OASCR Genomes to Life OBER Genomes to Life

OASCR Genomes to Life OBER Genomes to Life a partnership between Biology and Computing Gary Johnson John Houghton Office of Science http: //www. doegenomestolife. org/ 1
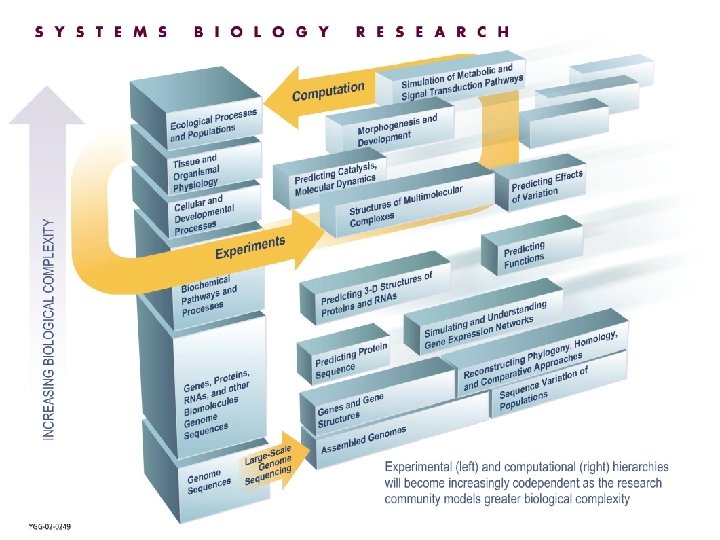
OASCR Genomes to Life OBER 2

OASCR Genomes to Life OBER Office of Advanced Scientific Computing Research: Mathematical, Information and Computational Sciences a brief overview http: //www. sc. doe. gov/production/octr/mics/index. html 3

MICS Mission OASCR Genomes to Life OBER Discover, develop, and deploy the computational and networking advances that enable researchers in the scientific disciplines to analyze, model, simulate, and predict complex physical, chemical, and biological phenomena important to the Department of Energy (DOE). support a broad research portfolio in advanced scientific computing – applied mathematics, computer science, networking and collaboratory software operate supercomputers, a high performance network, and related facilities. 4
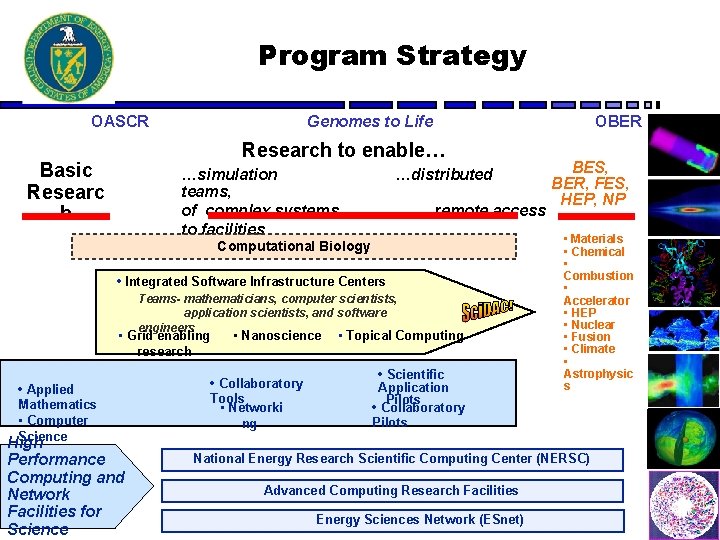
Program Strategy OASCR Genomes to Life OBER Research to enable… Basic Researc h …simulation teams, of complex systems to facilities …distributed remote access • Materials Computational Biology • Integrated Software Infrastructure Centers Teams- mathematicians, computer scientists, application scientists, and software engineers • Grid enabling research • Applied Mathematics • Computer Science High Performance Computing and Network Facilities for Science • Nanoscience • Collaboratory Tools • Networki ng BES, BER, FES, HEP, NP • Topical Computing • Scientific Application Pilots • Collaboratory Pilots • Chemical • Combustion • Accelerator • HEP • Nuclear • Fusion • Climate • Astrophysic s National Energy Research Scientific Computing Center (NERSC) Advanced Computing Research Facilities Energy Sciences Network (ESnet) 5

Budget Request OASCR Genomes to Life OBER FY 2003 - $166, 625, 000 SBIR/STTR Facilities Base Research Enhancements over FY 2002 • Computational Biology • Sci. DAC • Facilities Sci. DAC +$5. 6 M +$5. 3 M +$1. 3 M Comp. Bio. 6
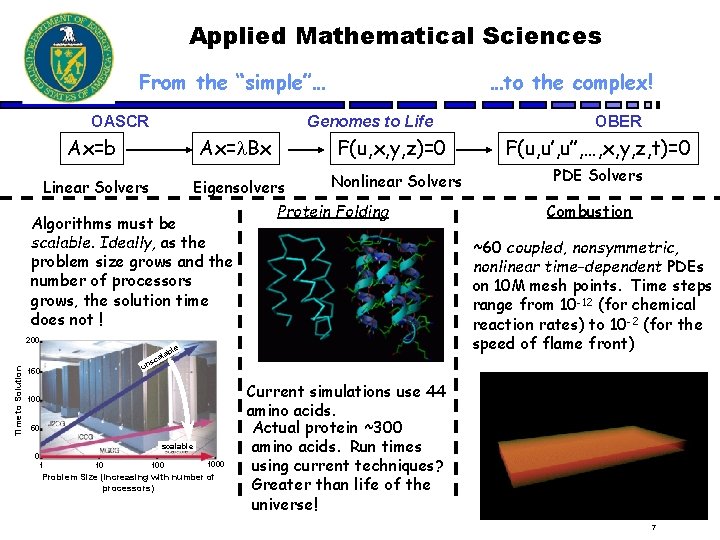
Applied Mathematical Sciences From the “simple”… OASCR …to the complex! Genomes to Life Ax=b Ax= Bx F(u, x, y, z)=0 Nonlinear Solvers Eigensolvers Protein Folding Linear Solvers Algorithms must be scalable. Ideally, as the problem size grows and the number of processors grows, the solution time does not ! Time to Solution 200 a sc OBER F(u, u’’, …, x, y, z, t)=0 PDE Solvers Combustion ~60 coupled, nonsymmetric, nonlinear time-dependent PDEs on 10 M mesh points. Time steps range from 10 -12 (for chemical reaction rates) to 10 -2 (for the speed of flame front) le lab un 150 100 50 scalable 0 1 10 1000 Problem Size (increasing with number of processors) Current simulations use 44 amino acids. Actual protein ~300 amino acids. Run times using current techniques? Greater than life of the universe! 7

AMS Base Research Program OASCR Genomes to Life Objectives Advance our understanding of science and technology by supporting research in basic applied mathematics and in computational research that facilitates the use of the latest high-performance computer systems. Ongoing Projects Applied Mathematics Research: Linear Algebra Grid Generation Fluid Dynamics Predictability Analysis Differential Eqs. & Optimization Uncertainty Quantification Advanced Numerical Algorithms: Automated Reasoning PETSc Hypre Aztec CHOMBO TAO Super. LU ADIFOR / PICO ADIC OBER Accomplishments Robust High-Performance Numerical Libraries Adaptive Mesh Refinement (AMR) Sustained Teraflop/s simulations Level Set / Fast Marching Methods Investment in Education Computational Sciences Graduate Fellowship Growth Opportunities Ultrascalable Algorithms (up to millions of PEs) Mathematical Microscopy These opportunities will be explored through • Genomes to Life (with BER) • Comp. Nanoscience (with BES) • Fusion Energy (FESAC-ASCAC workshop) 8

Computer Science Research OASCR Genomes to Life OBER • Challenge – HPC for Science is (still after fifteen years!) – Hard to use – Inefficient – Fragile – An unimportant vendor market • Vision – A comprehensive, integrated software environment which enables the effective application of high performance systems to critical DOE problems • Goal– Radical Improvement in – Application Performance – Ease of Use – Time to Solution System Admin Res. Mgt Software Scientific Developme Application nt s Framewr PSEs ks Schedule Compiler Viz/Data r s Chkpt/Rs Debugge Math trt rs Libs File Sys Perf Runtme Tools Tls User Space Runtime Support OS OS Kernel Bypass Node and System Hardware Arch HPC System Elements 9
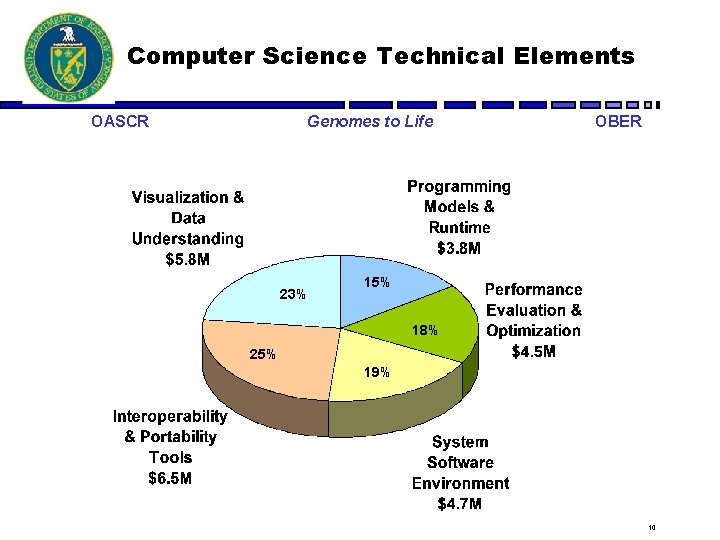
Computer Science Technical Elements OASCR Genomes to Life 23% OBER 15% 18% 25% 19% 10

Major Accomplishments OASCR Genomes to Life OBER • PVM – the first widely successful model for parallel computing • MPI – the lingua franca of today’s parallel computing • MPICH – the open source version of MPI that is the basis for all vendor adaptations • Global Arrays – the distributed shared memory programming model that is at the core of NWChem, the motivating application for Sci. DAC • CTSS – the first interactive operating system for high performance computers • SUNMOS/Puma/Cougar – the most successful high performance parallel operating system • OSCAR – a partnership with industry, the most 11

National Collaboratories OASCR Genomes to Life OBER Why? • The nature of how large scale science is done is changing – Distributed data, computing, people, instruments – Instruments integrated with large-scale computing – Human resources are seldom collocated with the resources needed for their science • Additional drivers – Large and international collaborations – Management of unique national user facilities – Large multi-laboratory science and engineering projects 12

An End-to-End Problem for Applications Many different types of objects need to be connected to and coordinated by the networks OASCR Genomes to Life OBER Scientist 13

Staff OASCR Genomes to Life OBER – Ed Oliver, Associate Director for Advanced Scientific Computing Research – Dan Hitchcock, Senior Scientific Advisor – Linda Twenty, Senior Budget & Financial Specialist – Walt Polansky, Acting Director MICS – – – – – Gary Johnson, ACRTs, Computational Biology Fred Johnson, Computer Science William (Buff) Miner, NERSC & Scientific Applications Thomas Ndousse-Fetter, Network Research Kimberly Rasar, Senior Info. Tech. (Sci. DAC) Chuck Romine, Applied Mathematics Mary Anne Scott, Collaboratories George Seweryniak, Esnet John van Rosendale, Computer Science- Visualization and Data Management – Vacancies- (2) – Jane Hiegel – Susan Kilroy Phone- 301 -903 -5800 Fax- 301 -903 -7774 http: //www. sc. doe. gov/production/octr/mics/index. html 14

OASCR Advisory Committee OASCR Genomes to Life • Committee Chair: • Subcommittee Chairs: – Biology: – Computing Infrastructure: • Members in common with BERAC: • Next Meeting: OBER Margaret Wright, NYU Juan Meza, LBNL Jill Dahlberg, General Atomics Warren Washington, NCAR 2 -3 May 2002 Crowne Plaza Hotel 14 th and K Streets Washington, DC 15

Genomes to Life Program History OASCR Genomes to Life OBER • Phased program startup – FY 2002: OBER – FY 2003: OASCR • Precursor activity – FN 01 -21: Advanced Modeling and Simulation of Biological Systems – 9 Awards, $3 M • Current solicitations – FN 02 -13: Genomes to Life • Program planning – 5 workshops – Goal 4 roadmap – Update to GTL roadmap 16

GTL Planning Activities OASCR • • 7 -8 August 6 -7 September 22 -23 January 6 -7 March 18 -19 March 19 April Future Genomes to Life OBER GTL Computing Workshop Systems Biology & GTL Workshop Computing Infrastructure Workshop Computer Science for GTL Workshop Mathematics for GTL Workshop Draft Goal 4 Roadmap New Edition of the GTL Roadmap 17

GTL Goal 4 Roadmap OASCR Genomes to Life OBER 18

Genomes to Life Goals OASCR Genomes to Life OBER Goal 1 Identify and Characterize the Molecular Machines of Life – the Multiprotein Complexes that Execute Cellular Functions and Govern Cell Form Goal 2 Characterize Gene Regulatory Networks Goal 3 Characterize the Functional Repertoire of Complex Microbial Communities in their Natural Environments at the Molecular Level Goal 4 Develop the Computational Methods and Capabilities to Advance Understanding of Complex Biological Systems and Predict their Behavior 19

Three Computing Domains OASCR Genomes to Life OBER • Bioinformatics/Data-Intensive Applications • Biophysics/Compute-Intensive Applications • Biosystems/Complex Systems Modeling 20
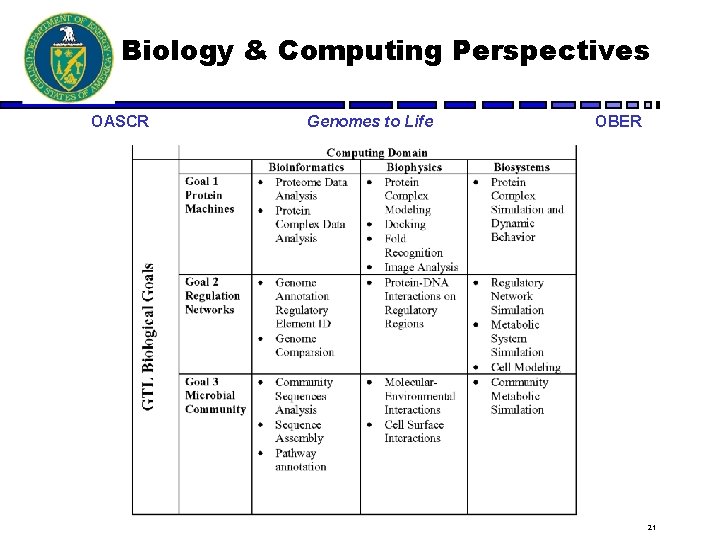
Biology & Computing Perspectives OASCR Genomes to Life OBER 21

Domain Challenges OASCR Genomes to Life OBER • Bioinformatics – Heterogeneous, large and growing data sets – Legacy systems that don’t interoperate and don’t scale • Biophysics – Already bumping up against computational resources • More computation, better algorithms, new theory • Biosystems – Too much data not to have models – Data-poor and biology-poor – Parts list short, but complex systems 22
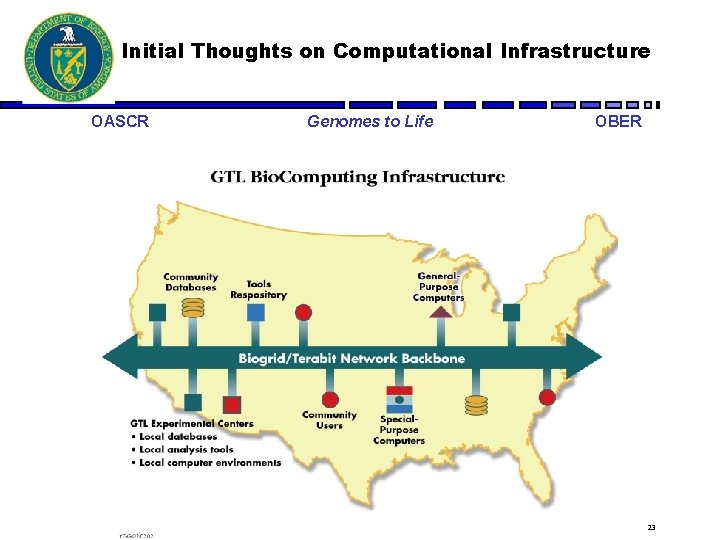
Initial Thoughts on Computational Infrastructure OASCR Genomes to Life OBER 23
- Slides: 23