any other • Frequencies obtained from protein blocks constructed regardless of evolutionary distance • Blocks represent regions of conserved sequence similarities • Conservation due to functional constraints, thus calculated frequencies reflect functional constraints • Much larger data set used than for the PAM matrix • BLOSUM 64 is roughly equivalent to PAM 120

Exercise 1 • Search the following sequence against Swiss. Prot database for similarity using Fasta and NCBI-Blast 2 programs at http: //www. ebi. ac. uk/services/index. html > unknown protein MAVACAVAVRPLVQVAVASAVSTAAPASSKPAVKLAASAVSAVALTTVSVSAGLLATTAVEDPRFHAADCQS RSADASASCEDLQPSTSTCTSAVRDANRPTRRVRRSGSKAQRRGSTTLTASVPSMAAAVVLPPKIALRRRHR LRLRAGHSATAAATDKTPREQPDKPAALPEDLLPADATSTSSTGKISSAAVCCGLLAHCSAAQLHAILCGLV QAVASSSVKGNNRKLLLGSKLRKLLEGVGVAPANGKAYTAADVAALSGPKLERLRATLKSQPGLLLWFLLFT APAKLQALQAALLPGGAGDRSFEEWRAAIDAVAGSGHEQLAAAQEVRGRQSACVEGSTAGNTATTATITTTN NNPASHGGVYTALTGTEVTGKKPAALPEDLLPADATSTSSTGKISSAAVCCGLLAHCSAAQLHAILCGLVQA VASSSVKGNNRKLLLGSKLRKLLEGVGVAPANGKAYTAADVAALSGPKLERLRATLKSQPGLLLWFLLFTAP AKLQALQAALLPGGAGDRSFEEWRAAIDAVAGSGHEQLAAAQEVRGRQSACVEGSTAGNTATTATITTTNNN PASHGGVYTALTGTEVTGKAAANKDLSRTRTTSHRNRCVSESGSTRNKSRSSSTHSVEYAEPKAGCS QPAATVPGCVPEIISAAIPPLAPLALHIRRAIVKELLEARPPGWNTFLYSWLQAAGLSEFLPANGTCRMYMA DRKQLVLRVGAMREEQVDAFLTCMCKAHGHSTWLARYLHMLGPEVSQLLS

GCG Introduction

GCG entry/data format • Entry – database: accessionnumber – genbank: zmzein; gb: * • The format of records – NCBI – EMBL • Editing – seqed [filename] – reformat [filename] – Reverse [filename]
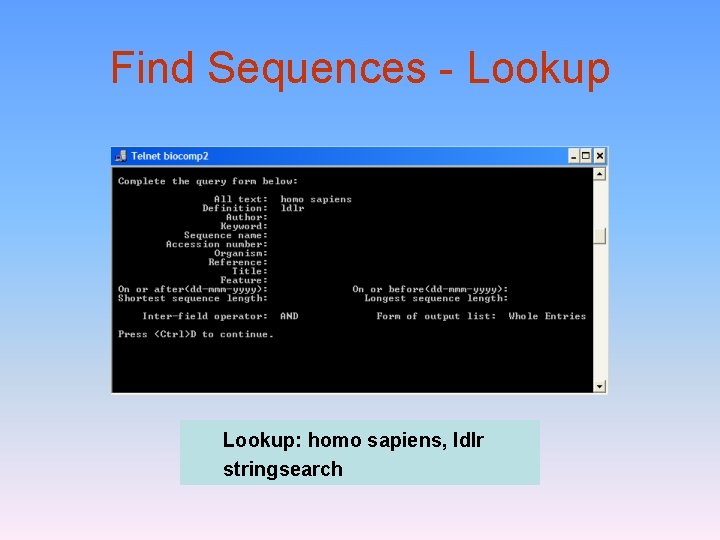
Find Sequences - Lookup: homo sapiens, ldlr stringsearch
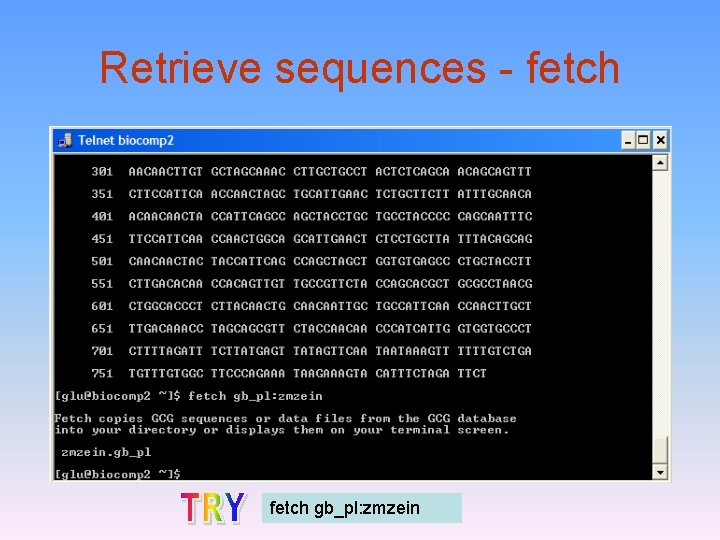
Retrieve sequences - fetch gb_pl: zmzein
![Sequence edit - seqed Ctrl+D : seqed zmzein. gb_pl reformat [filename] reverse [filename] Sequence edit - seqed Ctrl+D : seqed zmzein. gb_pl reformat [filename] reverse [filename]](http://slidetodoc.com/presentation_image_h2/1b3d8152a82641d84dcaaf0ca3172620/image-34.jpg)
Sequence edit - seqed Ctrl+D : seqed zmzein. gb_pl reformat [filename] reverse [filename]
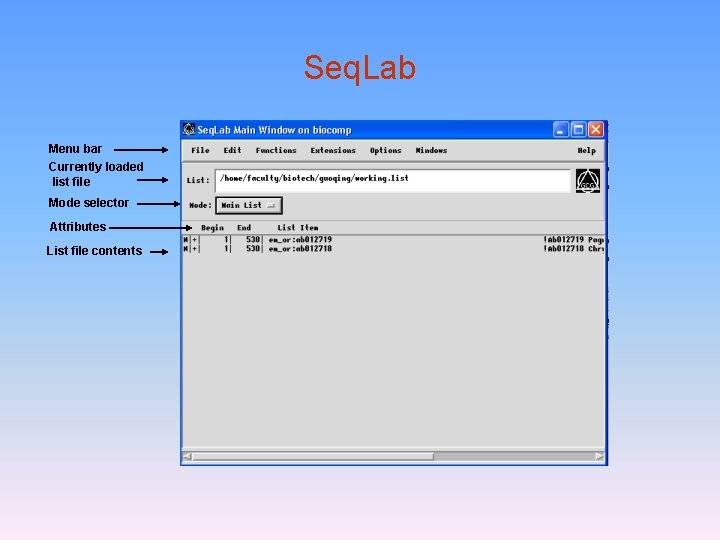
Seq. Lab Menu bar Currently loaded list file Mode selector Attributes List file contents
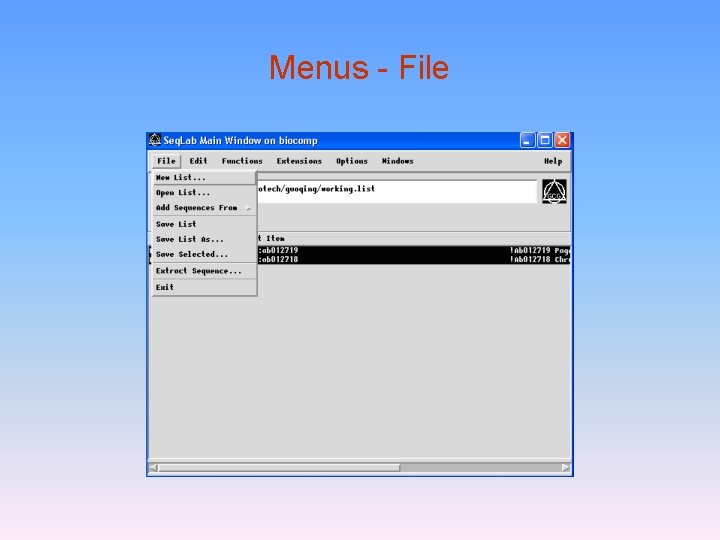
Menus - File
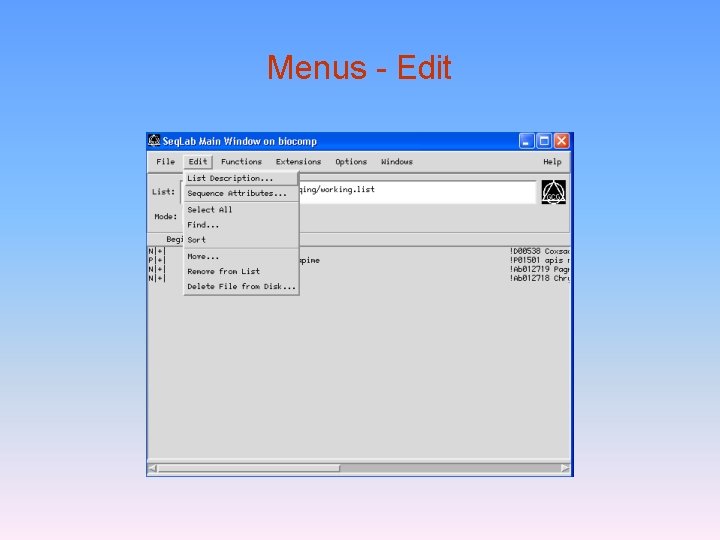
Menus - Edit
Menus - Functions
Menus - Options
Menus - Windows
Add sequence from databases
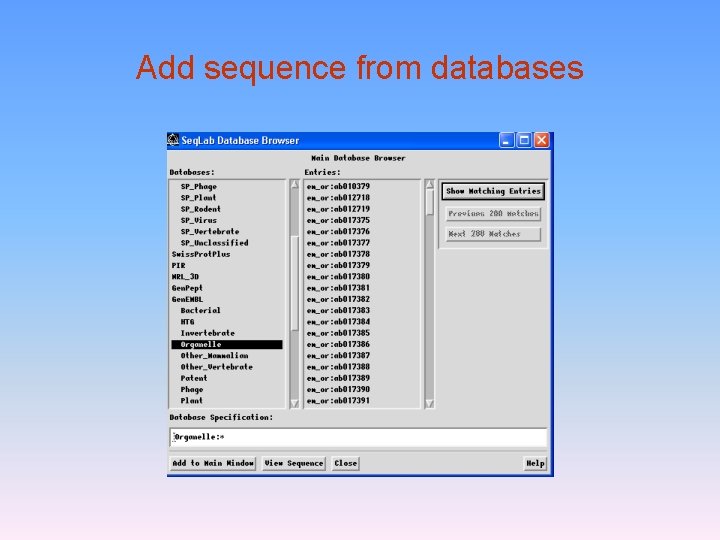
Add sequence from databases
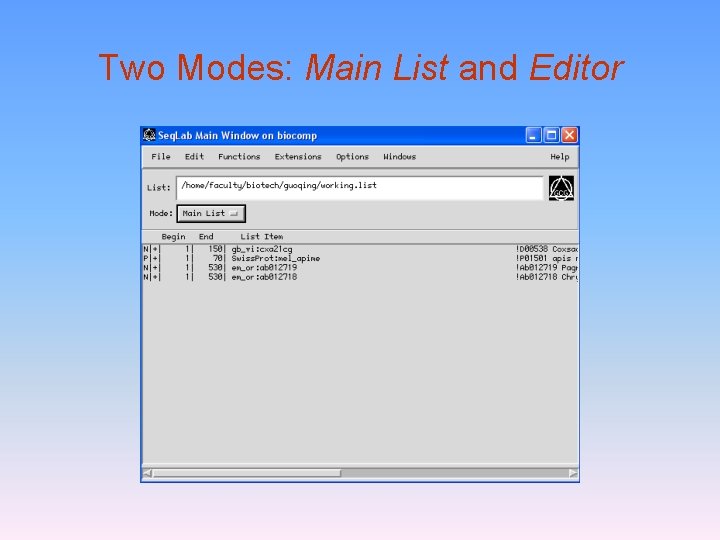
Two Modes: Main List and Editor
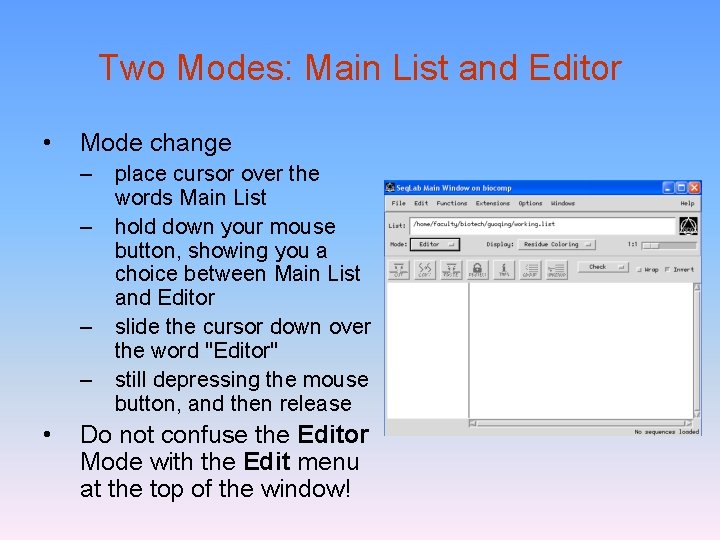
Two Modes: Main List and Editor • Mode change – place cursor over the words Main List – hold down your mouse button, showing you a choice between Main List and Editor – slide the cursor down over the word "Editor" – still depressing the mouse button, and then release • Do not confuse the Editor Mode with the Edit menu at the top of the window!

Adding files to Main List / Editor • Three kinds of files – files from your Unix directory – files in the sequence databanks – files you've retrieved from databanks on the net into your directory • If you have a file in Fasta format (a single line with a ">" sign and the name of the sequence, followed by as many lines of sequence letters), you can, in Editor Mode only, Import sequence from the File menu
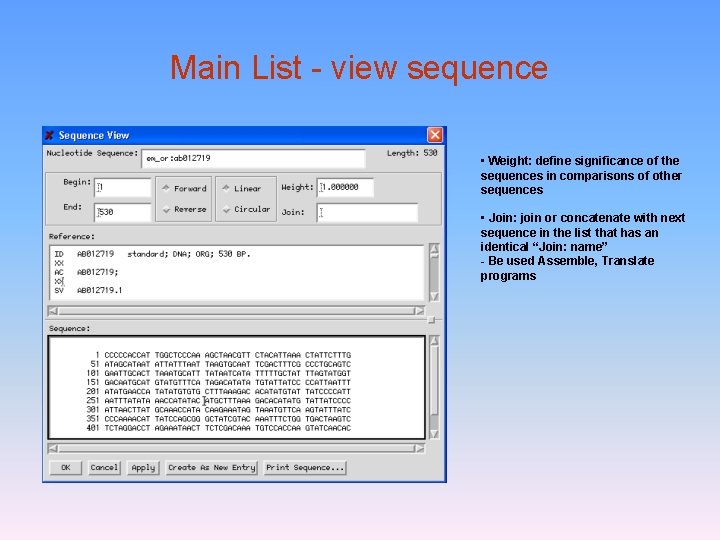
Main List - view sequence • Weight: define significance of the sequences in comparisons of other sequences • Join: join or concatenate with next sequence in the list that has an identical “Join: name” - Be used Assemble, Translate programs
Editing sequence • Editor – Cut, Copy, and Paste – Lock, Group, and un. Group
- Slides: 47