Computational Biology at Carnegie Mellon University A Quick

Computational Biology at Carnegie Mellon University A Quick Tour Jaime Carbonell Carnegie Mellon University December, 2008

Computational Biology at CMU: Educational History n n 1987 Undergraduate program in Computational Biology established 1991 Howard Hughes Medical Institute grant to build undergrad curriculum 2000 M. S. Program in Computational Biology established 2005 Joint CMU & U. of Pittsburgh PHD Program in Computational Biology

Computational Biology at CMU: History n n n 2002 NSF large ITR grant (CMU PI: Reddy & Carbonell) with U, Pitt, MIT, Boston U, NRC Canada Computational Biolinguistics 2003 NSF large ITR grant (CMU PI: Murphy) with UCSB, Berkeley, MIT Bioimage Informatics 2004 -2008 10 small grants from NSF, NIH, Merck, Gates on: Computational proteomics, viral evolution, HIV-human interactome, …

Joint CMU-Pitt Ph. D. Program in Computational Biology

Curriculum for Comp Bio Ph. D n Core graduate courses n n n n Molecular Biology Biochemistry Biophysics Advanced Algorithms & Language Tech. Machine Learning Methods Computational Genomics Computational Structural Biology Cellular and Systems Modeling

Curriculum n Elective Courses n n n Computational Genomics Computational Structural Biology Cellular and Systems Modeling Bioimage Informatics Computational Neurobiology Advanced Statistical Learning Methods

Example Books Used

Teaching & Advising Faculty n 30 faculty from CMU n n n 11 Computer Science 11. 5 Biology and Chemistry 3. 5 Bio-Engineering 3 Statistics and Mathematics 1 Business School 36 faculty from Pitt n n 19 Medical School 17 Biology, Chemistry, Physics

Faculty: Computational Genomics n n n n n Ziv Bar-Joseph* Jaime Carbonell Marie Dannie Durand* Jonathan Minden Ramamoorthi Ravi Kathryn Roeder Roni Rosenfeld Larry Wasserman Eric Xing* * = Primary research area Linguistics methods for elucidating sequence-structurefunction relations Machine Learning methods for annotation Modeling genome evolution through duplication

Faculty: Computational Structural Biology (Proteomics) n n n n Homologous structure determination by NMR Michael Erdmann Maria Kurnikova* Improving Chris Langmead* determination of protein structure and John Nagle dynamics using sparse data Gordon Rule Molecular dynamics of Robert Swendsen proteins and nucleic acids Jaime Carbonell*

Faculty: Cellular and Systems Modeling n n n n Computational modeling of mechanical properties of cells and tissues Ziv Bar-Joseph* Omar Ghattas Philip Le. Duc Modeling of Russell Schwartz* formation of protein complexes Joel Stiles* Multi-scale modeling Shlomo Ta’asan of excitable membranes Yiming Yang of large-scale gene Eric Xing Discoveryregulatory networks

Faculty: Bioimage Informatics n n n n Determining subcellular location from microscope images William Cohen Bill Eddy Christos Faloutsos Generative models of protein traffic Jelena Kovacevic Machine learning of Tom Mitchell* patterns of brain activity Robert Murphy* Statistical analysis of gel images for proteomics Eric Xing

Faculty: Computational Neurobiology n n n Justin Crowley Tom Mitchell Joel Stiles* David Touretzky* Nathan Urban Development of structure of neuronal circuits Machine learning of patterns of brain activity Multi-scale modeling of excitable membranes
Proteomics n Things to learn about proteins n n n n sequence activity Partners Structure Functions Expression level Location/motility

Examples of Cool Research n Computational Biolinguistics n n Sequence (DNA, Protein) Structure Function Language (Speech, Text) Syntax Semantics GPCRs (sensor/channel proteins, Klein CMU/Pitt) n n n Evolutionary Analysis (of genes, proteins, …) n n Conservation, replication, poly-functionality (Rosenberg) Immune System Modeling (just starting…) n n 60% of all targeted drugs affect GPCRs Language (information-theoretic) analysis Domain/Fold polymorphic modeling (Langmead) Cross-species Interactome (just starting…) n Human-HIV protein-protein (Carbonell, Klein)
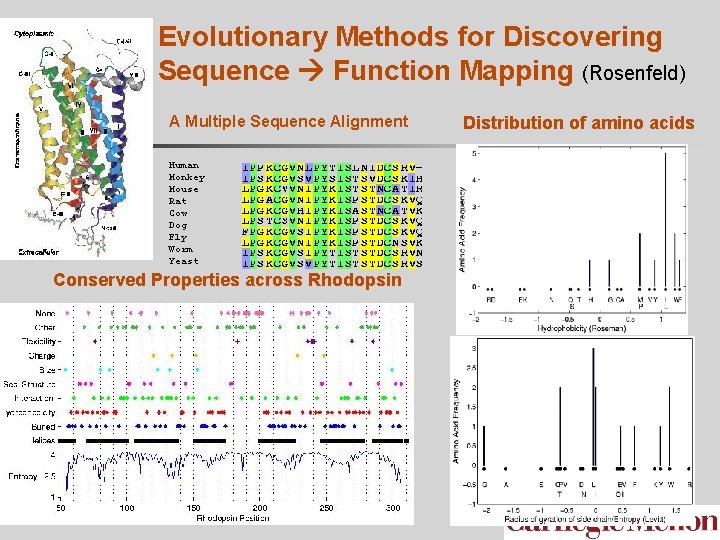
Evolutionary Methods for Discovering Sequence Function Mapping (Rosenfeld) A Multiple Sequence Alignment Human Monkey Mouse Rat Cow Dog Fly Worm Yeast Conserved Properties across Rhodopsin Distribution of amino acids
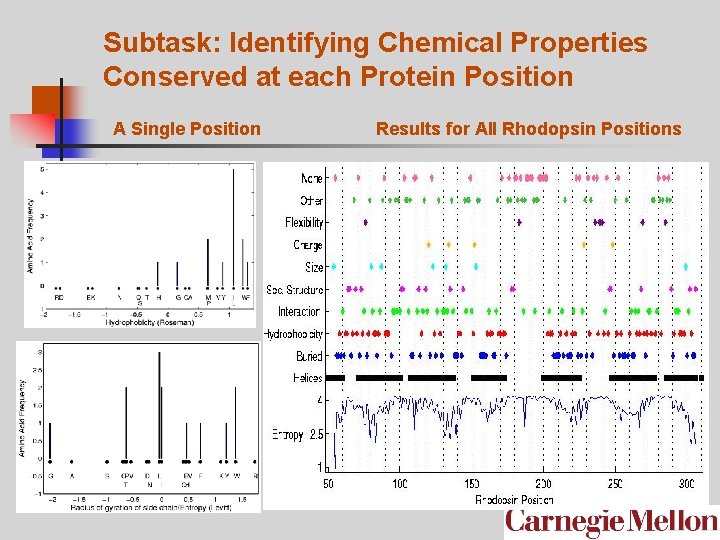
Subtask: Identifying Chemical Properties Conserved at each Protein Position A Single Position Results for All Rhodopsin Positions
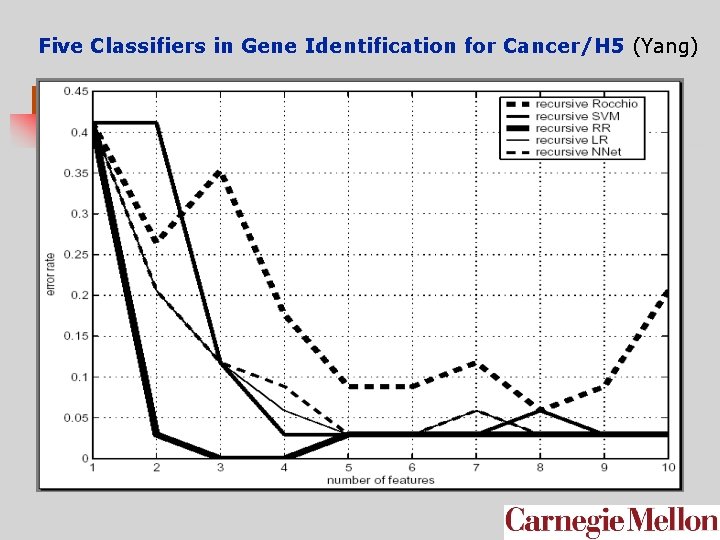
Five Classifiers in Gene Identification for Cancer/H 5 (Yang)

New Field: Location Proteomics (Langmead) n n n Can use CD-tagging (developed by Jonathan Jarvik and Peter Berget) to randomly tag many proteins Isolate separate clones, each of which produces one tagged protein Use RT-PCR to identify tagged gene in each clone Collect many live cell images for each clone using spinning disk confocal fluorescence microscopy Cluster proteins by their location patterns (automatically)
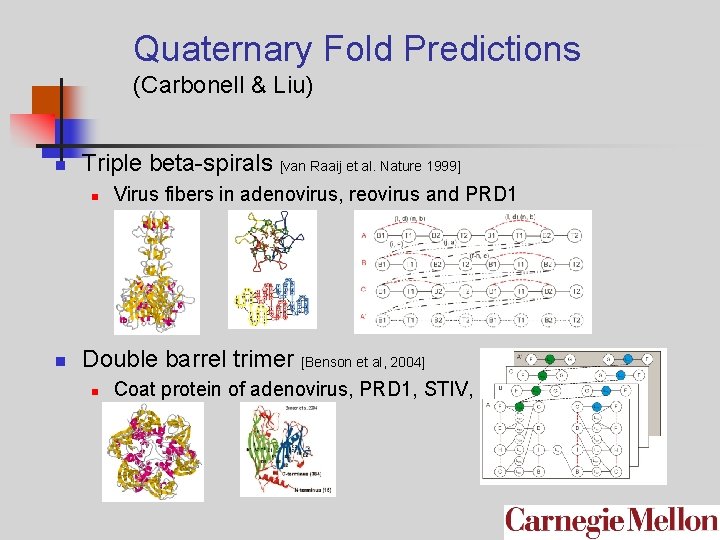
Quaternary Fold Predictions (Carbonell & Liu) n Triple beta-spirals [van Raaij et al. Nature 1999] n n Virus fibers in adenovirus, reovirus and PRD 1 Double barrel trimer [Benson et al, 2004] n Coat protein of adenovirus, PRD 1, STIV, PBCV
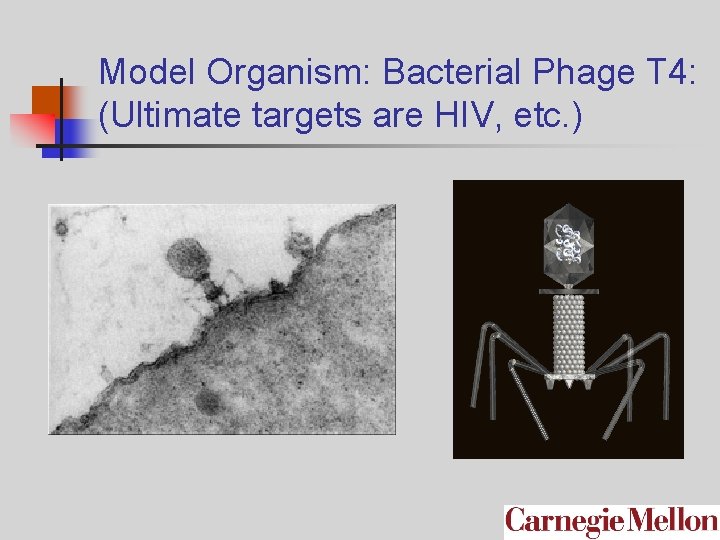
Model Organism: Bacterial Phage T 4: (Ultimate targets are HIV, etc. )
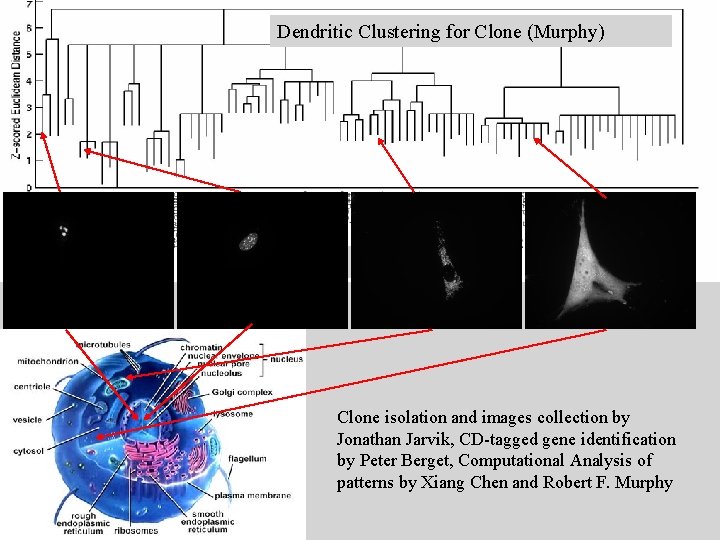
Dendritic Clustering for Clone (Murphy) Protein name Clone isolation and images collection by Jonathan Jarvik, CD-tagged gene identification by Peter Berget, Computational Analysis of patterns by Xiang Chen and Robert F. Murphy
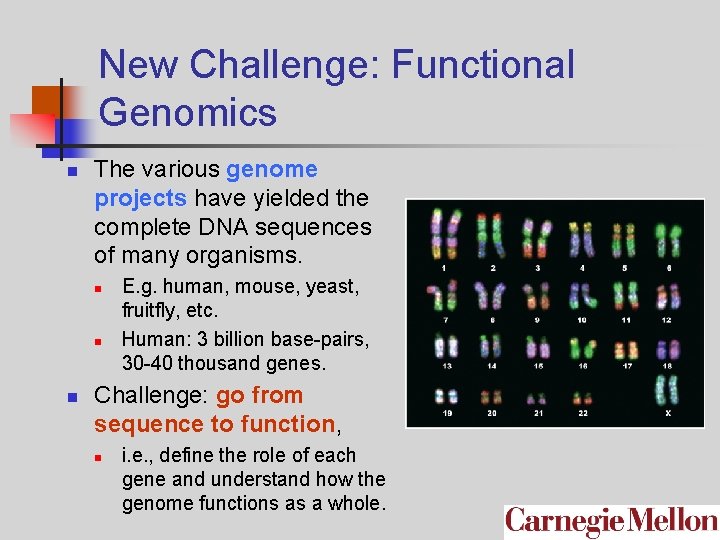
New Challenge: Functional Genomics n The various genome projects have yielded the complete DNA sequences of many organisms. n n n E. g. human, mouse, yeast, fruitfly, etc. Human: 3 billion base-pairs, 30 -40 thousand genes. Challenge: go from sequence to function, n i. e. , define the role of each gene and understand how the genome functions as a whole.
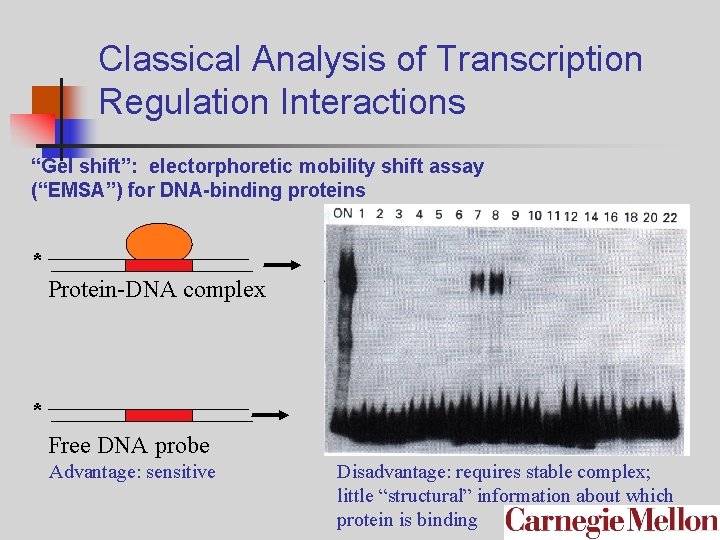
Classical Analysis of Transcription Regulation Interactions “Gel shift”: electorphoretic mobility shift assay (“EMSA”) for DNA-binding proteins * Protein-DNA complex * Free DNA probe Advantage: sensitive Disadvantage: requires stable complex; little “structural” information about which protein is binding
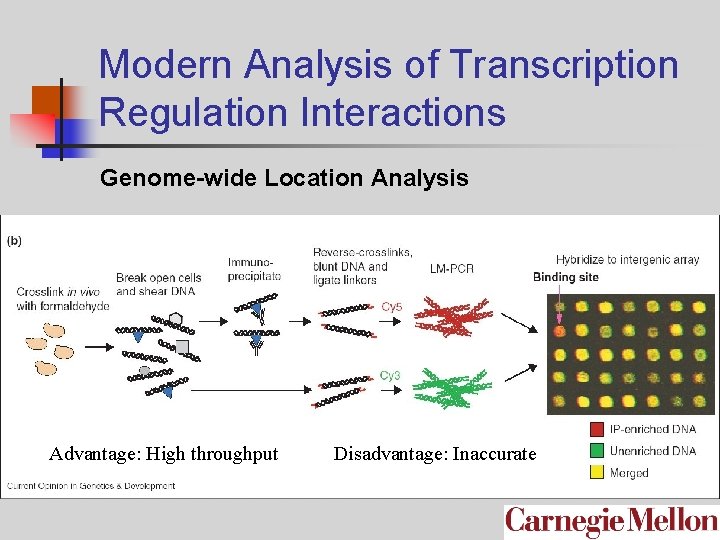
Modern Analysis of Transcription Regulation Interactions Genome-wide Location Analysis Advantage: High throughput Disadvantage: Inaccurate
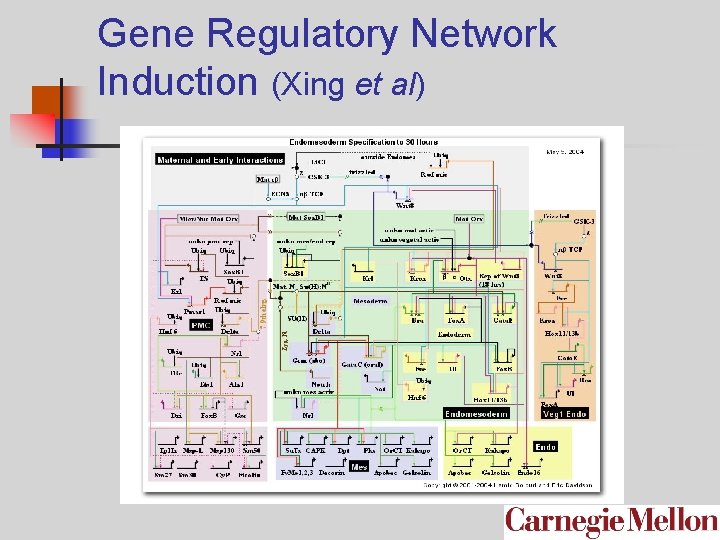
Gene Regulatory Network Induction (Xing et al)
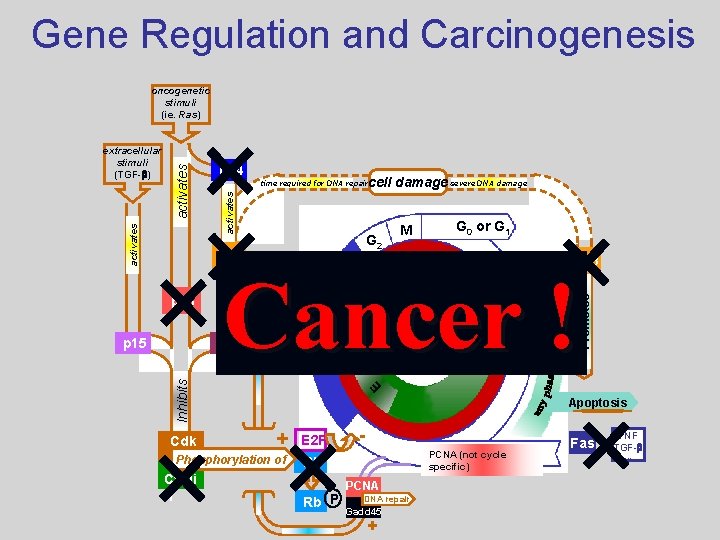
Gene Regulation and Carcinogenesis oncogenetic stimuli (ie. Ras) cell damage severe DNA damage activates time required for DNA repair M G 2 p 53 A G 0 or G 1 p 53 B Promotes p 16 S p 21 G 1 Cdk E Inhibits p 15 p 14 transcriptional activation activates Cancer ! extracellular stimuli (TGF-b) Apoptosis + Phosphorylation of Cycli n E 2 F - PCNA (not cycle specific) Rb Rb P PCNA DNA repair Gadd 45 + Fas TNF TGF-b. . .
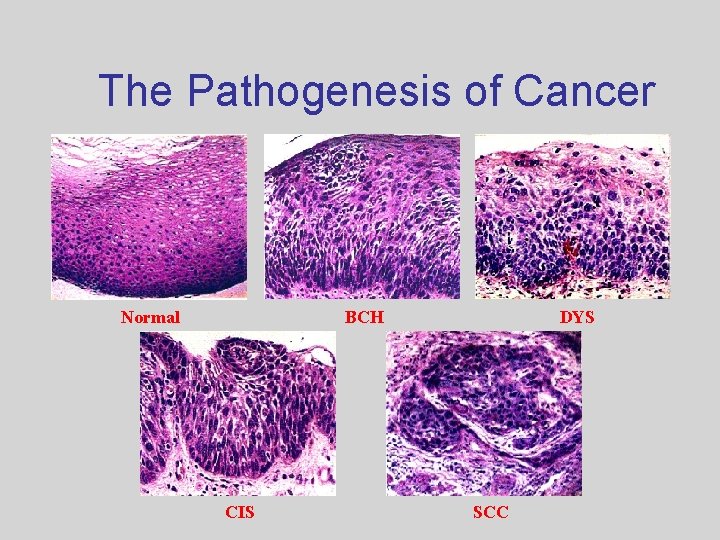
The Pathogenesis of Cancer Normal BCH CIS DYS SCC
- Slides: 28