Coding for DNA storage Shubham Chandak Kedar Tatwawadi

Coding for DNA storage Shubham Chandak, Kedar Tatwawadi EE 388 course project

Outline • DNA storage model • Capacity computation • Two coding strategies • Experimental results • Conclusion and future work

DNA storage • DNA as a storage medium • High density: 215 petabytes/gram 1 • High durability 2 • Synthesis (writing) and sequencing (reading) error‐prone and expensive • Error correction needed for reliable data recovery 1. Erlich, Y. , & Zielinski, D. (2017). DNA Fountain enables a robust and efficient storage architecture. Science, 355(6328), 950‐ 954. 2. Grass, R. N. , Heckel, R. , Puddu, M. , Paunescu, D. , & Stark, W. J. (2015). Robust Chemical Preservation of Digital Information on DNA in Silica with Error‐Correcting Codes. Angewandte Chemie International Edition, 54(8), 2552‐ 2555.
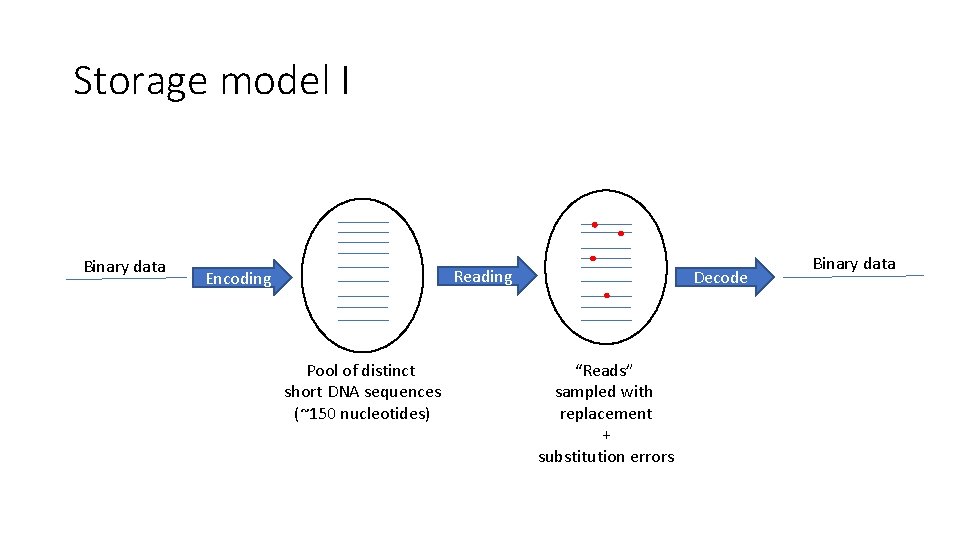
Storage model I Binary data Encoding P Pool of distinct short DNA sequences (~150 nucleotides) Reading P “Reads” sampled with replacement + substitution errors Decode Binary data

Storage model II • For this talk: • Ignore DNA symbols and constraints – work with binary sequences • Assume that the index of each sequence is transmitted without error
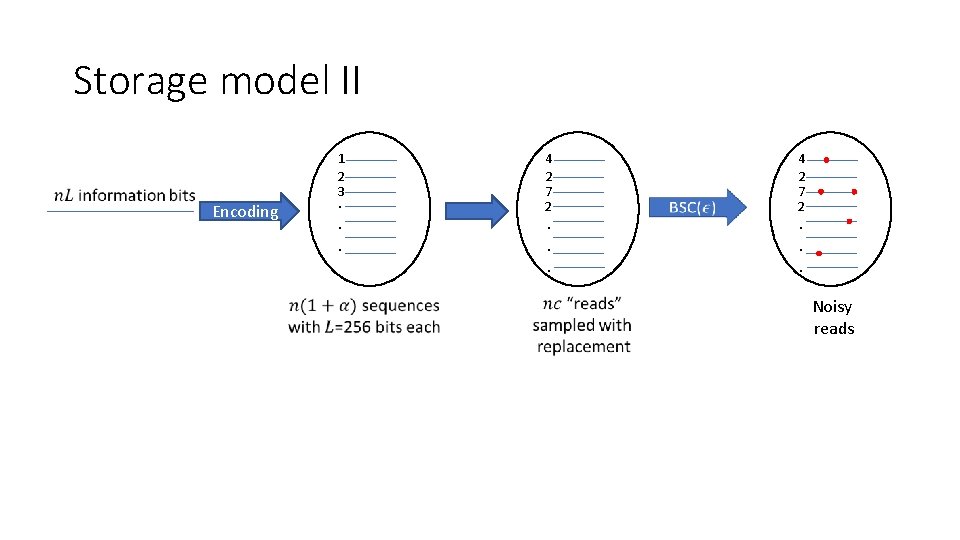
Storage model II 1 2 3 Encoding . . . P 4 2 7 2 . . . P Noisy reads
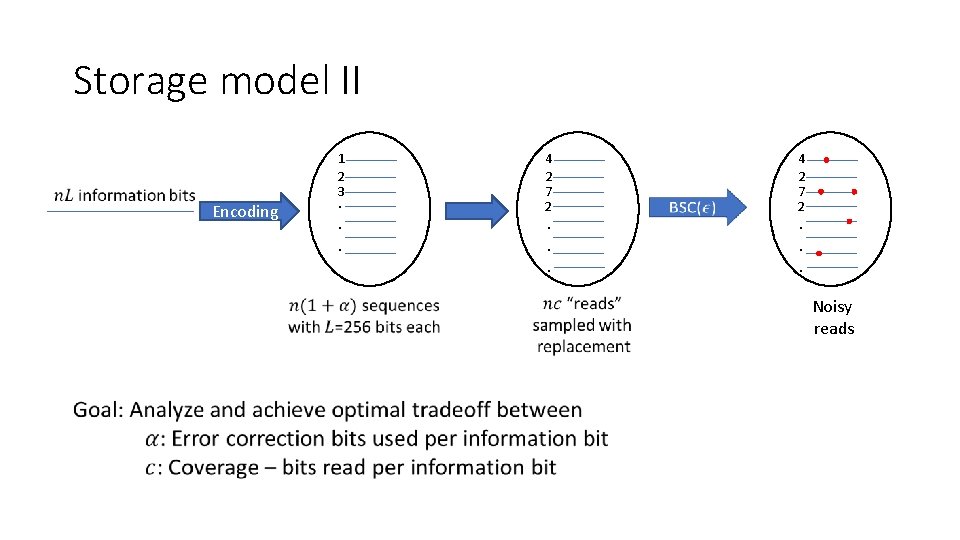
Storage model II 1 2 3 Encoding . . . P 4 2 7 2 . . . P Noisy reads

Capacity: error free reads •

Capacity: error free reads •


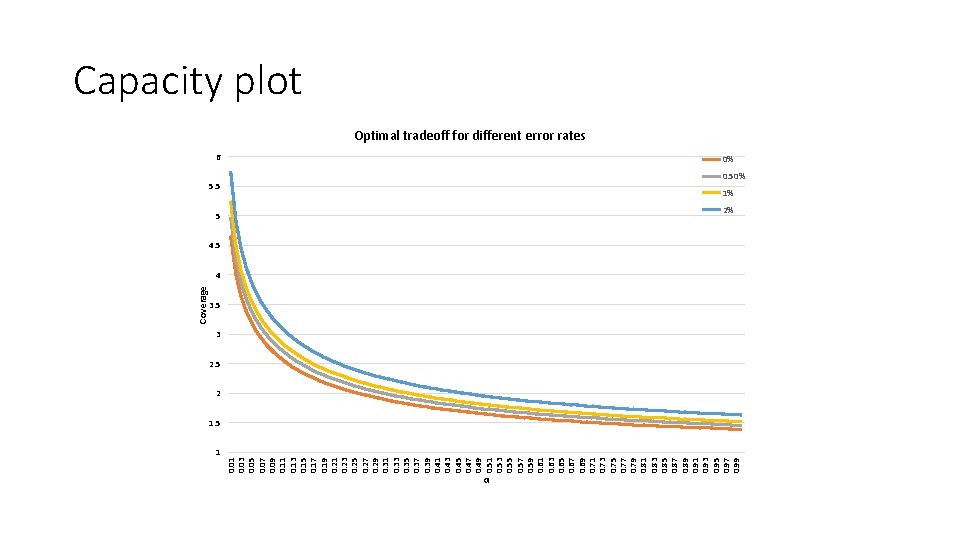
1 0. 03 0. 05 0. 07 0. 09 0. 11 0. 13 0. 15 0. 17 0. 19 0. 21 0. 23 0. 25 0. 27 0. 29 0. 31 0. 33 0. 35 0. 37 0. 39 0. 41 0. 43 0. 45 0. 47 0. 49 0. 51 0. 53 0. 55 0. 57 0. 59 0. 61 0. 63 0. 65 0. 67 0. 69 0. 71 0. 73 0. 75 0. 77 0. 79 0. 81 0. 83 0. 85 0. 87 0. 89 0. 91 0. 93 0. 95 0. 97 0. 99 Coverage Capacity plot Optimal tradeoff for different error rates 6 0% 5. 5 0. 50% 1% 5 2% 4. 5 4 3. 5 3 2. 5 2 1. 5 α
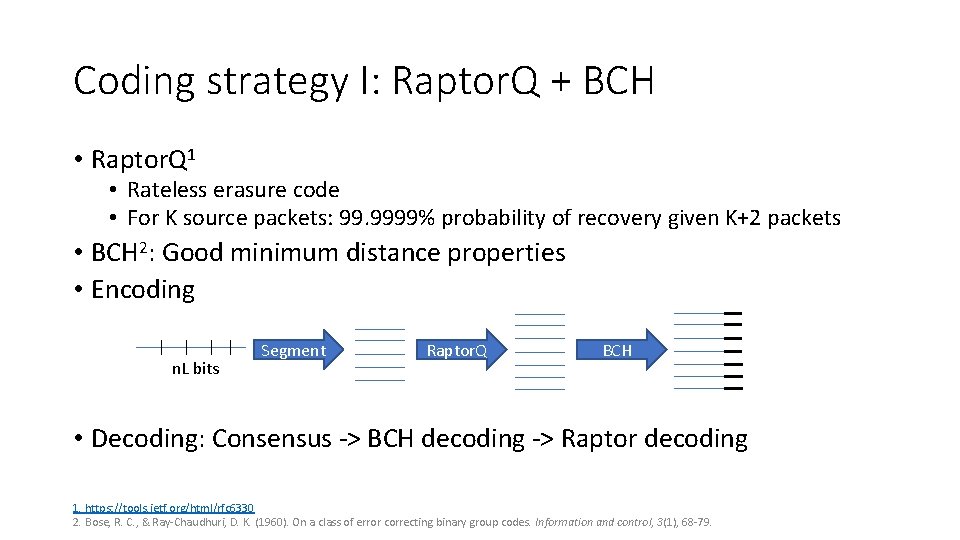
Coding strategy I: Raptor. Q + BCH • Raptor. Q 1 • Rateless erasure code • For K source packets: 99. 9999% probability of recovery given K+2 packets • BCH 2: Good minimum distance properties • Encoding n. L bits Segment Raptor. Q BCH • Decoding: Consensus ‐> BCH decoding ‐> Raptor decoding 1. https: //tools. ietf. org/html/rfc 6330 2. Bose, R. C. , & Ray‐Chaudhuri, D. K. (1960). On a class of error correcting binary group codes. Information and control, 3(1), 68‐ 79.
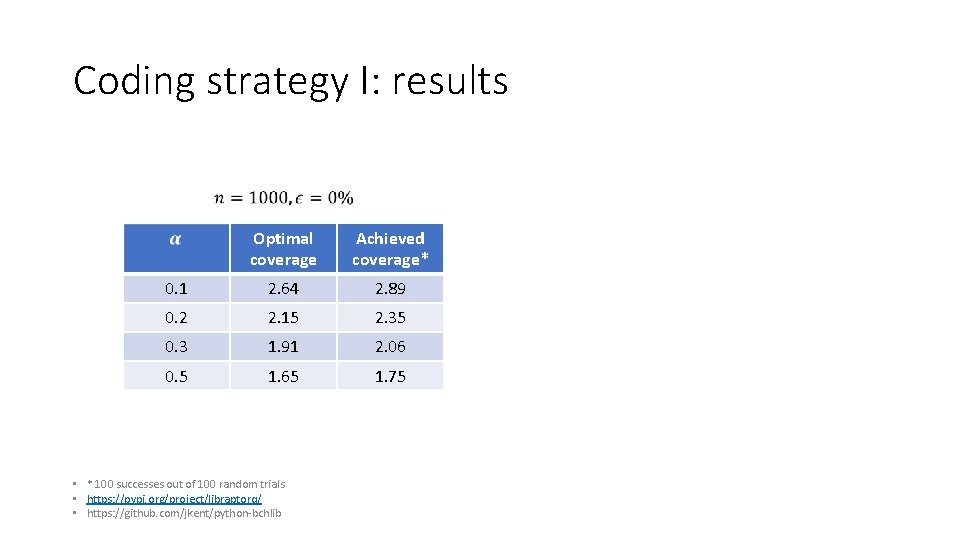
Coding strategy I: results Optimal coverage Achieved coverage* 0. 1 2. 64 2. 89 0. 2 2. 15 2. 35 0. 3 1. 91 2. 06 0. 5 1. 65 1. 75 • * 100 successes out of 100 random trials • https: //pypi. org/project/libraptorq/ • https: //github. com/jkent/python‐bchlib
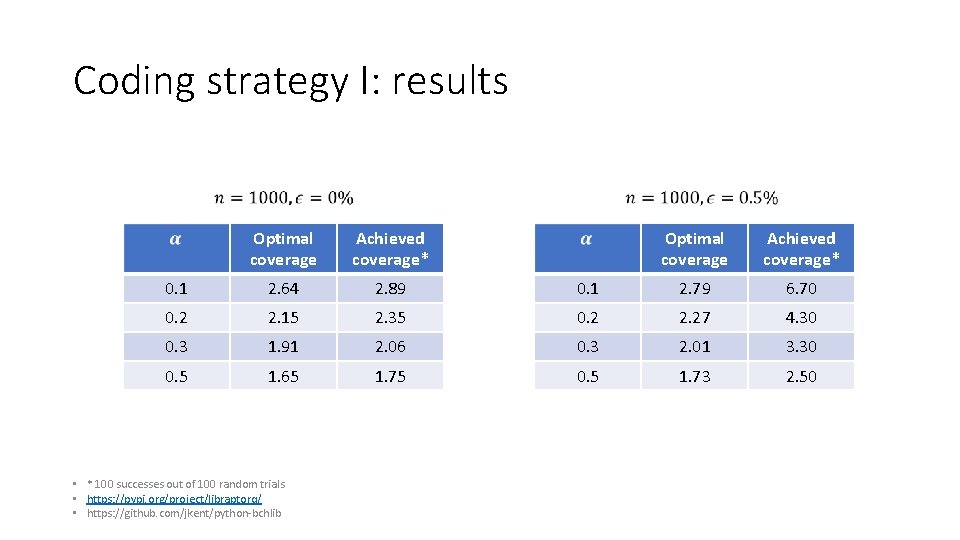
Coding strategy I: results Optimal coverage Achieved coverage* 0. 1 2. 64 2. 89 0. 1 2. 79 6. 70 0. 2 2. 15 2. 35 0. 2 2. 27 4. 30 0. 3 1. 91 2. 06 0. 3 2. 01 3. 30 0. 5 1. 65 1. 75 0. 5 1. 73 2. 50 • * 100 successes out of 100 random trials • https: //pypi. org/project/libraptorq/ • https: //github. com/jkent/python‐bchlib
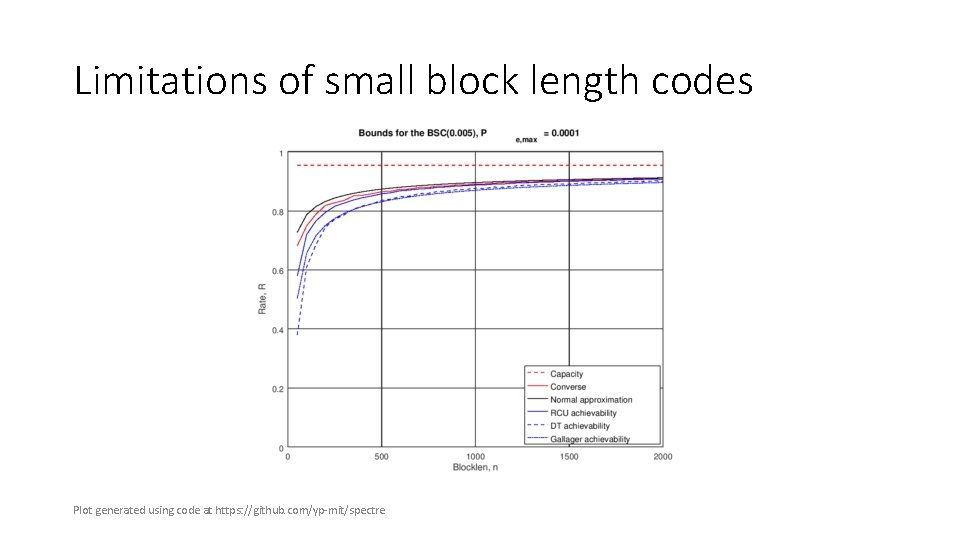
Limitations of small block length codes Plot generated using code at https: //github. com/yp‐mit/spectre

Coding strategy II: LDPC • n. L bits LDPC Segment
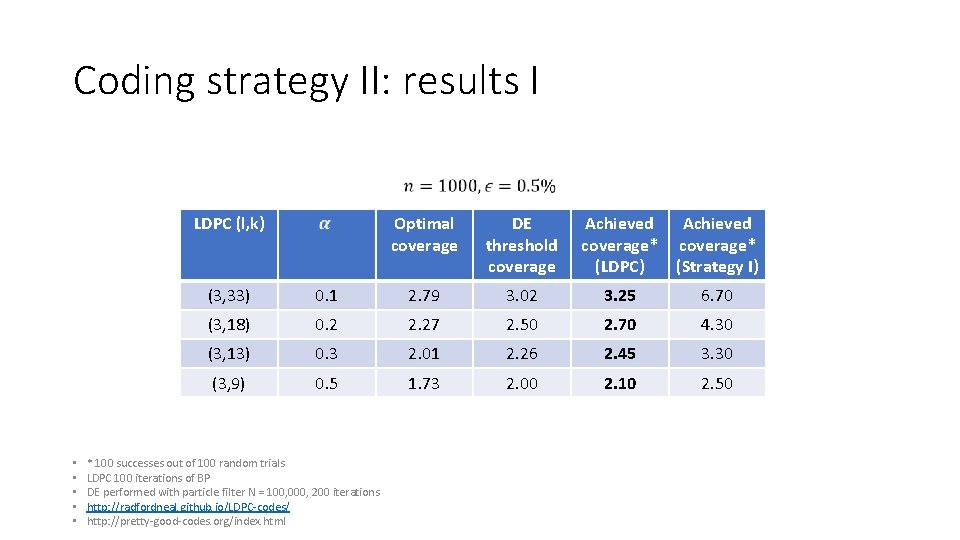
Coding strategy II: results I LDPC (l, k) • • • Optimal coverage DE threshold coverage Achieved coverage* (LDPC) Achieved coverage* (Strategy I) (3, 33) 0. 1 2. 79 3. 02 3. 25 6. 70 (3, 18) 0. 2 2. 27 2. 50 2. 70 4. 30 (3, 13) 0. 3 2. 01 2. 26 2. 45 3. 30 (3, 9) 0. 5 1. 73 2. 00 2. 10 2. 50 * 100 successes out of 100 random trials LDPC 100 iterations of BP DE performed with particle filter N = 100, 000, 200 iterations http: //radfordneal. github. io/LDPC‐codes/ http: //pretty‐good‐codes. org/index. html
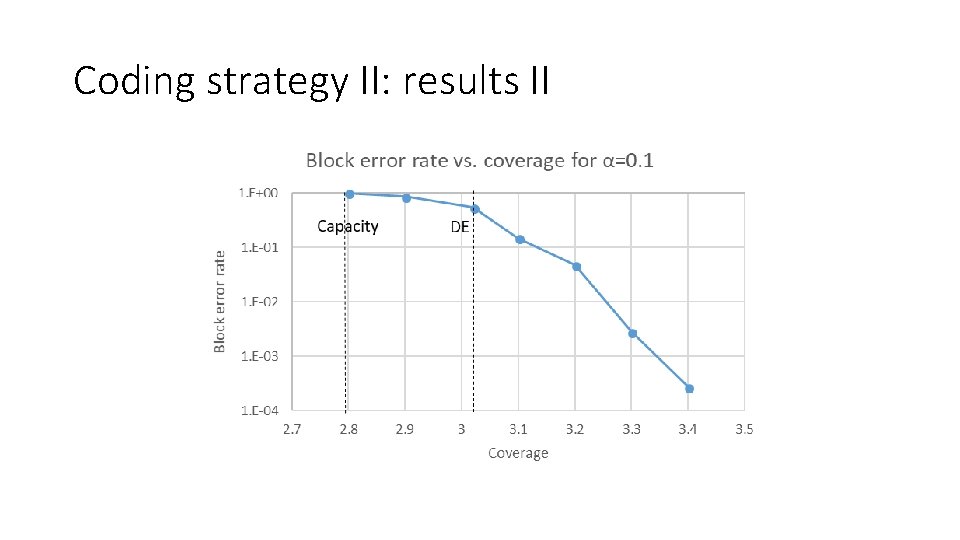
Coding strategy II: results II

Conclusion and future work • Analyzed DNA storage problem for simplified model • Implemented schemes to achieve close‐to‐optimum performance

Conclusion and future work • Analyzed DNA storage problem for simplified model • Implemented schemes to achieve close‐to‐optimum performance • Adding index to segments • Protect index with BCH code • Converting binary data to DNA symbols {A, C, G, T} • Constraint: Runs of 3 or more not allowed, e. g. , AAA • Interaction between error correction and constraint coding • Exploiting non‐IID noise in reads

Thank You!
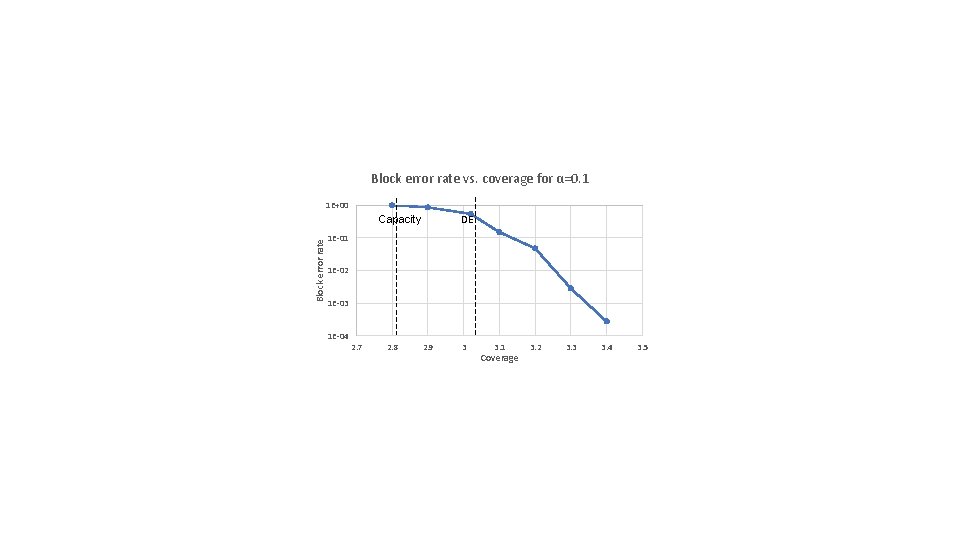
Block error rate vs. coverage for α=0. 1 1 E+00 DE Block error rate Capacity 1 E‐ 01 1 E‐ 02 1 E‐ 03 1 E‐ 04 2. 7 2. 8 2. 9 3 3. 1 Coverage 3. 2 3. 3 3. 4 3. 5
- Slides: 23