UCSC Genome Tools and Databases Jim Kent Genome

UCSC Genome Tools and Databases Jim Kent - Genome Bioinformatics Group University of California Santa Cruz
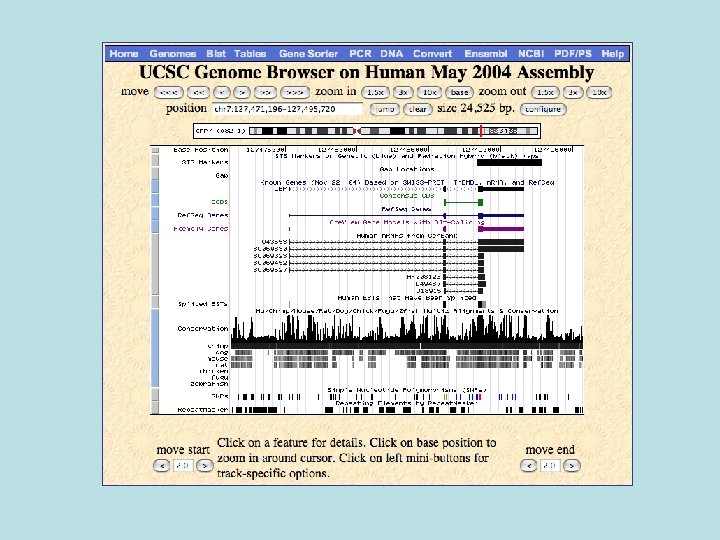

Behind the Genome Browser • ‘Genome’ database, one for each assembly of each genome. – hg 17 (human genome assembly 17) – mm 6 (mus musculus 6) – can. Fam 1 (canis familiaris 1) • hg 17 has 1616 tables, but not really – Some tables split across chromosomes for speed – 228 logical tables – Only ~30 different types of tables
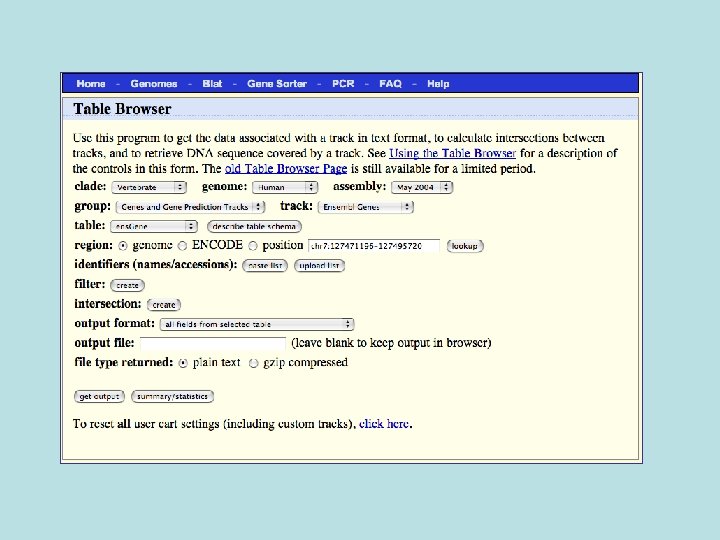
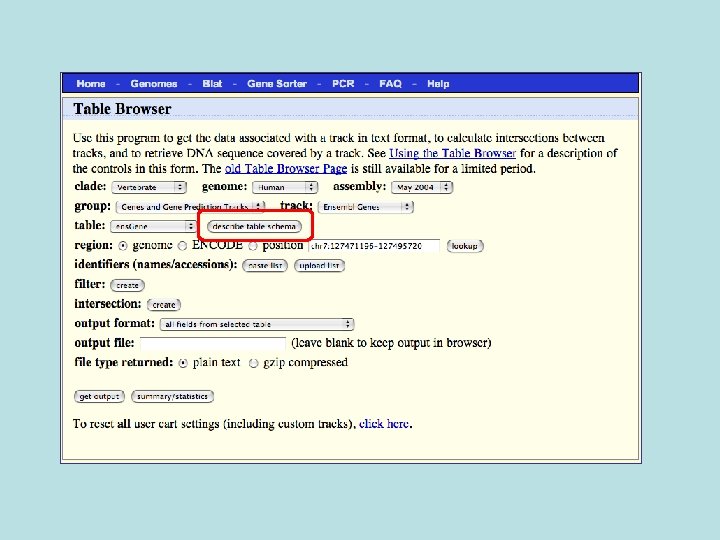
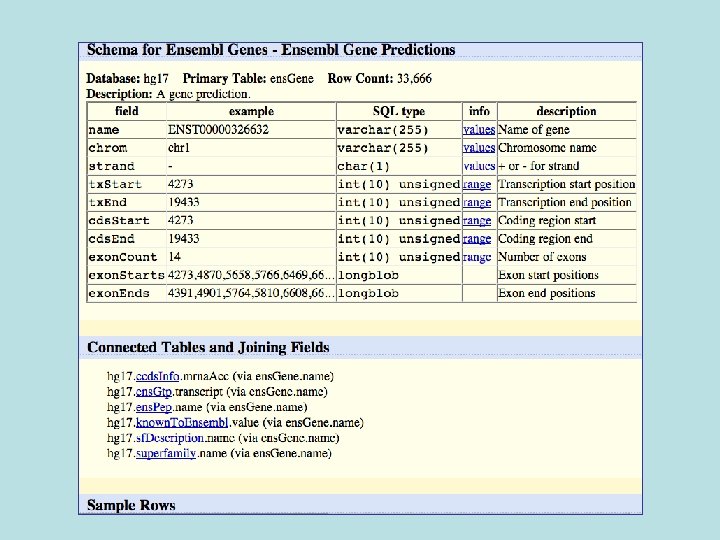
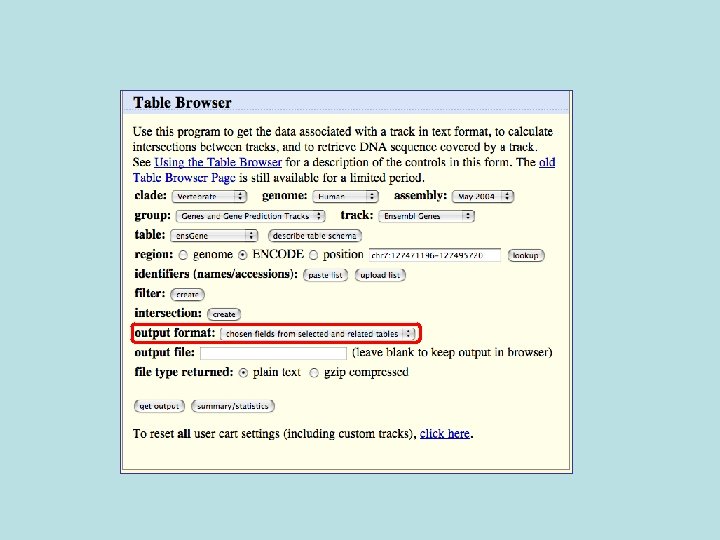
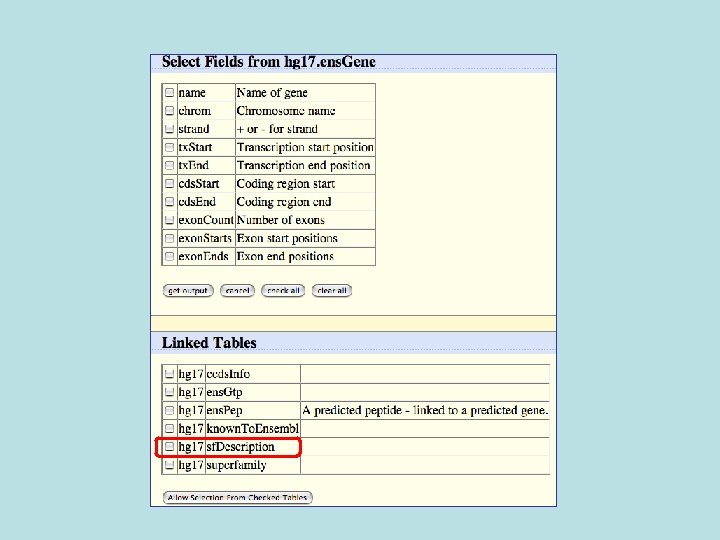
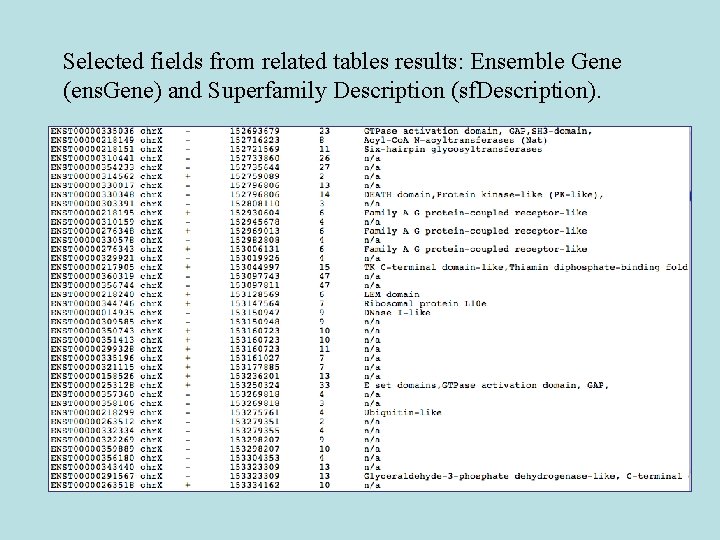
Selected fields from related tables results: Ensemble Gene (ens. Gene) and Superfamily Description (sf. Description).
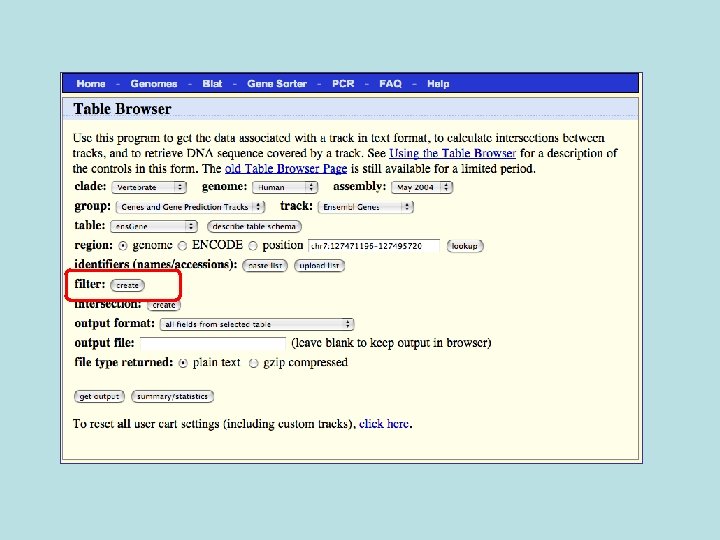
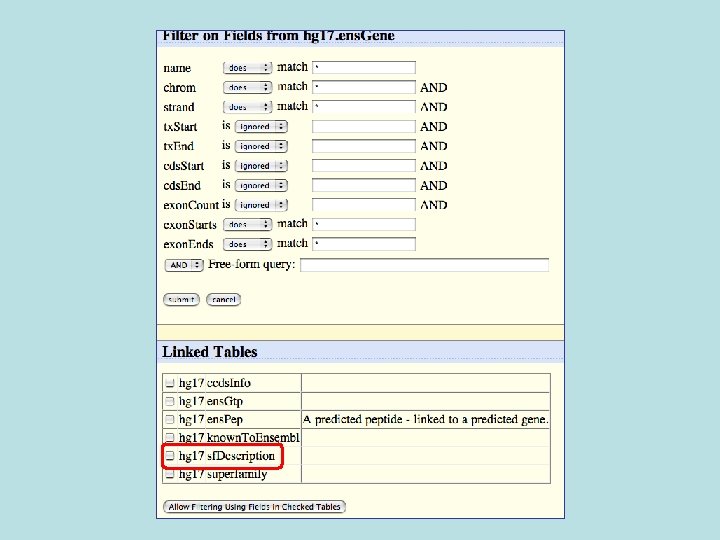
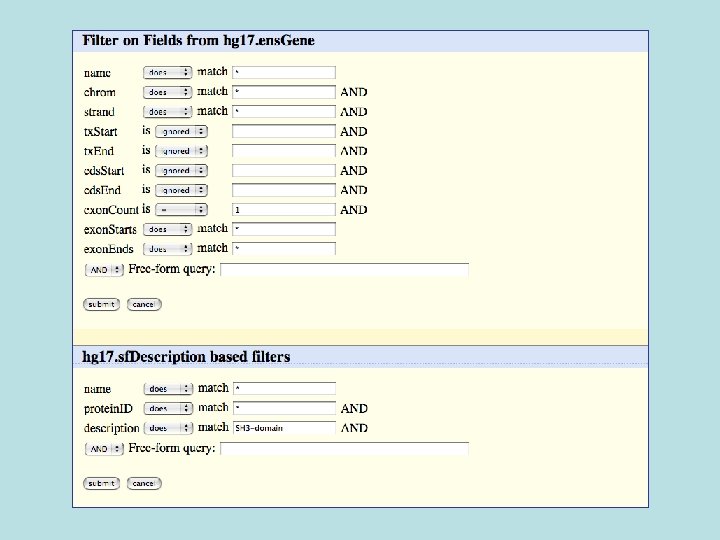
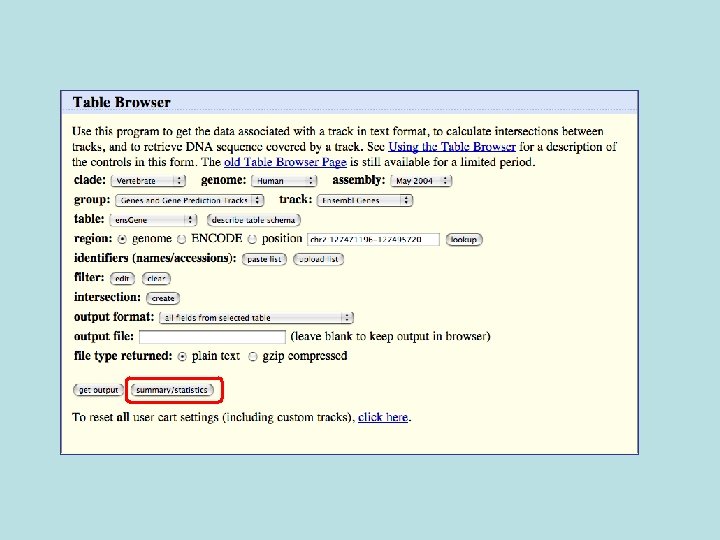
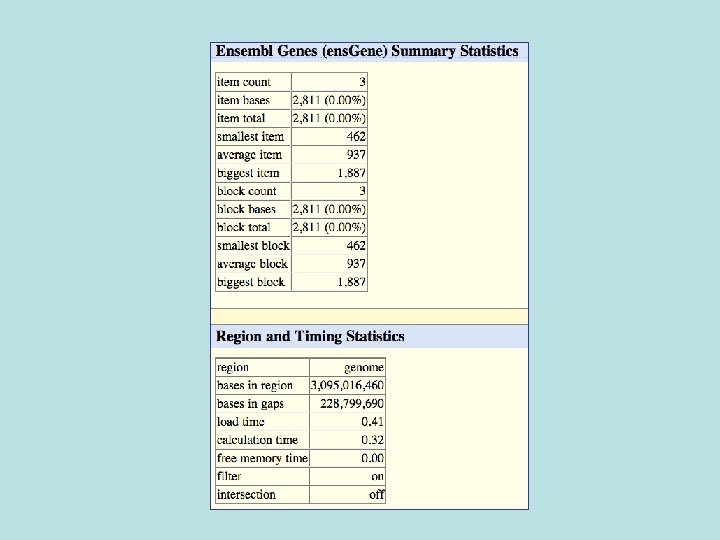
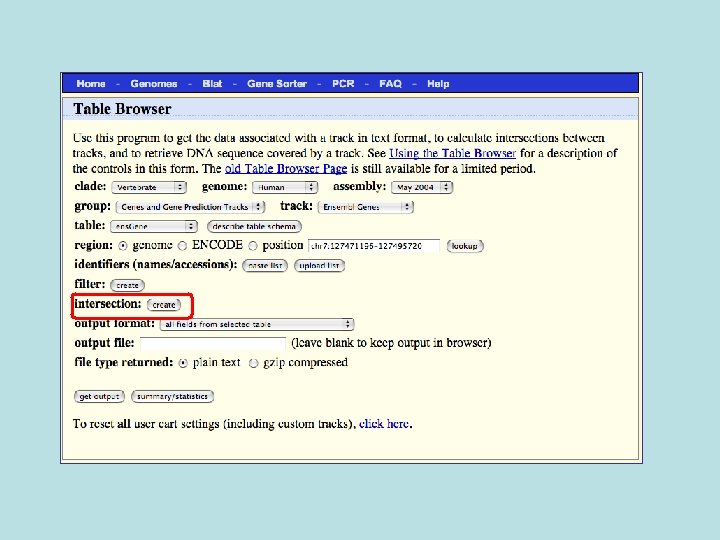
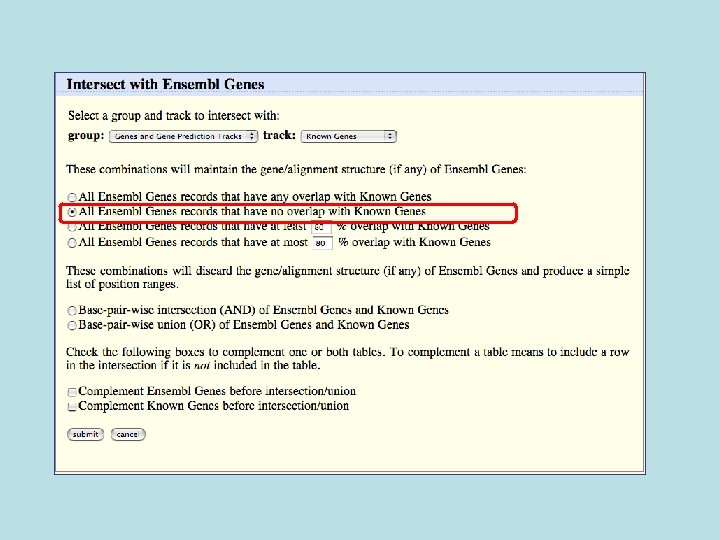

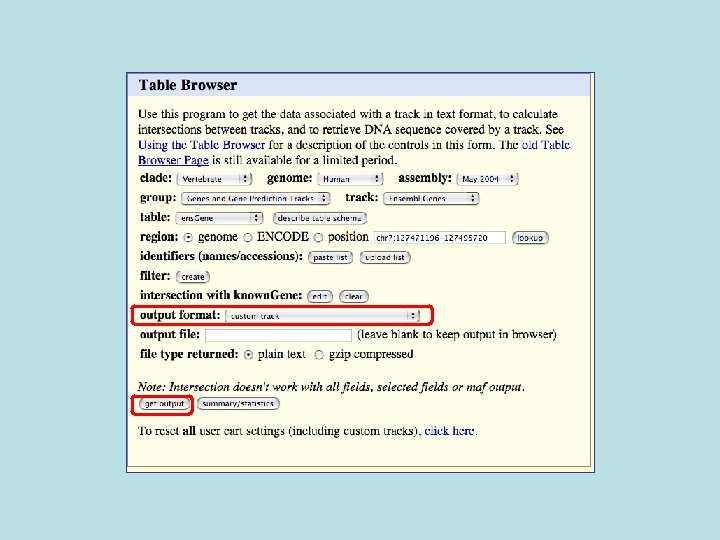
Custom Track Output • Useful for visualizing results of queries in genome browser • The way to produce more complex queries.
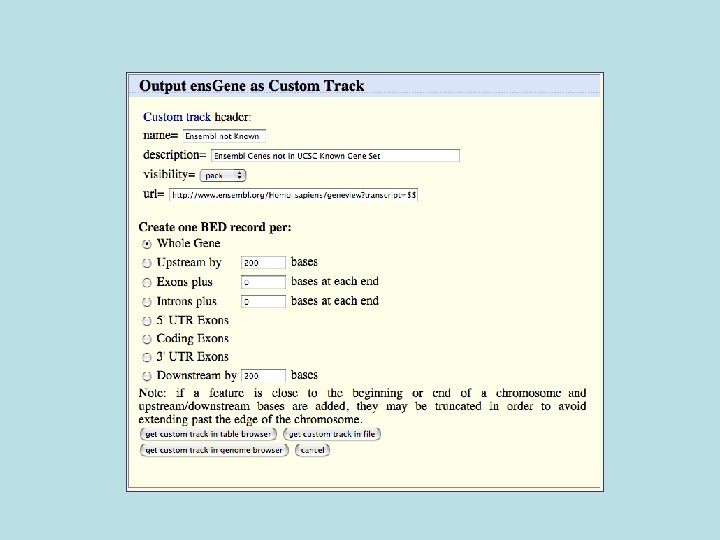
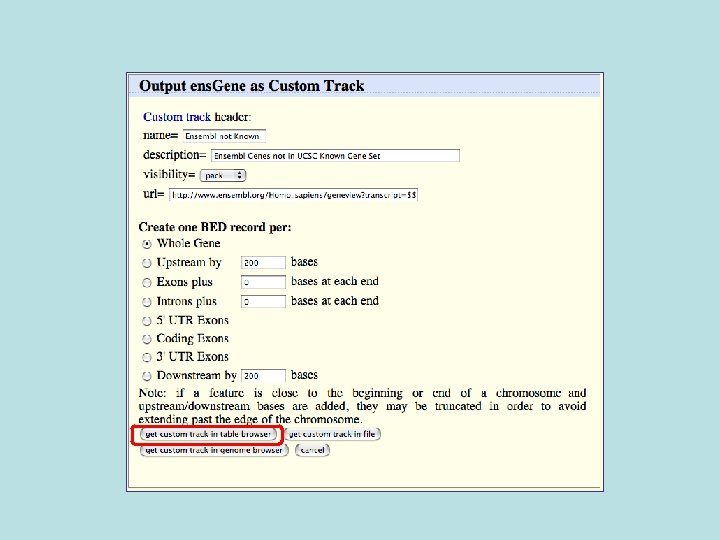
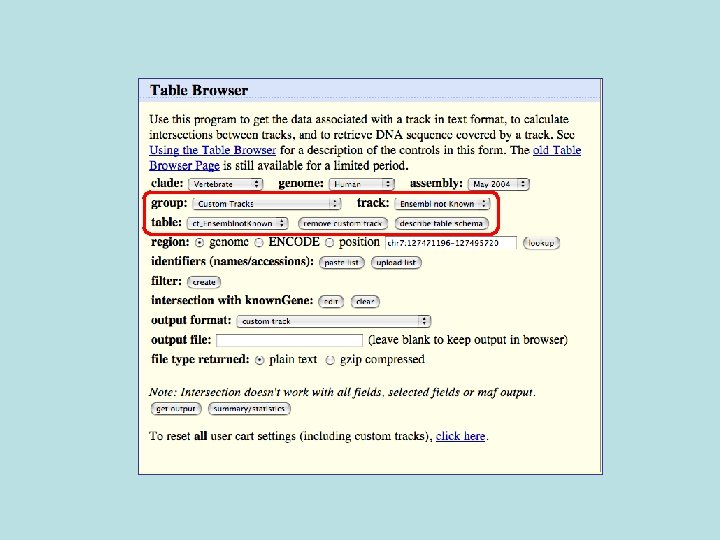


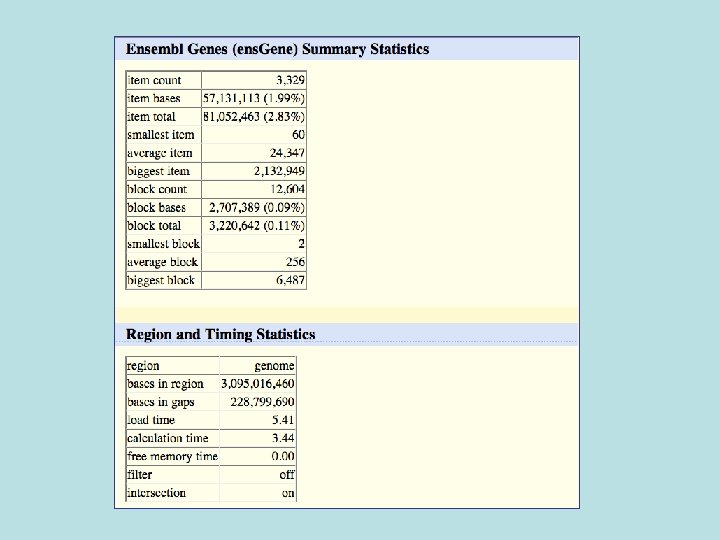
681/3329 (20%) of Ensemble not known also not conserved 1728/33, 666 (5%) of Ensembl in general not conserved

Meta-data behind Table Browser • The track. Db table describes each track. • Table and field descriptions in Auto. Sql. as files, which also generate SQL code and C code to load/save from database and tabseparated files. • Descriptions of how tables are connected in all. joiner file, which along with joiner. Check program checks database integrity.

. as Files - table and field docs table cpg. Island "Describes the Cp. G Islands" ( string chrom; "Human chromosome or FPC contig" uint chrom. Start; "Start position in chromosome" uint chrom. End; "End position in chromosome" string name; "Cp. G Island" uint length; "Island Length" uint cpg. Num; "Number of Cp. Gs in island" uint gc. Num; "Number of C and G in island" float per. Cpg; "Percentage of island that is Cp. G" float per. Gc; "Percentage of island that is C or G" ) auto. Sql generates code from these. They also help document.

all. joiner - basic example identifier softberry. Gene. Name "Link together Fshgene++ gene structure, peptide, and homolog" $gbd. softberry. Gene. name $gbd. softberry. Pep. name $gbd. softberry. Hom. name • The central concept is an identifier that appears in fields in multiple table, sometimes even multiple databases. • $gbd is a variable that contains a comma-separated list of databases. • An identifier record ends with a blank line.

# Genbank/tr. EMBL Accessions and meaningful subsets thereof identifier genbank. Accession external=genbank "Generic Genbank Accession. More specific Genbank accessions follow $gbd. seq. acc identifier bac. End. Accession type. Of=genbank. Accession "Genbank accession of a BAC end read. " $gbd. all_bacends. q. Name dupe. Ok $gbd. bac. End. Pairs. lf. Names comma $hg. fish. Clones. be. Names comma min. Check=0. 70 type. Of - allows joins between parent and child, but not between siblings. dupe. Ok - allows more than one row with same identifier in primary table comma - indicates field is comma separated list of identifiers min. Check - indicates only a portion identifiers in field is in the primary table

identifier hugo. Name external=HUGO fuzzy "International Human Gene Identifier" $hg. ref. Link. name $hg. atlas. Onco. Gene. locus. Symbol $hg. kg. Alias. alias $hg. kg. Xref. gene. Symbol $hg. ref. Flat. gene. Name $hg. jax. Ortholog. human. Symbol hg 13, hg 15. gene. Bands. name “Biological” names for human genes are so messy, no validation is done (note ‘fuzzy’ keyword).

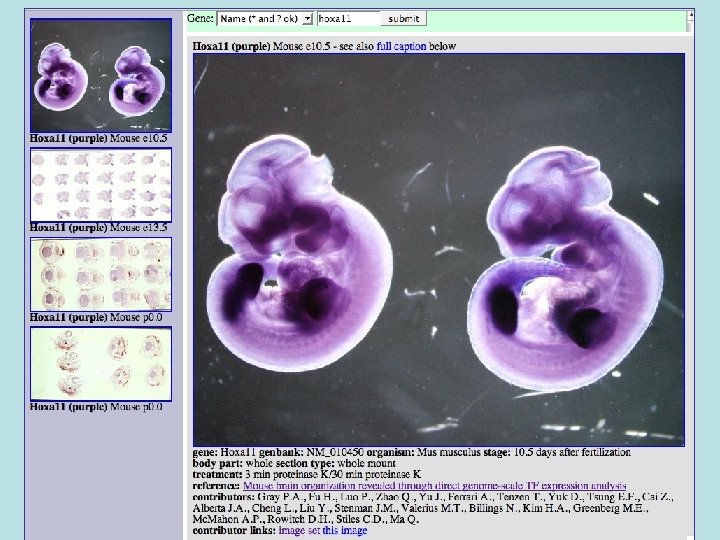
Other Databases • Genome databases - one for each assembly of each organism: hg 17, mm 6, can. Fam 1, etc. • hg. Central - home to db. Db and user settings info. One database shared by all web servers. • hg. Fixed - mostly microarray data. • uni. Prot - Relationalized Swiss. Prot/tr. EMBL database. • go - Gene ontology terms and term/gene associations. • gene. Pix - gene image database

Gene Pix • Image browser for in-situ and other geneoriented pictures • Hopefully in the long run will have a million images covering almost all vertebrate genes. • (Needs new name, Gene Pix is a microarray analysis program. Visi. Gene? )
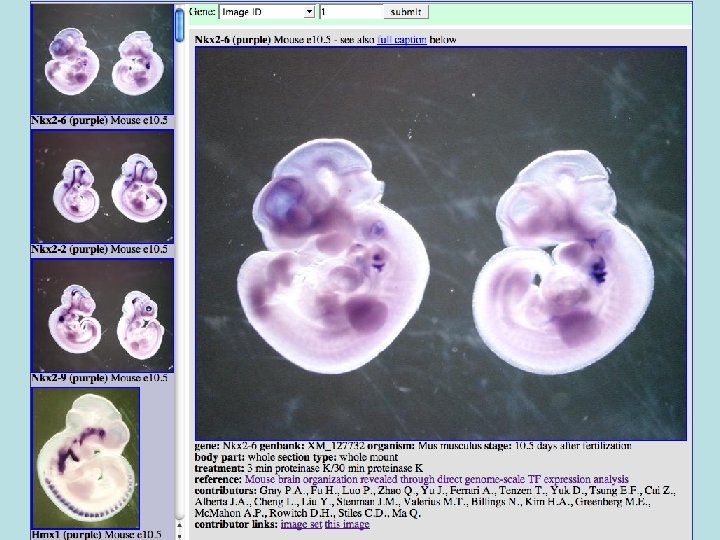

Data Sets • Paul Gray - ~1000 mouse transcription factor genes - whole embryo & sections. These are in the database now. • Other potential sources: – – German Axel. DB frog in situs Japanese NIBB frog in situs (have nice browser) Genepaint. org - mouse stuff EMAGE and Jackson Lab mouse images • From development and other journals, copyright issues. – Nathaniel Heintz BAC expression constructs – Eddy Rubin lab mouse embryos – UCSF cell-localization stuff?

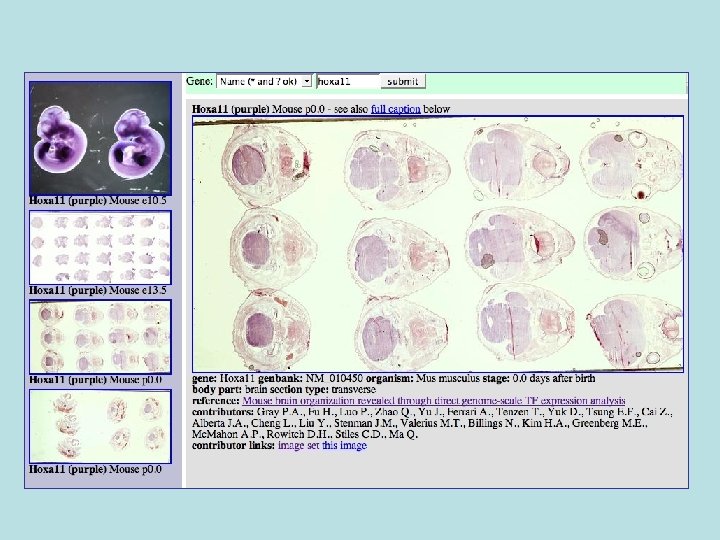
Types of images • Whole animal vs. sectioned tissues, vs. single cell. • Single vs. multiple probes within same image. • Single image vs. image series (movies even). • RNA, Antibody, Fusion protein. Mitotic cell 3 stains

Gene Pix Programs • gene. Pix. Load - loads SQL database from a well defined format involving a. ra file and a tab separated file. See gene. Pix. Load. doc • load. Mahoney - converts Paul Gray (Mahoney center) spreadsheet and image directory into gene. Pix. Load format • Hg/lib/gene. Pix. c - interface with SQL database. • hg. Gene. Pix - cgi script to display images • known. To. Gene. Pix - makes table in mm 5 (or other) genome database to connect known genes to gene. Pix Ids.

Gene Pix Database • Just a single database for all assemblies of all organisms. • A known. To. Gene. Pix table in the assembly database.

Gene. Pix tables • Gene - gene info • gene. Synonym • Antibody - info on an • file. Location - directory antibody • body. Part - whole, brain etc. • slice. Type - transverse, sagital • probe. Type - antibody, RNA, fusion protein • treatment - tech details • Probe - links gene, primers, • contributor - who done it sequence Ab. • Journal - scientific journal • submission. Set - info about a • probe. Color - color probe is whole set of images from one • image. File - file containing author image • section. Set - links together • Image - a single image. separate sections of same • image. Probe links image and specimen. probe

Some Anatomy Required

Especially with slices

Edinburgh mouse atlas

Theiler Stages

Later Stages

NIBB Japanese Frog Site

Earlier Stages

Who you gonna call? Angie Hinrichs - developer of 2 nd and 4 th versions of Table Browser. Genome browser hacker extraordinaire. Hiram Clawson - main mouse man at the moment. Developed ‘wiggle’ tracks.

Kate Rosenbloom - ENCODE project and multiple alignment display. Bob Kuhn - Software and database quality assurance. David Haussler - Ideas. Money. Comparative genomics.

More Acknowledgements • UCSC - Robert Baertsch, Gill Bejerano, Galt Barber, Ron Chao, Mark Diekhans, Jorge Garcia, Patrick Gavin, Rachel Harte, Fan Hsu, Yontoa Lu, Crystal Lynch, Donna Karolchik, Jennifer Jackson, Ann Pace, Jacob Pedersen, Andy Pohl, Katie Pollard, Ali Sultan-Qurraie, Brian Raney, Krishna Roskin, Adam Siepel, Chuck Sugnet, Paul Tatarsky, Daryl Thomas, Heather Trumbower • Penn State - Scott Schwartz, Laura Elnitski, Belinda Giardine, Ross Hardison, Minmei Hou, Webb Miller, Anton Nekrutenko • Funding - NHGRI, HHMI, NCI, UCSC
- Slides: 49