Computational Biology Lecture 5 Sequencing Bud Mishra Professor

Computational Biology Lecture #5: Sequencing Bud Mishra Professor of Computer Science and Mathematics 10 ¦ 15 ¦ 2001 12/17/2021 ©Bud Mishra, 2001 1
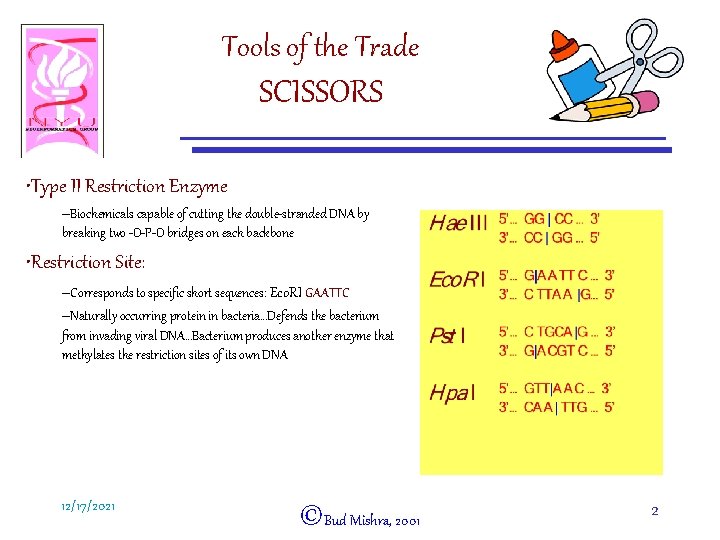
Tools of the Trade SCISSORS • Type II Restriction Enzyme –Biochemicals capable of cutting the double-stranded DNA by breaking two -O-P-O bridges on each backbone • Restriction Site: –Corresponds to specific short sequences: Eco. RI GAATTC –Naturally occurring protein in bacteria…Defends the bacterium from invading viral DNA…Bacterium produces another enzyme that methylates the restriction sites of its own DNA 12/17/2021 ©Bud Mishra, 2001 2

Tools of the Trade GLUE • DNA Ligase –Cellular Enzyme: Joins two strands of DNA molecules by repairing phosphodiester bonds –T 4 DNA Ligase (E. coli infected with bacteriophage T 4) • Hybridization –Hydrogen bonding between two complementary single stranded DNA fragments, or an RNA fragment and a complementary single stranded DNA fragment… results in a double stranded DNA or a DNA-RNA fragment 12/17/2021 ©Bud Mishra, 2001 3
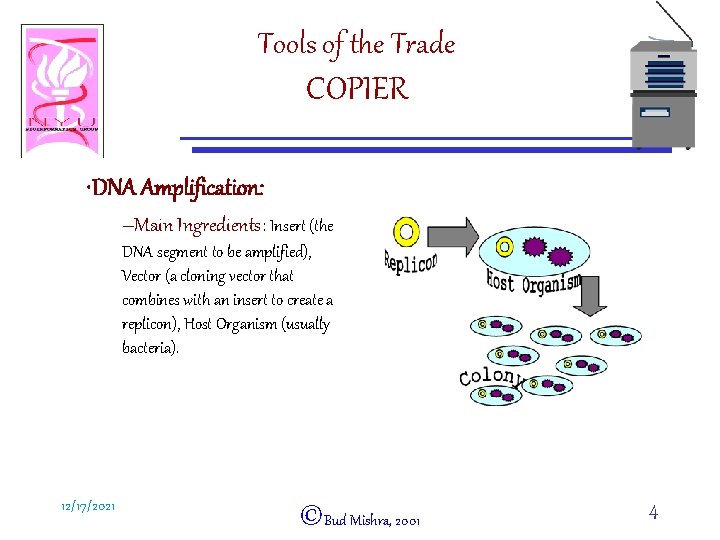
Tools of the Trade COPIER • DNA Amplification: –Main Ingredients: Insert (the DNA segment to be amplified), Vector (a cloning vector that combines with an insert to create a replicon), Host Organism (usually bacteria). 12/17/2021 ©Bud Mishra, 2001 4

Tools of the Trade COPIER PCR (Polymerase Chain Reaction): Main Ingredients: Primers, Catalysts, Templates, and the d. NTPs. 12/17/2021 ©Bud Mishra, 2001 5
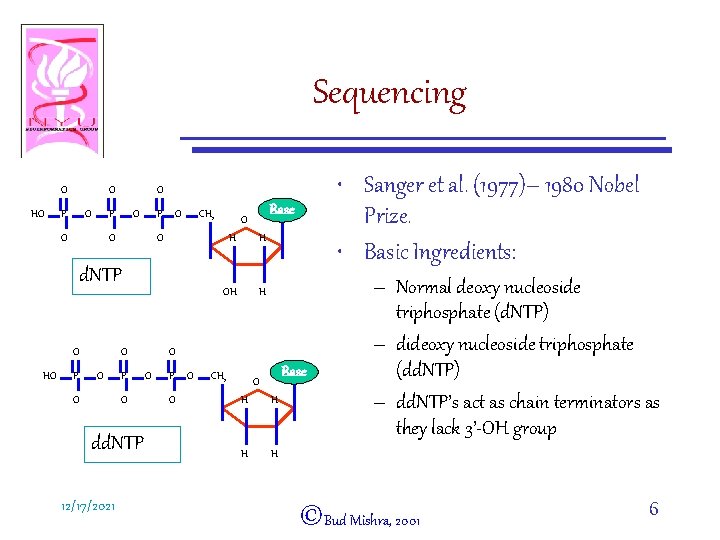
Sequencing O HO O P O- O CH 2 HO P O O O- P O- dd. NTP 12/17/2021 Base O O- d. NTP O • Sanger et al. (1977)– 1980 Nobel Prize. • Basic Ingredients: O H H O O P O- O CH 2 Base O H H – Normal deoxy nucleoside triphosphate (d. NTP) – dideoxy nucleoside triphosphate (dd. NTP) – dd. NTP’s act as chain terminators as they lack 3’-OH group ©Bud Mishra, 2001 6
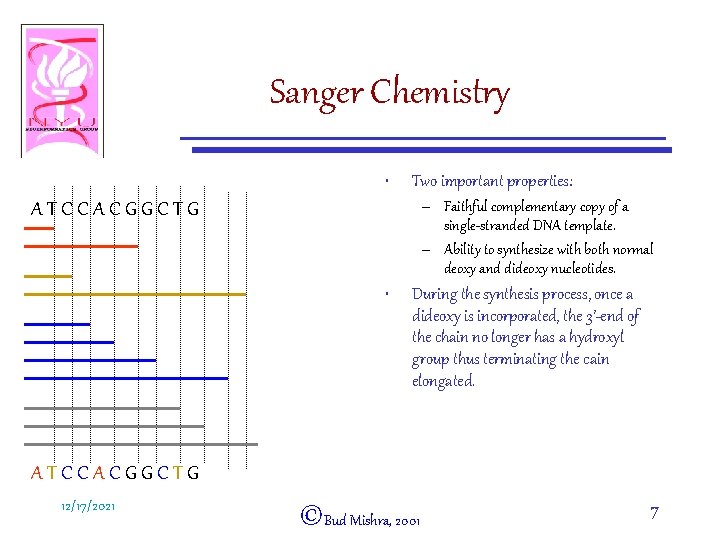
Sanger Chemistry • Two important properties: – Faithful complementary copy of a single-stranded DNA template. – Ability to synthesize with both normal deoxy and dideoxy nucleotides. ATCCACGGCTG • During the synthesis process, once a dideoxy is incorporated, the 3’-end of the chain no longer has a hydroxyl group thus terminating the cain elongated. ATCCACGGCTG 12/17/2021 ©Bud Mishra, 2001 7

Alternative Approaches • Mass Spectroscopy: – MALDI-TOF (Matrix Assisted Laser Desorption Ionization » Time-Of-Flight) • STM Sequencing: – Scanning Tunneling Microscopes (Read base pair information directly from the electron micrographs. ) • ELIDA Sequencing: – Enzymatic Luminometric Inorganic Pyrophosphate Detection Assay • • Exonuclease Sequencing: Sequencing by Hybrdization: Sequencing with Bacterial Ion Channels/Nanopores: Single Molecule Sequencing/Optical Sequencing: 12/17/2021 ©Bud Mishra, 2001 8

Sequencing a Genome • A divide-and-conquer approach: • DIVIDE: Create a “high-coverage” clone library by choosing many randomly located clones. (E. g. , 96, 000 BAC clones, each of length 180 Kb, from a human geome of length 3. 3 Gb. 6 £ coverage BAC library) • CONTIG: Use the clone overlap information to create the contigs. (E. g. , 6£ coverage BAC library would yield 96, 000 £ e-6 ¼ 200 contigs. ) • PRUNE: Remove “non-essential” from the contig to form minimal tiling paths. (Minimal tiling path would consist of about 32, 000 BAC clones. ) 12/17/2021 ©Bud Mishra, 2001 9

Sequencing a Genome • Shotgun Sequencing: 1. Subclone a BAC on the minimal tiling path into M 13’s; 2. Generate sequence reads from M 13 subclones. Each sequence read ¼ 500 bps, with 6 – 12 incorrect base reads. • Contig the Sequence Reads: • Assemble the Sequences and Close the Gaps: 12/17/2021 ©Bud Mishra, 2001 10

Greedy Algorithm • Find overlaps between pairs of sequence reads (only consider overlaps that span at least 15 bps). • Sort overlaps by decreasing length. • Merge read contigs according to the sorted list. 12/17/2021 ©Bud Mishra, 2001 11

Practical Problems for Sequence Assembly • • It is possible to read a contiguous subsequence of DNA up to 500 bps. These short “sequence reads” are to be combined to create sequenced DNA up to – 50 Kb (Cosmids) – 200 Kb (BACs) – 3, 000 Mb (Whole Genome) • As the size of the “Sequenced DNA” scales up to the whole genome, several other issues are faced: – – 12/17/2021 Segmental duplications, homologies, etc. Repeats, compression and gaps. False positive overlap probability. Computational complexity. ©Bud Mishra, 2001 12
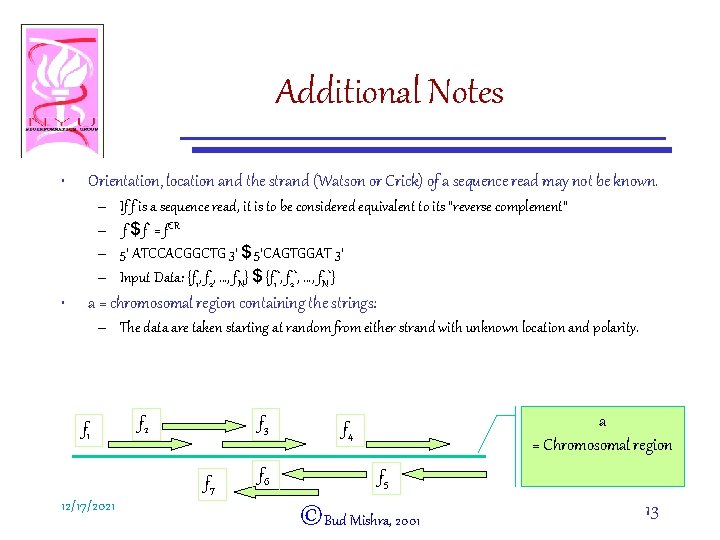
Additional Notes • Orientation, location and the strand (Watson or Crick) of a sequence read may not be known. – – • If f is a sequence read, it is to be considered equivalent to its “reverse complement” f $ f* = f. CR 5’ ATCCACGGCTG 3’ $ 5’CAGTGGAT 3’ Input Data: {f 1, f 2, …, f. N} $ {f 1*, f 2*, …, f. N*} a = chromosomal region containing the strings: – The data are taken starting at random from either strand with unknown location and polarity. f 1 f 2 f 3 f 7 12/17/2021 f 6 a = Chromosomal region f 4 f 5 ©Bud Mishra, 2001 13

Sequence Reconstruction Problem • Given: F = {f 1, f 2, …, f. N} Infer: a (chromosomal region) from F. – Caveat: The data may have errors: Deletions, Insertions and Substitutions. • Simplest Model: – All fi’s have same polarity. – fi’s are free of errors. • Sequence Reconstruction Problem (SRP): • Shortest Superstring Problem (SSP): – SSP is NP-complete. a Transformation from Vertex Cover for Cubic Graphs. – SRP is NP-complete. a Transformation from SSP. 12/17/2021 ©Bud Mishra, 2001 14

SSP • Given: A set of N strings over an alphabet {A, T, C, G}: F = {f 1, f 2, …, f. N} • Find: A string S of minimal length such that each fi ( i = 1, …, N) is a substring of S. S = Shortest Common Superstring 12/17/2021 ©Bud Mishra, 2001 15

SRP • Given: F = {f 1, f 2, …, f. N} e 2 [0, 1) = An error rate d(¢, ¢) = Distance function between two sequences [e. g. , edit distance] Notation: x Æ y = min(x, y) • Find: A string S such that 8 i 9 a v S d(a, fi) Æ d(a, fi*) 5 e |a|. 12/17/2021 ©Bud Mishra, 2001 16

Relation between SSP & SRP • Consider the following homomorphism: si 2 {A, T, C, G}* h: si = x 1 x 2 L xm a – – • • – fi =AA x 1 CC AA X 2 CC AA xm CC fi* = GG xm. C TT GG xm-1 C TT L GG x 1 C TT 8 i ¹ j fi* cannot overlap with fj Assume that e = 0. Hence, S is a sloution to SSP for {s 1, s 2, …, s. N} iff S = h(S) & S* = h(S)* are solutions to SRP for {f 1 = h(s 1), f 2 = h(s 2), …, f. N = h(s. N)} • • 12/17/2021 ©Bud Mishra, 2001 17

A Greedy Algorithm • F = {f 1, f 2, …, f. N} while 9 fi and fj in F s. t. i ¹ j and fi Å fj ¹ ; loop Choose the substrings fi and fj with MOST overlap F = (F n {fi, fj}) [ {S(fi, fj)} /* S(fi, fj) = Shortest Common Superstring of fi and fj; this can be done in time O(|fi| |fj|). */ end loop return F = {S} • Time Complexity = O(N 3 max(|fi|)2) = polynomial. 12/17/2021 ©Bud Mishra, 2001 18
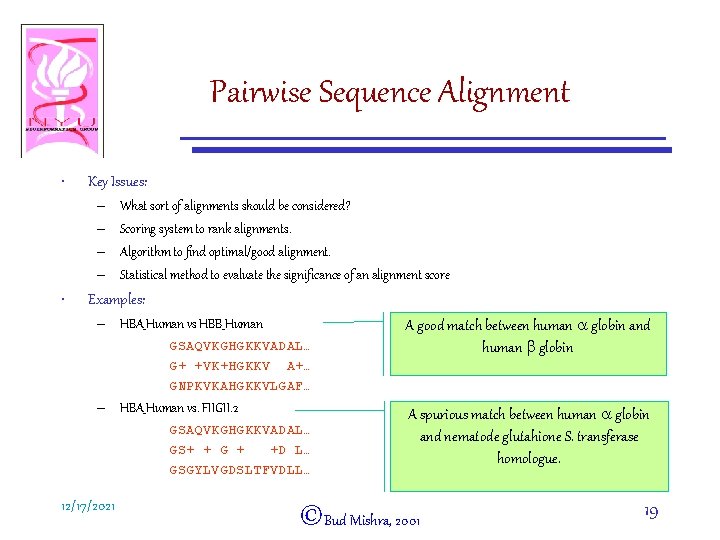
Pairwise Sequence Alignment • Key Issues: – – • What sort of alignments should be considered? Scoring system to rank alignments. Algorithm to find optimal/good alignment. Statistical method to evaluate the significance of an alignment score Examples: – HBA_Human vs HBB_Human GSAQVKGHGKKVADAL… G+ +VK+HGKKV A+… GNPKVKAHGKKVLGAF… – HBA_Human vs. FIIGII. 2 GSAQVKGHGKKVADAL… GS+ + G + +D L… GSGYLVGDSLTFVDLL… 12/17/2021 A good match between human a globin and human b globin A spurious match between human a globin and nematode glutahione S. transferase homologue. ©Bud Mishra, 2001 19

Dynamic Programming • Simple Score Function: – d(xi, yj) = {1, if xi ¹ yj {0, – d(xi, -) = d(-, yj) = 1 if xi = yj • Defn: For two strings x and y, D(i, j) is defined to be the edit distance of the prefixes x[1. . i] and y[1. j] = Minimum number of atomic operations (substitutions and indels) needed to transform the first i letters of x into irst j letters of y. |x| = n, |y| = m ) S(x, y) = D(n, m) 12/17/2021 ©Bud Mishra, 2001 20
![Recurrence Relation • D(i, j) = Edit distance of x[1. . i] and y[1. Recurrence Relation • D(i, j) = Edit distance of x[1. . i] and y[1.](http://slidetodoc.com/presentation_image_h2/13cc6799848171417dabcee5d27b147e/image-21.jpg)
Recurrence Relation • D(i, j) = Edit distance of x[1. . i] and y[1. . j] – – 12/17/2021 D(0, 0) = 0, D(i, 0) = i, (i deletions from x) D(0, j) = j, (j insertions into y) i = 1, j = 1: D(i, j) = min{D(i-1, j) + 1, (1 deletion from x) {D(i, j-1) +1, (1 insertion into y) {D(i-1, j-1) + d(xi, yj). (1 substitution xi a yj) ©Bud Mishra, 2001 21
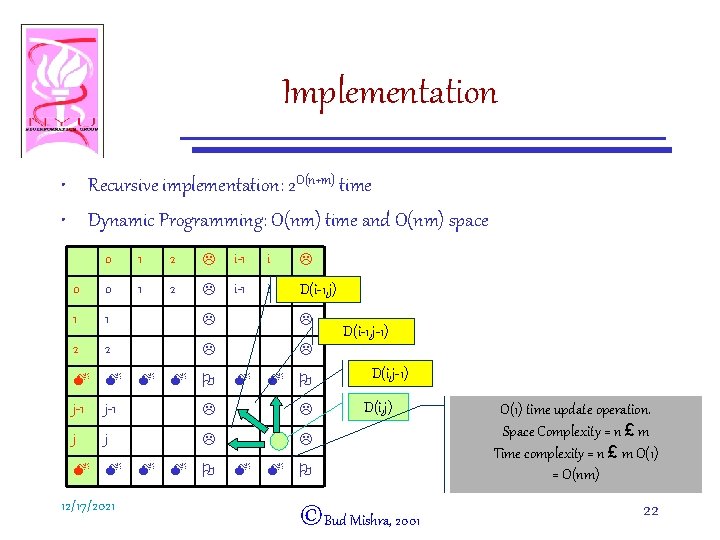
Implementation • Recursive implementation: 2 O(n+m) time • Dynamic Programming: O(nm) time and O(nm) space 0 1 2 L i-1 i L 0 0 1 2 L i-1 i D(i-1, j) L 1 1 L L 2 2 L L M M O j-1 L L j j L L M M O 12/17/2021 D(i-1, j-1) D(i, j) ©Bud Mishra, 2001 O(1) time update operation. Space Complexity = n £ m Time complexity = n £ m O(1) = O(nm) 22
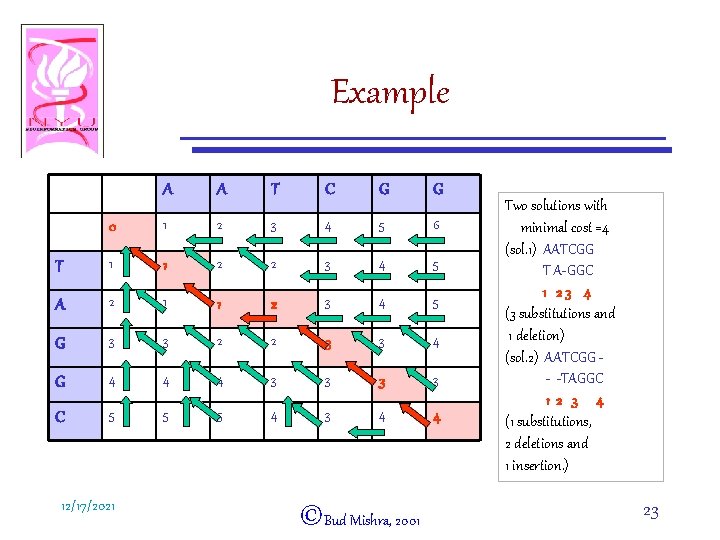
Example A A T C G G 0 1 2 3 4 5 6 T 1 1 2 2 3 4 5 A 2 1 1 2 3 4 5 G 3 3 2 2 3 3 4 G 4 4 4 3 3 C 5 5 5 4 3 4 4 12/17/2021 ©Bud Mishra, 2001 Two solutions with minimal cost =4 (sol. 1) AATCGG T A-GGC 1 23 4 (3 substitutions and 1 deletion) (sol. 2) AATCGG - -TAGGC 12 3 4 (1 substitutions, 2 deletions and 1 insertion. ) 23

Performance of the Greedy Algorithm • Greedy Algorithm: Repeatedly combine the pairs of “most overlapping” sequence contigs. • F = {f 1, f 2, …, f. N} – Assume that no substring contains another. – fi = uv, fj = vw ) fi Å fj = v (possibly, emptystring l) Note: u ¹ l. – Let v be the longest possible such string • Overlap Length, ov(i, j) = |v| • Prefix Length, pf(i, j) = |u|. 12/17/2021 ©Bud Mishra, 2001 24

Example • fi = ATAT • fj = TATA ov(i, j) =3, pf(i, j) =1, ov(j, i) = 3 pf(j, i) = 1 ov(i, j) = 3, pf(i, j) = 1, ov(j, i) = 1 pf(j, i) = 3 – --------- • fi = ATTA • fj = TTAA – --------- • Triangle Inequality: Note that 8 i, j, k pf(i, j) + pf(j, k) = pf(i, k) 12/17/2021 ©Bud Mishra, 2001 25

Prefix Graph • Directed Labeled Graph: (V, E, pf) – V ' F, E ' F £ F – pf(i, j) = weight on each edge connecting fi & fj • A Hamiltonian cycle in G is a simple cycle that visits every vertex of G: – HC = (vp(1), vp(2), …, vp(N)) – Weight(HC) = pf(p(1), p(2)) + L + pf(p(N-1), p(N)) + pf(p(N), p(1)) • MWHC(G) = Minimum Weight Hamiltonian Cycle of G. 12/17/2021 ©Bud Mishra, 2001 26

Cycle Cover of G • • (1) (2) A set of simple cycles of G such that each vertex belongs exactly one cycle. CYC(G) = Minimum Total Weight Cycle Cover of G. CYC(G) 5(1) MWHC(G) 5(2) OPT(S) ------Note: Hamiltonian Cycle is a cycle cover Note: Shortest Superstring has a length larger than that of the Min. Wt. Hamiltonian Cycle. 12/17/2021 ©Bud Mishra, 2001 27
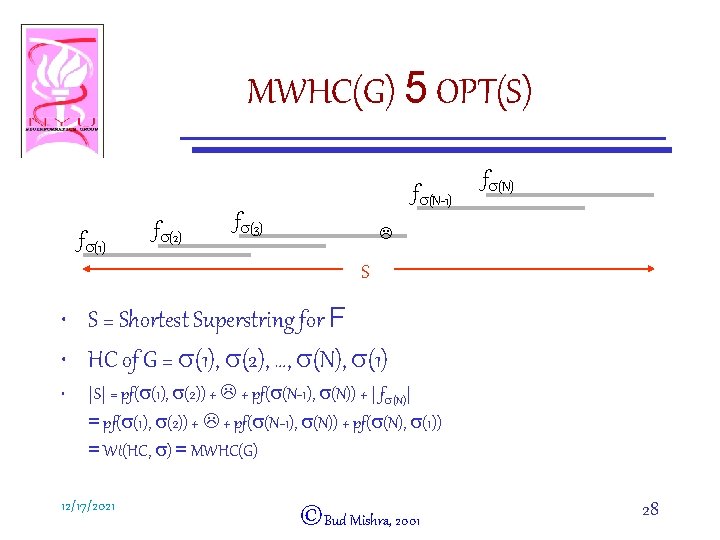
MWHC(G) 5 OPT(S) fs(N-1) fs(2) fs(3) fs(N) L S • S = Shortest Superstring for F • HC of G = s(1), s(2), …, s(N), s(1) • |S| = pf(s(1), s(2)) + L + pf(s(N-1), s(N)) + | fs(N)| = pf(s(1), s(2)) + L + pf(s(N-1), s(N)) + pf(s(N), s(1)) = Wt(HC, s) = MWHC(G) 12/17/2021 ©Bud Mishra, 2001 28

Greedy Algorithm • Output = T. • Claim: |T| · 3 CYC(G) + OPT(S) · 4 OPT(S) fk-1 f 2 f 3 • Wt(ci) = wi & • Longest String in Ci = li. L Contig 12/17/2021 fk • Consider a contig of the Greedy Algorithm. • Cycle =f 1, f 2, …, fk, f 1 = C 1 • |T| 5 å(wi+li) · å li + OPT(S) ©Bud Mishra, 2001 29

Technical Lemma • Claim: fi 2 Ci & fj 2 Cj ) ov(i, j) 5 wi + wj • Proof: – Suppose ov(i, j) > wi+wj ) Ci and Cj can be combined into a ew cycle of smaller weight ) Greedy Algorithm failed to combine two contigs into a new contig of smaller weight ) Contradiction # 12/17/2021 ©Bud Mishra, 2001 30

The Competitive Bound • |T| 5 å (wi+ li) = å (li – 2 wi) + å 3 wi 5 å li - å (wi+wj) + 3 å wi 5 å li - å ov(i, j) + 3 å wi 5 OPT(S) + 3 OPT(S) = 4 OPT(S) • The greedy algorithm computes a solution that is never worse than 4 times the best possible solution. � 12/17/2021 ©Bud Mishra, 2001 31
- Slides: 31