Genomes at NCBI Genomes at NCBI Database and
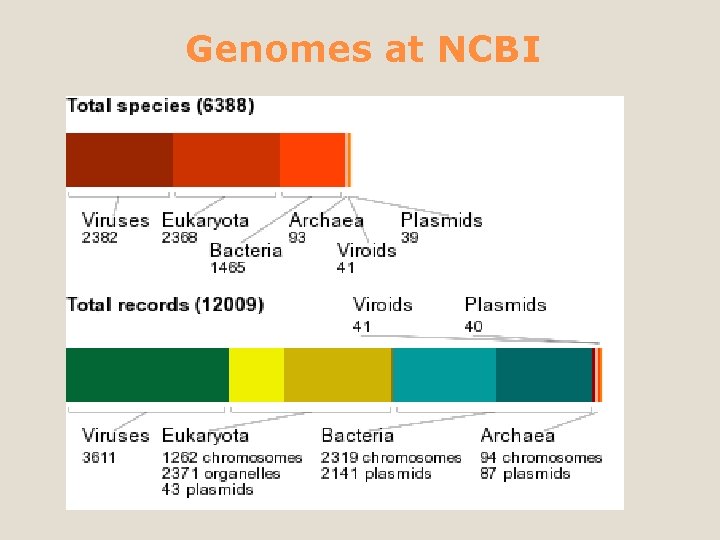
Genomes at NCBI
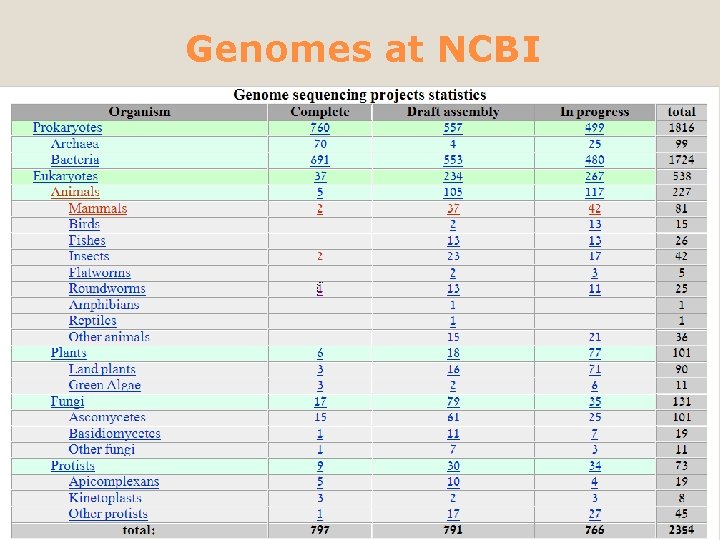
Genomes at NCBI

Database and Tool Explosion The annual database issue of NAR (Nucleic Acids Research) has grown exponentially 1996: first annual compilation of databases and tools lists 57 databases and tools 2009: 1170 databases and tools 2000: 230 databases and tools 3
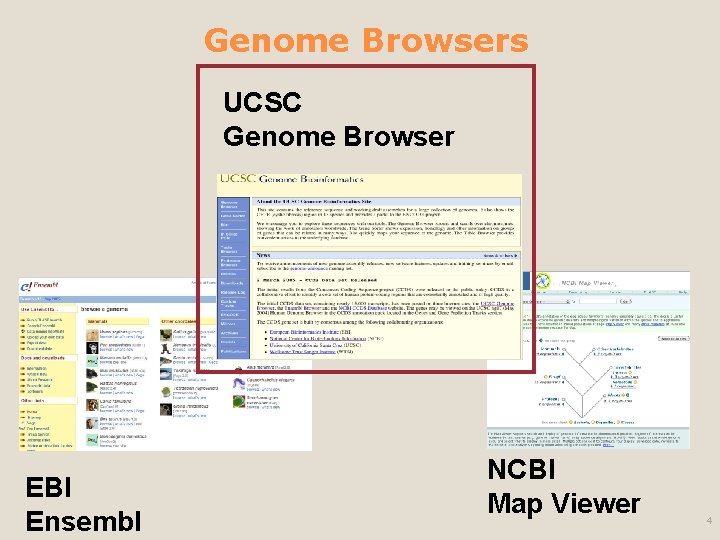
Genome Browsers UCSC Genome Browser EBI Ensembl NCBI Map Viewer 4
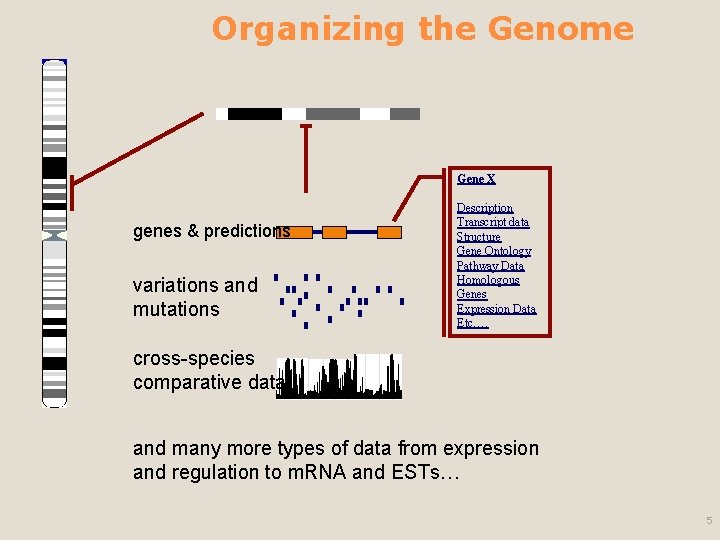
Organizing the Genome Gene X genes & predictions variations and mutations Description Transcript data Structure Gene Ontology Pathway Data Homologous Genes Expression Data Etc…. cross-species comparative data and many more types of data from expression and regulation to m. RNA and ESTs… 5
UCSC Genome Browser: genome. ucsc. edu

UCSC Genome Browser 7
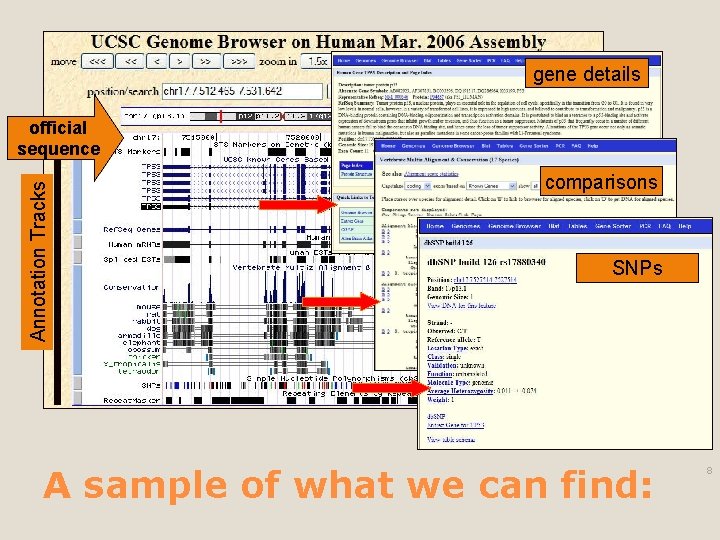
gene details Annotation Tracks official sequence comparisons SNPs A sample of what we can find: 8
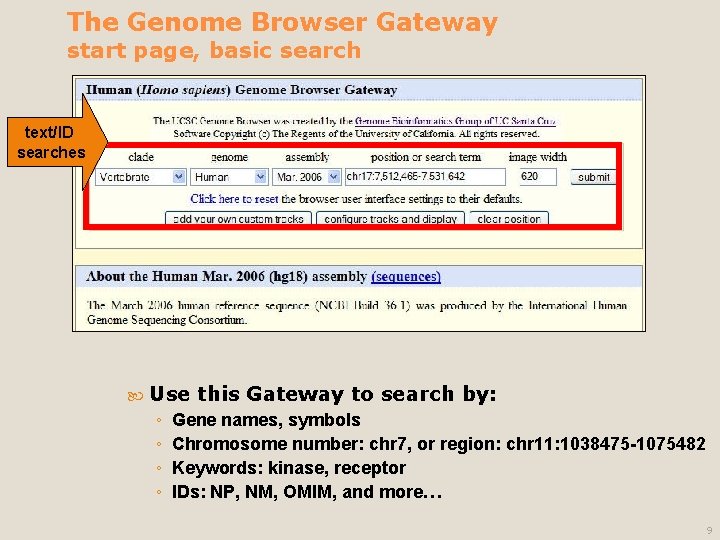
The Genome Browser Gateway start page, basic search text/ID searches Use ◦ ◦ this Gateway to search by: Gene names, symbols Chromosome number: chr 7, or region: chr 11: 1038475 -1075482 Keywords: kinase, receptor IDs: NP, NM, OMIM, and more… 9
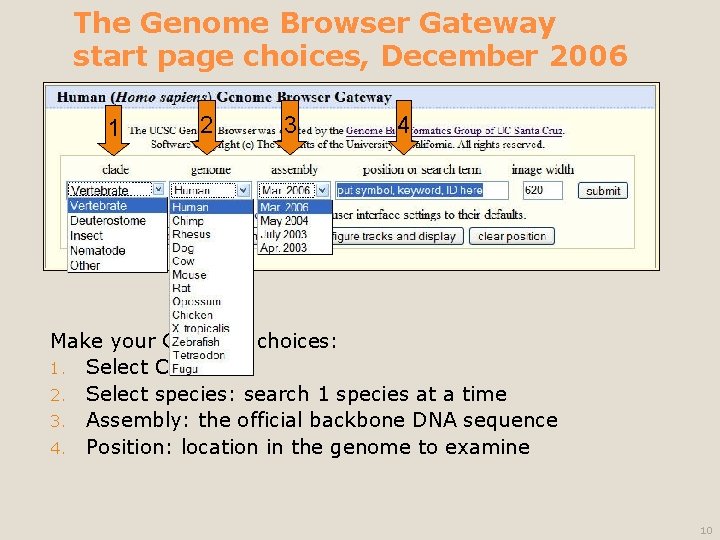
The Genome Browser Gateway start page choices, December 2006 1 2 3 4 Make your Gateway choices: 1. Select Clade 2. Select species: search 1 species at a time 3. Assembly: the official backbone DNA sequence 4. Position: location in the genome to examine 10
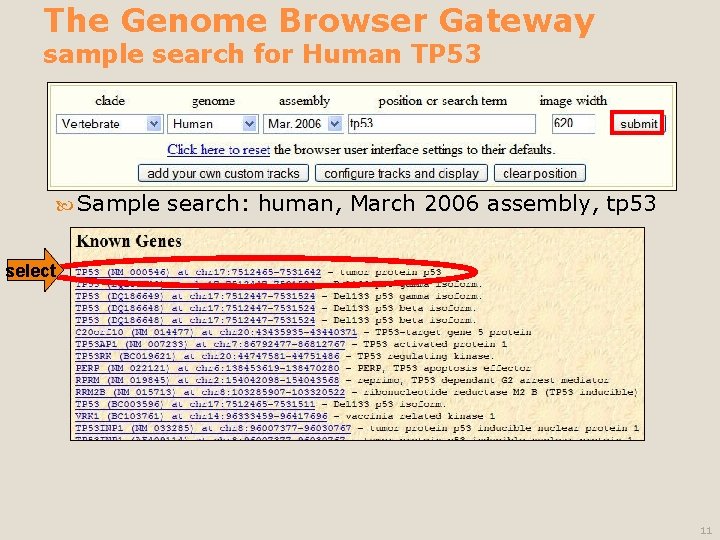
The Genome Browser Gateway sample search for Human TP 53 Sample search: human, March 2006 assembly, tp 53 select 11
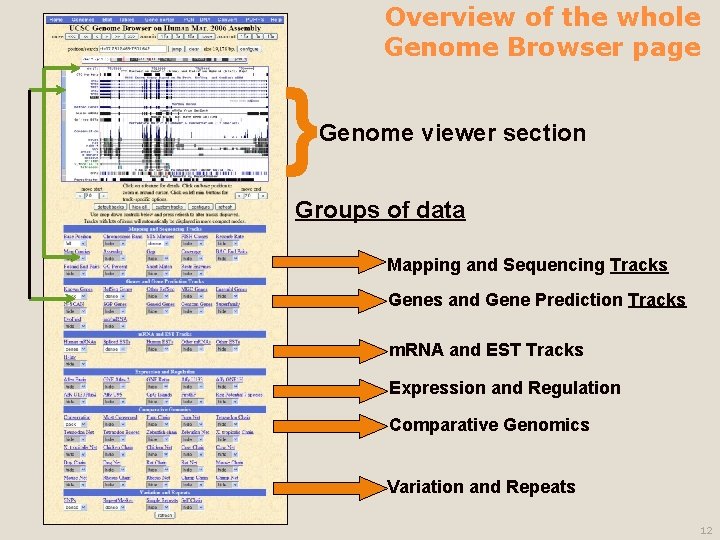
Overview of the whole Genome Browser page } Genome viewer section Groups of data Mapping and Sequencing Tracks Genes and Gene Prediction Tracks m. RNA and EST Tracks Expression and Regulation Comparative Genomics Variation and Repeats 12
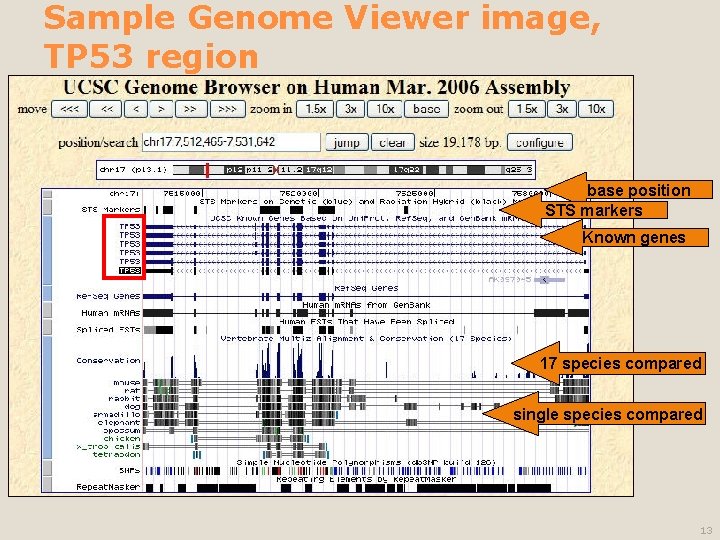
Sample Genome Viewer image, TP 53 region base position STS markers Known genes 17 species compared single species compared 13
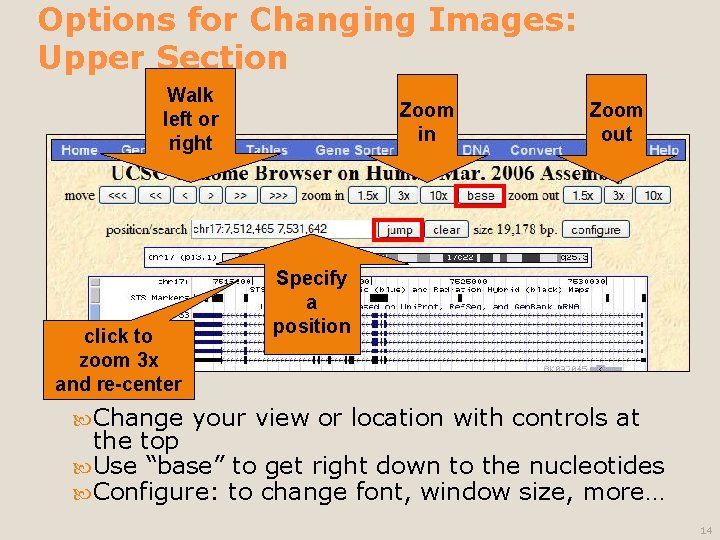
Options for Changing Images: Upper Section Walk left or right click to zoom 3 x and re-center Change Zoom in Zoom out Specify a position your view or location with controls at the top Use “base” to get right down to the nucleotides Configure: to change font, window size, more… 14
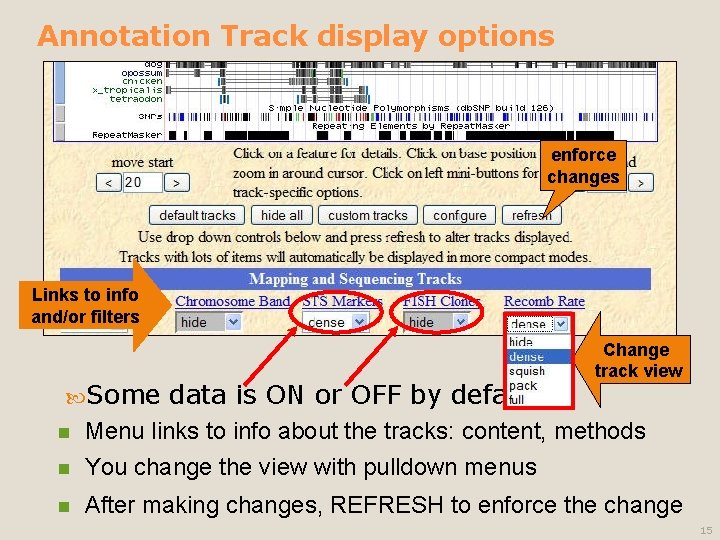
Annotation Track display options enforce changes Links to info and/or filters Some data is ON or OFF by default Change track view n Menu links to info about the tracks: content, methods You change the view with pulldown menus n After making changes, REFRESH to enforce the change n 15

Annotation Track options, defined Hide: removes a track from view n Dense: all items collapsed into a single line n Squish: each item = separate line, but 50% height + packed n Pack: each item separate, but efficiently stacked (full height) n Full: each item on separate line 16

Reset, Hide, Configure or Refresh to change settings enforce any changes (hide, full, squish…) reset, back to defaults start from scratch 17
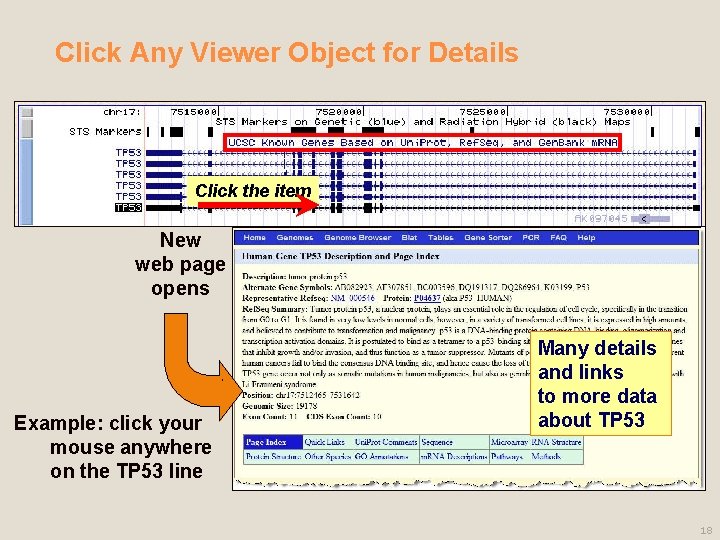
Click Any Viewer Object for Details Click the item New web page opens Example: click your mouse anywhere on the TP 53 line Many details and links to more data about TP 53 18
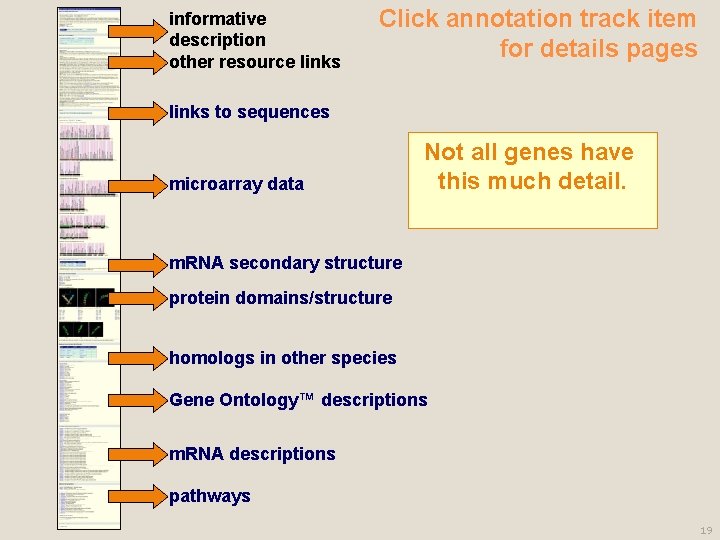
informative description other resource links Click annotation track item for details pages links to sequences microarray data Not all genes have this much detail. m. RNA secondary structure protein domains/structure homologs in other species Gene Ontology™ descriptions m. RNA descriptions pathways 19
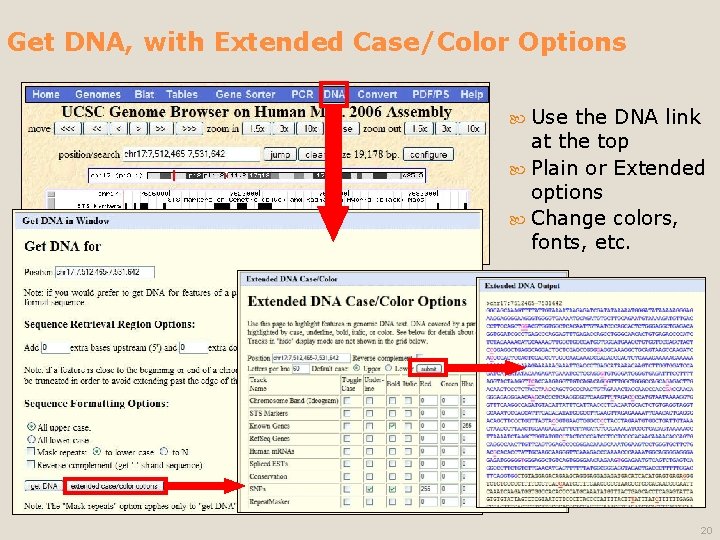
Get DNA, with Extended Case/Color Options Use the DNA link at the top Plain or Extended options Change colors, fonts, etc. 20
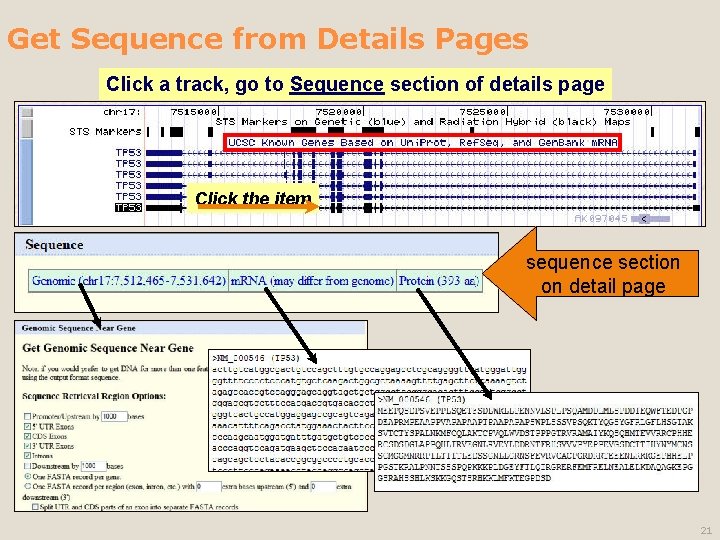
Get Sequence from Details Pages Click a track, go to Sequence section of details page Click the line Click the item sequence section on detail page 21

Accessing the BLAT tool BLAT = BLAST-like Alignment Tool 22
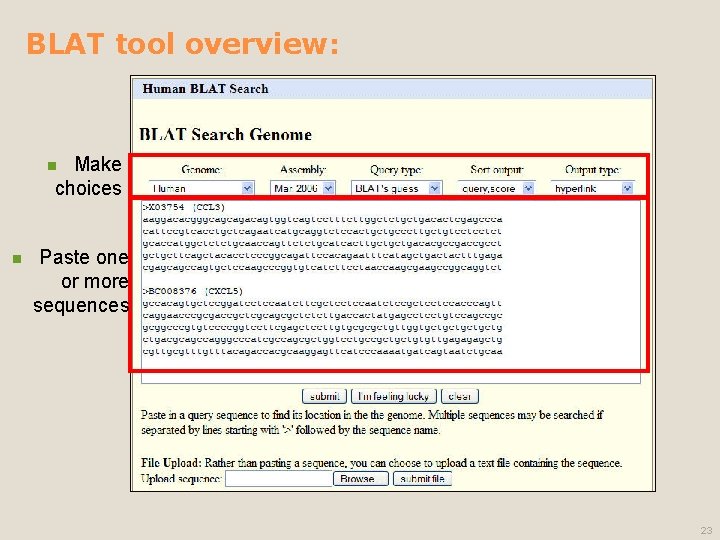
BLAT tool overview: Make choices n n Paste one or more sequences 23
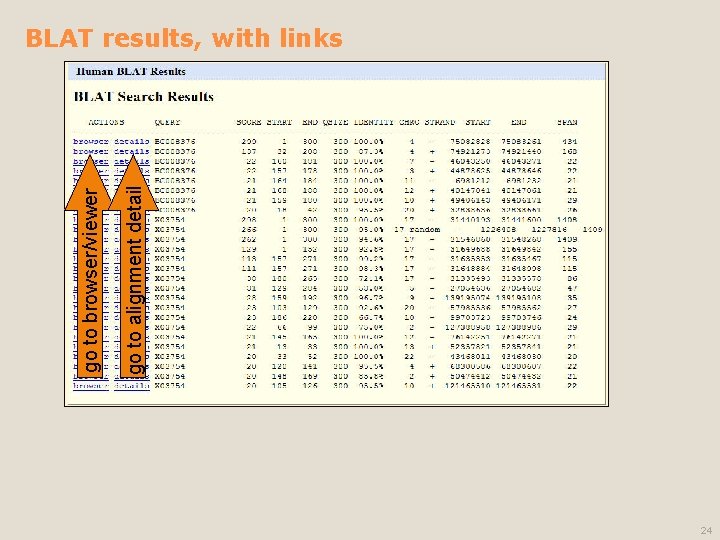
go to alignment detail go to browser/viewer BLAT results, with links 24
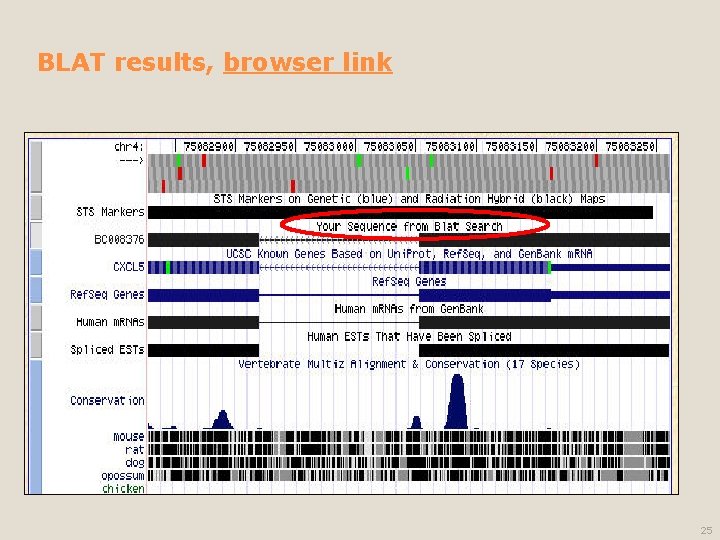
BLAT results, browser link 25
BLAT results, alignment details 26
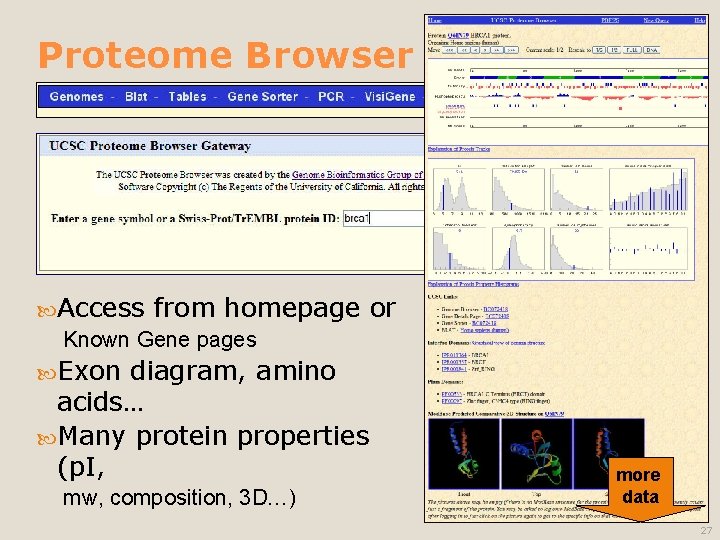
Proteome Browser Access from homepage Known Gene pages diagram, amino acids… Many protein properties (p. I, or Exon mw, composition, 3 D…) more data 27
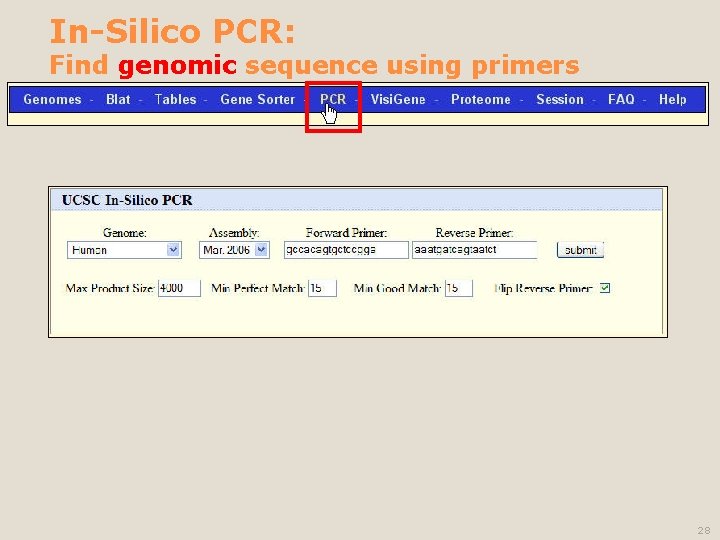
In-Silico PCR: Find genomic sequence using primers 28
- Slides: 28