Udine Lectures Lecture 1 Introduction to Biology Bud

Udine Lectures Lecture #1: Introduction to Biology ¦ Bud Mishra Professor of Computer Science and Mathematics (Courant, NYU) Professor (Watson School, CSHL) 7 ¦ 5 ¦ 2002 12/11/2021 ©Bud Mishra, 2002 1

Goal • The goal of this course is to understand, design and create a large-scale computational system centered on the biology of – – individual cells, population of cells, intra-cellular processes, and realistic simulation and visualization of these processes at multiple spatiotemporal scales. 12/11/2021 ©Bud Mishra, 2002 2

Why? • Such a reasoning system, in the hands of a working biologist, can then be used to – gain insight into the underlying biology, – design refutable biological experiments, and – ultimately, discover intervention schemes to suitably modify the biological processes for therapeutic purposes. • The course will focus primarily on two biological processes: – genome-evolution and – cell-to-cell communication. 12/11/2021 ©Bud Mishra, 2002 3

Genomics 12/11/2021 ©Bud Mishra, 2002 4

Introduction to Biology • Genome: – Hereditary information of an organism is encoded in its DNA and enclosed in a cell (unless it is a virus). All the information contained in the DNA of a single organism is its genome. • DNA molecule can be thought of as a very long sequence of nucleotides or bases: S = {A, T, C, G} 12/11/2021 ©Bud Mishra, 2002 5

Complementarity • DNA is a double-stranded polymer and should be thought of as a pair of sequences over S. However, there is a relation of complementarity between the two sequences: – A , T, C , G – That is if there is an A (respectively, T, C, G) on one sequence at a particular position the other sequence must have a T (respectively, A, G, C) at the same position. • We will measure the sequence length (or the DNA length) in terms of base pairs (bp): for instance, human (H. sapiens) DNA is 3. 3 £ 109 bp measuring about 6 ft of DNA polymer completely stretched out! 12/11/2021 ©Bud Mishra, 2002 6
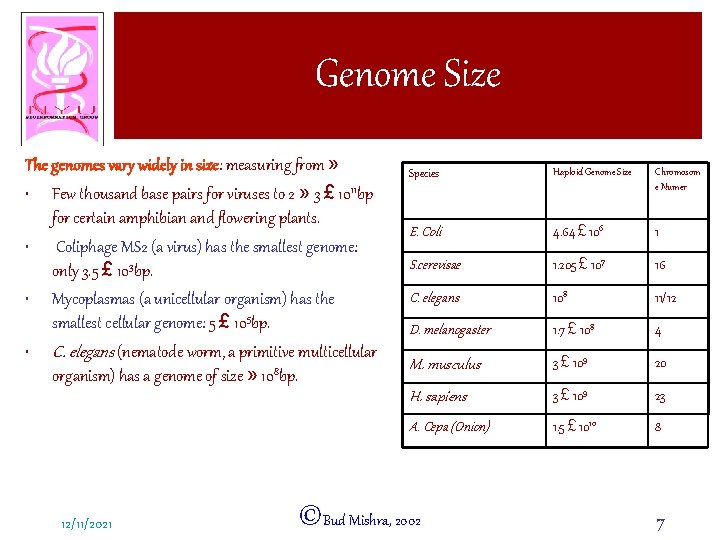
Genome Size The genomes vary widely in size: measuring from » • Few thousand base pairs for viruses to 2 » 3 £ 1011 bp for certain amphibian and flowering plants. • Coliphage MS 2 (a virus) has the smallest genome: only 3. 5 £ 103 bp. • Mycoplasmas (a unicellular organism) has the smallest cellular genome: 5 £ 105 bp. • C. elegans (nematode worm, a primitive multicellular organism) has a genome of size » 108 bp. 12/11/2021 Species Haploid Genome Size Chromosom e Numer E. Coli 4. 64 £ 106 1 S. cerevisae 1. 205 £ 107 16 C. elegans 108 11/12 D. melanogaster 1. 7 £ 108 4 M. musculus 3 £ 109 20 H. sapiens 3 £ 109 23 A. Cepa (Onion) 1. 5 £ 1010 8 ©Bud Mishra, 2002 7

DNA ) Structure and Components • The usual configuration of DNA is in terms of a double helix consisting of two chains or strands coiling around each other with two alternating grooves of slighltly different spacing. The “backbone” in each strand is made of alternating big sugar molecules (Deoxyribose residues: C 5 O 4 H 10) and small phosphate ((P O 4)-3) molecules. • Now, one of the four bases (the letters in our alphabet S), each one an almost planar nitrogenic organic compound, is connected to the sugar molecule. The bases are: – – Adenine ) A Thymine ) T Cytosine ) C Guanine ) G 12/11/2021 ©Bud Mishra, 2002 8

Genome in Detail The Human Genome at Four Levels of Detail. Apart from reproductive cells (gametes) and mature red blood cells, every cell in the human body contains 23 pairs of chromosomes, each a packet of compressed and entwined DNA (1, 2). 12/11/2021 ©Bud Mishra, 2002 9

DNA ) Structure and Components (contd. ) • The sequence of bases defines the information encoded by the DNA. • Complementary base pairs (A-T and C-G) are connected by hydrogen bonds and the base-pair forms a coplanar “rung” connecting the two strands. • – Cytosine and thymine are smaller (lighter) molecules, called pyrimidines – Guanine and adenine are bigger (bulkier) molecules, called purines. – Adenine and thymine allow only for double hydrogen bonding, while cytosine and guanine allow for triple hydrogen bonding. Thus the chemical (through hydrogen bonding) and the mechanical (purine to pyrimidine) constraints on the pairing lead to the complementarity and makes the double stranded DNA both chemically inert and mechanically quite rigid and stable. 12/11/2021 ©Bud Mishra, 2002 10
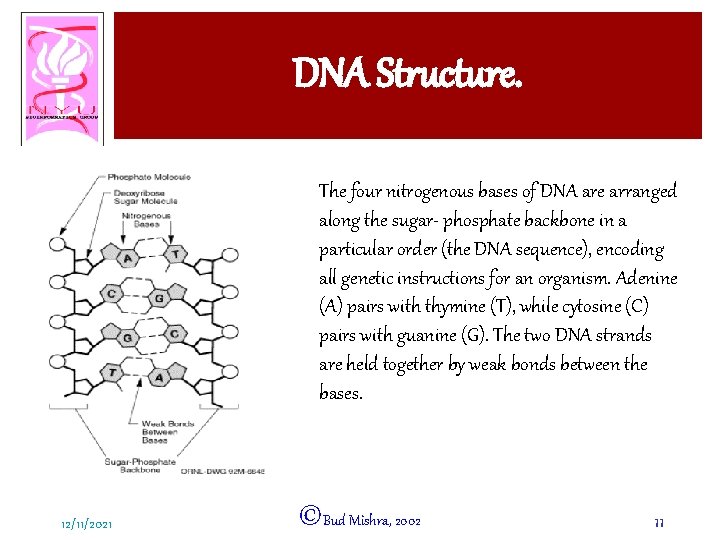
DNA Structure. The four nitrogenous bases of DNA are arranged along the sugar- phosphate backbone in a particular order (the DNA sequence), encoding all genetic instructions for an organism. Adenine (A) pairs with thymine (T), while cytosine (C) pairs with guanine (G). The two DNA strands are held together by weak bonds between the bases. 12/11/2021 ©Bud Mishra, 2002 11

DNA ) Structure and Components • • • (contd. ) The building blocks of the DNA molecule are four kinds of deoxyribonucleotides, – where each deoxyribonucleotide is made up of a sugar residue, a phosphate group and a base. – From these building blocks (or related, d. NTPs deoxyribonucleoside triphosphates) one can synthesize a strand of DNA. The sugar molecule in the strand is in the shape of a pentagon (4 carbons and 1 oxygen) in a plane parallel to the helix axis and with the 5 th carbon (5' C) sticking out. The phosphodiester bond (-O-P-O-) between the sugars connects this 5' C to a carbon in the pentagon (3' C) and provides a directionality to each strand. The strands in a double-stranded DNA molecule are antiparallel. Most of the enzymes moving along the backbone moves in the 5'-3' direction. 12/11/2021 ©Bud Mishra, 2002 12

The Central Dogma 12/11/2021 ©Bud Mishra, 2002 13

The Central Dogma • • • The intermediate molecule carrying the information out of the nucleus of an eukaryotic cell is RNA, a single stranded polymer. RNA also controls the translation process in which amino acids are created making up the proteins. The central dogma(due to Francis Crick in 1958) states that these information flows are all unidirectional: “The central dogma states that once `information' has passed into protein it cannot get out again. The transfer of information from nucleic acid to nucleic acid, or from nucleic acid to protein, may be possible, but transfer from protein to protein, or from protein to nucleic acid is impossible. Information means here the precise determination of sequence, either of bases in the nucleic acid or of amino acid residues in the protein. ” 12/11/2021 ©Bud Mishra, 2002 14

RNA and Transcription • • The polymer RNA (ribonucleic acid) is similar to DNA but differ in several ways: – it's single stranded; – its nucleotide has a ribose sugar (instead of deoxyribose) and – it has the pyrimidine base uracil, U, substituting thymine, T-- U is complementary to A like thymine. RNA molecule tends to fold back on itself to make helical twisted and rigid segments. – For instance, if a segment of an RNA is 5' - GGGGAAAACCCC - 3', – then the C's fold back on the G's to make a hairpin structure (with a 4 bp stem and a 5 bp loop). – The secondary RNA structure can even be more complicated, for instance, in case of E. coli, Ala t. RNA (transfer RNA) forms a cloverleaf shape. – Prediction of RNA structure is an interesting computational problem. 12/11/2021 ©Bud Mishra, 2002 15

RNA, Genes and Promoters • • • A specific region of DNA that determines the synthesis of proteins (through the transcription and translation) is called a gene – Originally, a gene meant something more abstract---a unit of hereditary inheritance. – Now a gene has been given a physical molecular existence. Transcription of a gene to a messenger RNA, m. RNA, is keyed by an RNA polymerase enzyme, which attaches to a core promoter (a specific sequence adjacent to the gene). Regulatory sequences such as silencers and enhancers control the rate of transcription – by their influence on the RNA polymerase through a feedback control loop involving many large families of activator and repressor proteins that bind with DNA and – which in turn, transpond the RNA polymerase by coactivator proteins and basal factors. 12/11/2021 ©Bud Mishra, 2002 16

Transcriptional Regulation • The entire structure of transcriptional regulation of gene expression is rather dispersed and fairly complicated: – The enhancer and silencer sequences occur over a wide region spanning many Kb's from the core promoter on either directions; – A gene may have many silencers and enhancers and can be shared among the genes; – They are not unique---different genes may have different combinations; – The proteins involved in control of the RNA polymerase number around 50 and – Different cliques of transcriptional factors operate in different cliques. • Any disorder in their properation can lead to cancer, immune disorder, heart disease, etc. 12/11/2021 ©Bud Mishra, 2002 17

Transcription • • The transcription of DNA in to m. RNA is performed with a single strand of DNA (the sense strand) around a gene. The double helix – Untwists momentarily to create a transcriptional bubble which moves along the DNA in the 3' - 5' direction (of the sense strand) – As the complementary m. RNA synthesis progresses adding one RNA nucleotide at a time at the 3' end of the RNA, attaching an U (respectively, A, G and C) for the corresponding DNA base of A (respectively, T, C and G), – Ending when a termination signal (a special sequence) is encountered. • • This newly synthesized m. RNA are capped by attaching special nucleotide sequences to the 5' and 3‘ ends. This molecule is called a pre-m. RNA. 12/11/2021 ©Bud Mishra, 2002 18
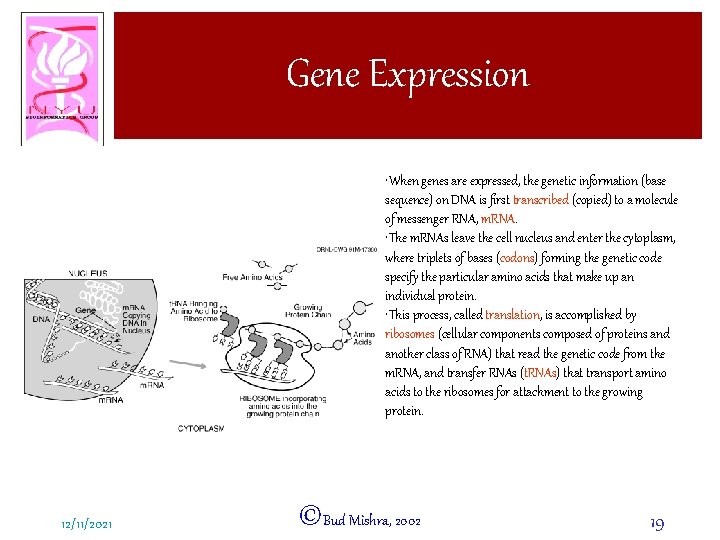
Gene Expression • When genes are expressed, the genetic information (base sequence) on DNA is first transcribed (copied) to a molecule of messenger RNA, m. RNA. • The m. RNAs leave the cell nucleus and enter the cytoplasm, where triplets of bases (codons) forming the genetic code specify the particular amino acids that make up an individual protein. • This process, called translation, is accomplished by ribosomes (cellular components composed of proteins and another class of RNA) that read the genetic code from the m. RNA, and transfer RNAs (t. RNAs) that transport amino acids to the ribosomes for attachment to the growing protein. 12/11/2021 ©Bud Mishra, 2002 19

The Genome Structure 12/11/2021 ©Bud Mishra, 2002 20

Exons and Introns • • In eukaryotic cells, the region of DNA transcribed into a pre-m. RNA involves more than just the information needed to synthesize the proteins. The DNA containing the code for protein are the exons, which are interrupted by the introns, the non-coding regions. Thus pre-m. RNA contains both exons and introns and is altered to excise all the intronic subsequences in preparation for the translation process---this is done by the spliceosome. The location of splice sites, separating the introns and exons, is dictated by short sequences and simple rules such as – “introns begin with the dinucleotide GT and end with the dinucleotide AG” (the GT-AG rule). 12/11/2021 ©Bud Mishra, 2002 21

Protein and Translation • • • The translation process begins at a particular location of the m. RNA called the translation start sequence (usually AUG) and is mediated by the transfer RNA (t. RNA), made up of a group of small RNA molecules, each with specificity for a particular amino acid. The t. RNA's carry the amino acids to the ribosomes, the site of protein synthesis, where they are attached to a growing polypeptide. The translation stops when one of the three trinucleotides UAA, UAG or UGA is encountered. Each 3 consecutive (nonoverlapping) bases of m. RNA (corresponding to a codon codes for a specific amino acid. There are 43 = 64 possible trinucleotide codons belonging to the set {U, A, G, C}3 12/11/2021 ©Bud Mishra, 2002 22

Genetic Codes • The codon AUG is the start codon and the codons UAA, UAG and UGA are the stop codons. – That leaves 60 codons to code for 20 amino acids with an expected redundancy of 3! – Multiple codons (one to six) are used to code a single amino acid. • The line of nucleotides between and including the start and stop codons is called an open reading frame (ORF) • All the information of interest to us resides in the ORF's. • The mapping from the codons to amino acid (and naturally extended to a mapping from ORF's polypeptides by a homomorphism) given by FP : {U, A, G, C}3 ! {A, R, D, N, C, E, Q, G, H, I, L, K, M, F, P, S, T, W, Y, V} 12/11/2021 ©Bud Mishra, 2002 23
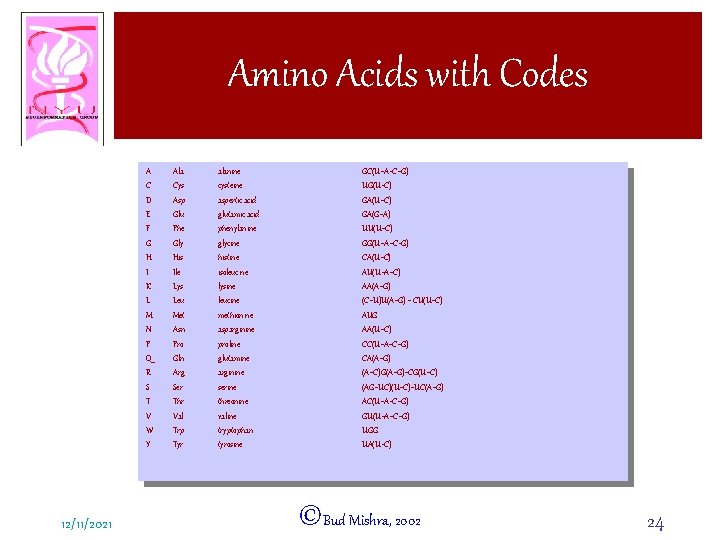
Amino Acids with Codes A C D E F G H I K L M N P Q R S T V W Y 12/11/2021 Ala Cys Asp Glu Phe Gly His Ile Lys Leu Met Asn Pro Gln Arg Ser Thr Val Trp Tyr alanine cysteine aspertic acid glutamic acid phenylanine glycine histine isoleucine lysine leucine methionine asparginine proline glutamine arginine serine threonine valine tryptophan tyrosine GC(U+A+C+G) UG(U+C) GA(G+A) UU(U+C) GG(U+A+C+G) CA(U+C) AU(U+A+C) AA(A+G) (C+U)U(A+G) + CU(U+C) AUG AA(U+C) CC(U+A+C+G) CA(A+G) (A+C)G(A+G)+CG(U+C) (AG+UC)(U+C)+UC(A+G) AC(U+A+C+G) GU(U+A+C+G) UGG UA(U+C) ©Bud Mishra, 2002 24

Interrupted Genes: • An open reading frame (containing a gene) consists of – INTRONS: Intervening sequences a Noncoding regions – EXONS: Protein coding regions • Introns are abundant in eukaryotes and certain animal viruses. 12/11/2021 ©Bud Mishra, 2002 25
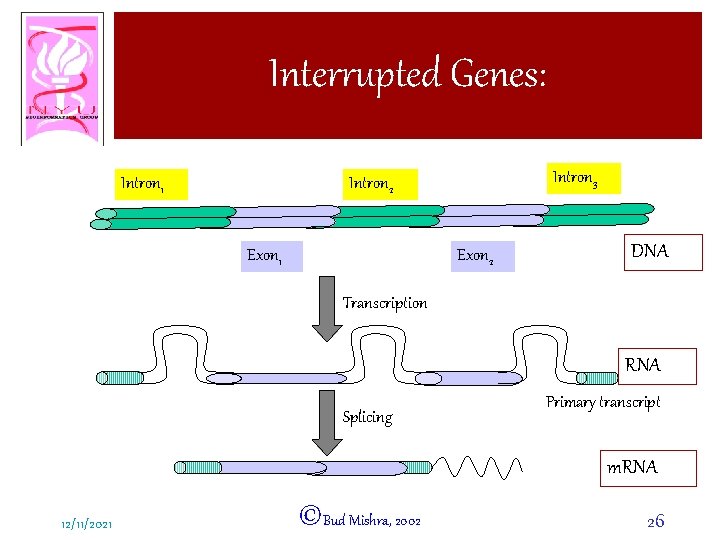
Interrupted Genes: Intron 1 Intron 3 Intron 2 Exon 1 Exon 2 DNA Transcription RNA Splicing Primary transcript m. RNA 12/11/2021 ©Bud Mishra, 2002 26
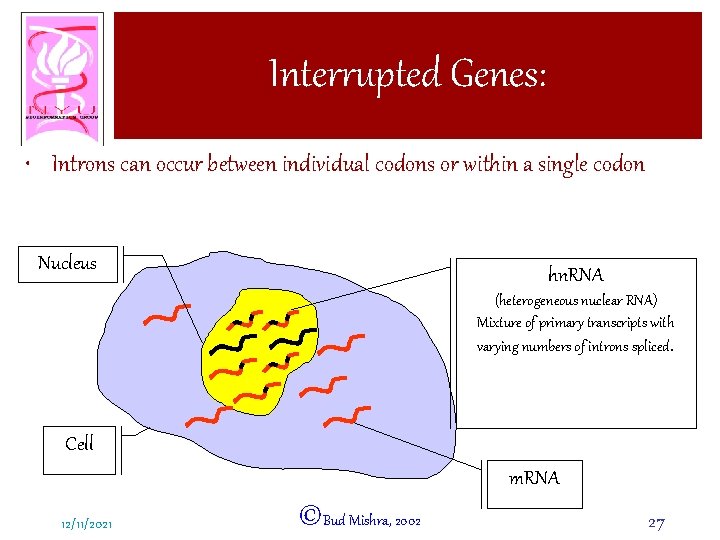
Interrupted Genes: • Introns can occur between individual codons or within a single codon Nucleus hn. RNA (heterogeneous nuclear RNA) Mixture of primary transcripts with varying numbers of introns spliced. Cell m. RNA 12/11/2021 ©Bud Mishra, 2002 27
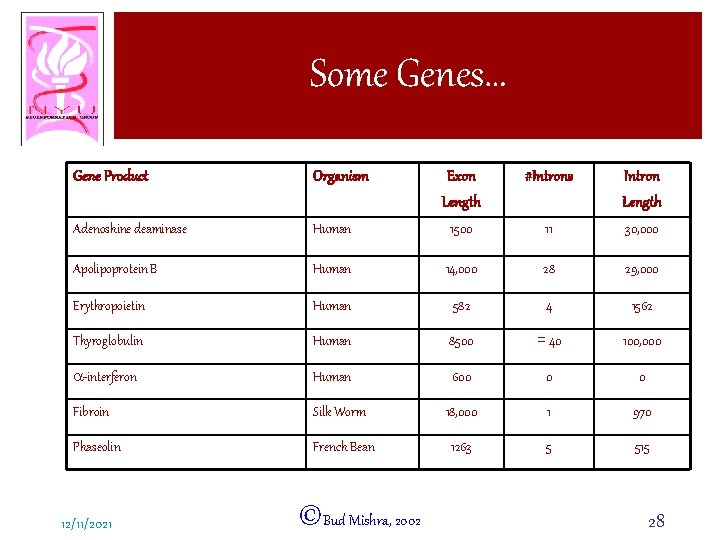
Some Genes… Gene Product Organism Exon Length #Introns Intron Length Adenoshine deaminase Human 1500 11 30, 000 Apolipoprotein B Human 14, 000 28 29, 000 Erythropoietin Human 582 4 1562 Thyroglobulin Human 8500 = 40 100, 000 a-interferon Human 600 0 0 Fibroin Silk Worm 18, 000 1 970 Phaseolin French Bean 1263 5 515 12/11/2021 ©Bud Mishra, 2002 28
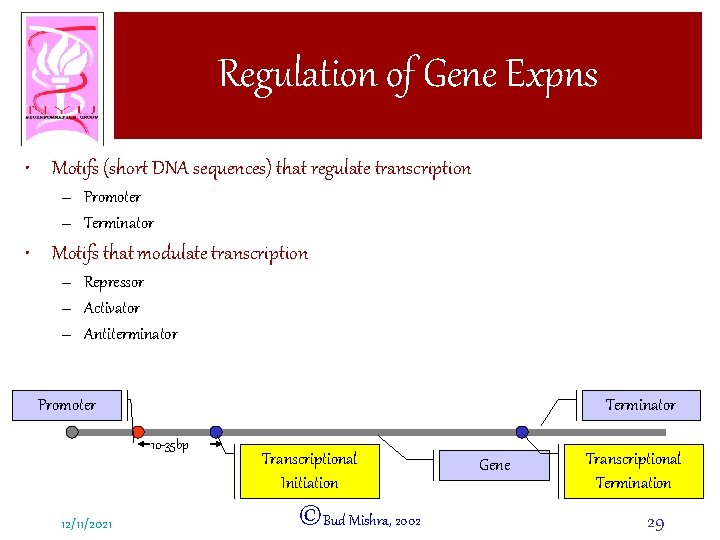
Regulation of Gene Expns • Motifs (short DNA sequences) that regulate transcription – Promoter – Terminator • Motifs that modulate transcription – Repressor – Activator – Antiterminator Promoter Terminator 10 -35 bp 12/11/2021 Transcriptional Initiation ©Bud Mishra, 2002 Gene Transcriptional Termination 29

Promoters • pol I (RNA polymerase I) – Transcribes ribosomal RNA genes 100 » 1000 bp in front of the gene • pol II (RNA polymerase II) – Transcribes genes encoding polypeptides – Complex and variable regulatory regions • pol III (RNA polymerase III) – Transcribes transfer RNA and other small RNAs – Both up and down stream 12/11/2021 ©Bud Mishra, 2002 30

Motifs • Each motif is a binding site for a specific protein • Transcription Factor: – Transcription factors (specific to a cell/environmental conditions) bind to regulatory regions and facilitate • Assembly of RNA polymerase into a transcriptional complex • Activation of a transcriptional complex. • Termination Factor: • – Assembly of proteins for termination and modification of the end of the RNA Epigenetic Changes – Methylation of the cytosine in the 5’ region – Structural changes in cromatin 12/11/2021 ©Bud Mishra, 2002 31

Organization of Genetic Info • Bacterial Genome: – Genes are closely spaced along the DNA. – The sequences of genes may overlap. – Related genes (encoding enzymes whose functions are part of the same pathway or whose activities are related) are linked as a single transcription unit. 12/11/2021 ©Bud Mishra, 2002 32

Organization of Genetic Info • Eukaryotic Genome: – – – Genes are separated by long stretches of noncoding DNA sequences. Multiple genes in a single transcription unit is extremely rare. Multiple chromosomes – Linear Chloroplasts and mitochondria – Circular Genes appearing on the same chromosome are syntenic. 12/11/2021 ©Bud Mishra, 2002 33
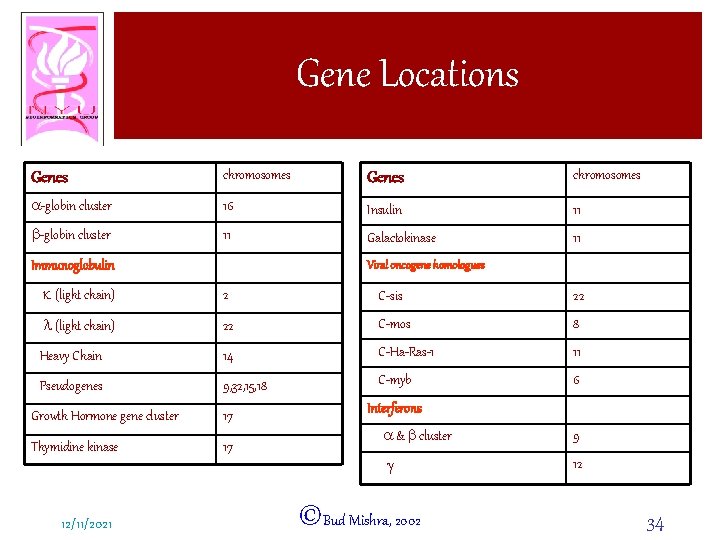
Gene Locations Genes chromosomes a-globin cluster 16 Insulin 11 b-globin cluster 11 Galactokinase 11 Immunoglobulin Viral oncogene homologues k (light chain) 2 C-sis 22 l (light chain) 22 C-mos 8 Heavy Chain 14 C-Ha-Ras-1 11 Pseudogenes 9, 32, 15, 18 C-myb 6 Growth Hormone gene cluster 17 Thymidine kinase 17 12/11/2021 Interferons a & b cluster 9 g 12 ©Bud Mishra, 2002 34

Eukaryotic Genome • Multiple copies of the same gene – Solve “supply problem” – There are several hundred ribosomal RNA genes I mammals • Pseudogenes – Nonfunctional copies of genes…(Deletions or alterations in the DNA sequence) – Number of pseudo genes for a particular gene varies greatly…Different from one organism to another. 12/11/2021 ©Bud Mishra, 2002 35

Genes in Eukaryotes • A gene may appear exactly once • It may be part of a family of repeated sequence. Members of a family may be clustered or dispersed. • Members of a gene family may be related and functional (expressed at different times in development, or in different cells) or may be pseudo genes. • Chromosomal Morphology: – Nucleolar organizers (genes for ribosomal RNA) – Telomeric and Centromeric regions (Tandemly repeated sequences) 12/11/2021 ©Bud Mishra, 2002 36

Genome Rearrangement • Reshuffling of genes between homologous chromosomes via reciprocal crossing-over during both meiosis and mitosis. • Gene synteny and linkages are usually preserved. • Most rearrangements are random. • Some rearrangements are normal processes altering gene expressions in an orderly and programmed manner. 12/11/2021 ©Bud Mishra, 2002 37

Chromosomal Aberrations • • Breakage Translocation (Among non-homologous chromosomes. ) Formation of acentric and dicentric chromosomes. Gene Conversions Amplification and deletions Point mutations Jumping genes a Transposition of DNA segments Programmed rearrangements a E. g. , antibody responses. 12/11/2021 ©Bud Mishra, 2002 38

Repeat Structure • Copy Number: 2 » 106 • Direct Repeats “head-to-tail” – Tandem repeats or separated by other sequences • Inverted Repeats “head-to-head” – Stem-and-loop structure – Hairpin structure • Reverse Palindrome • True Palindrome 12/11/2021 ©Bud Mishra, 2002 39
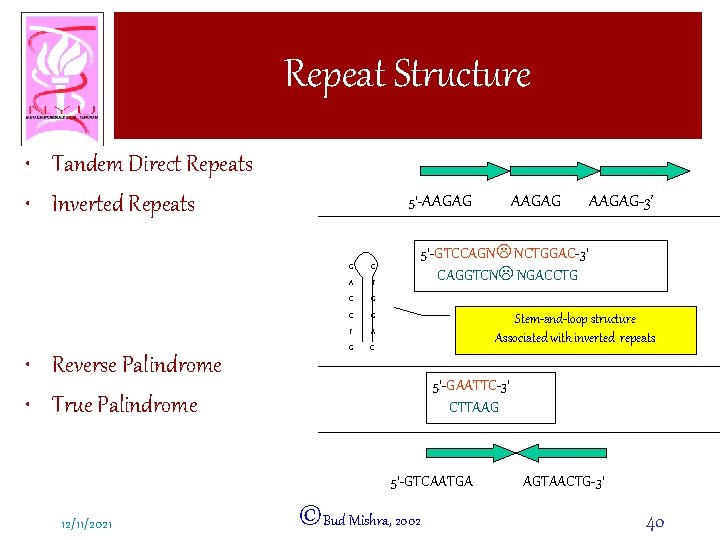
Repeat Structure • Tandem Direct Repeats • Inverted Repeats • Reverse Palindrome • True Palindrome 5’-AAGAG G C A T C G T A G C AAGAG-3’ 5’-GTCCAGNL NCTGGAC-3’ CAGGTCNL NGACCTG Stem-and-loop structure Associated with inverted repeats 5’-GAATTC-3’ CTTAAG 5’-GTCAATGA 12/11/2021 AAGAG ©Bud Mishra, 2002 AGTAACTG-3’ 40

Repeats within the Genome • Gene Family – Genes and its cognate pseudogenes • Satellite: Repeats made of noncoding units – Minisatellites: Tandem repeats…Mostly in centromeric regions – Satellite repeat units vary in length freom 2 base pairs to several thousands. 12/11/2021 ©Bud Mishra, 2002 41

Interspersed Repeats • SINES: Short Interspersed Repeats – Each repeat unit is of length 100 – 500 bps – Processed pseudogenes derived from class III genes – Example: Alu repeats…dimeric head-to-tail repeats of 130 bp • LINES: Long Interspersed Repeats – Each unit is of length > 6 Kb. 12/11/2021 ©Bud Mishra, 2002 42

The Cell • A cell is a small coalition of a set of genes held together in a set of chromosomes (and even perhaps unrelated extrachromosomal elements). • They also have set of machinery made of proteins, enzymes, lipids and organelles taking part in a dynamic process of information processing. – In eukaryotic cells the genetic materials are enclosed in the cell nucleus separated from the other organelles in the cytoplasm by a membrane. – In prokaryotic cells the genetic materials are distributed homogeneously as it does not have a nucleus. – Example of prokaryotic cells are bacteria with a considerably simple genome. 12/11/2021 ©Bud Mishra, 2002 43

Organelles • The organelles common to eukaryotic plant and animal cells include – Mitochondria in animal cells and chloroplasts in plant cells (responsible for energy production); – A Golgi apparatus (responsible for modifying, sorting and packaging various macromolecules for distribution within and outside the cell); – Endpolastic reticulum (responsible for synthesizing protein); and – Nucleus (responsible for holding the DNA as chromosomes and replication and transcription). 12/11/2021 ©Bud Mishra, 2002 44

Chromosomes • The entire cell is contained in a sack made of plasma membrane. In plant cells, they are further surrounded by a cellulose cell wall. • The nucleus of the eukaryotic cells contain its genome in several chromosomes, where each chromosome is simply a single molecule of DNA as well as some proteins (primarily histones). 12/11/2021 ©Bud Mishra, 2002 45

Chromosomes • The chromosomes can be a circular molecule or linear, in which case the ends are capped with special sequence of telomeres. • The protein in the nucleus binds to the DNA and effects the compaction of the very long DNA molecules. • In somatic cells (as opposed to gametes: egg and sperm cells) of most eukaryotic organisms, the chromosomes occur in homologous pairs, with the only exceptions being the X and Y chromosomes---sex chromosomes. 12/11/2021 ©Bud Mishra, 2002 46
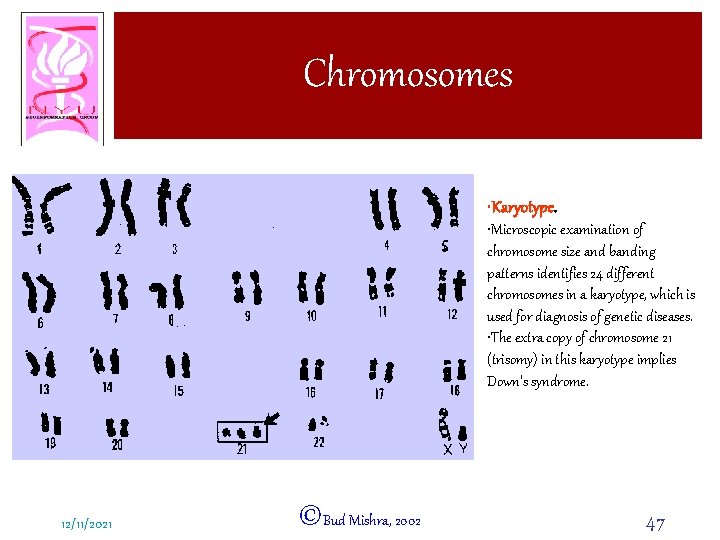
Chromosomes • Karyotype. • Microscopic examination of chromosome size and banding patterns identifies 24 different chromosomes in a karyotype, which is used for diagnosis of genetic diseases. • The extra copy of chromosome 21 (trisomy) in this karyotype implies Down's syndrome. 12/11/2021 ©Bud Mishra, 2002 47

Ploidy • Gametes contain only unpaired chromosomes; the egg cell contains only X chromosome and the sperm cell either an X or an Y chromosome. The male has X and Y chromosomes; the female, 2 X's. • Cells with single unpaired chromosomes are called haploid; the cells with homologous pairs, diploid; the cells with homologous triplet, quadruplet, etc. , chromosomes are called polyploid---many plant cells are polyploid. 12/11/2021 ©Bud Mishra, 2002 48

The dynamics of cell: • The cell cycle ) the set of events that occur within a cell between its birth by mitosis and its division into daughter cells again by mitosis – interphase period when DNA is synthesized and – mitotic phase • The cell division by mitosis (into 2 daughter cells) and meiosis (into 4 gametes from germ-line cells); • Working of the machinery within the cell---mainly the ones involving replication of DNA, transcription of DNA into RNA and translation of RNA into protein. 12/11/2021 ©Bud Mishra, 2002 49

The Cell Cycle 12/11/2021 ©Bud Mishra, 2002 50

The Cell Cycle: • In growing cells, the four phases proceed successively, taking from 10 -20 hrs. • Interphase: comprises the G 1, S, and G 2 phases. DNA is synthesized in S and other cellular macromolecules are synthesized throughout interphase, roughly doubling cell’s mass. • During G 2 the cell is prepared for mitotic (M) phase when the genetic material is evenly proportioned and the cell divides. G 0 M G 1 G 2 S • Nondividing cells exit the normal cycle, entering the quiesecent G 0 state. 12/11/2021 ©Bud Mishra, 2002 51

Differentiation & Suicide • Cellular dynamics controls how a cell changes (or differentiates) to carry out a specialized functions – Structural or morphological changes (muscles, neural, skin. . ) – Immune systems: Many cell types come together in organized tissues designed to let the body distinguish self from non-self. • Programmed Cell Death/Apoptosis: – Condensation of the nucleus. – Fragmentation of the DNA. – Morphological changes followed by consumption by macrophages. 12/11/2021 ©Bud Mishra, 2002 52

Cell Talk 12/11/2021 ©Bud Mishra, 2002 53
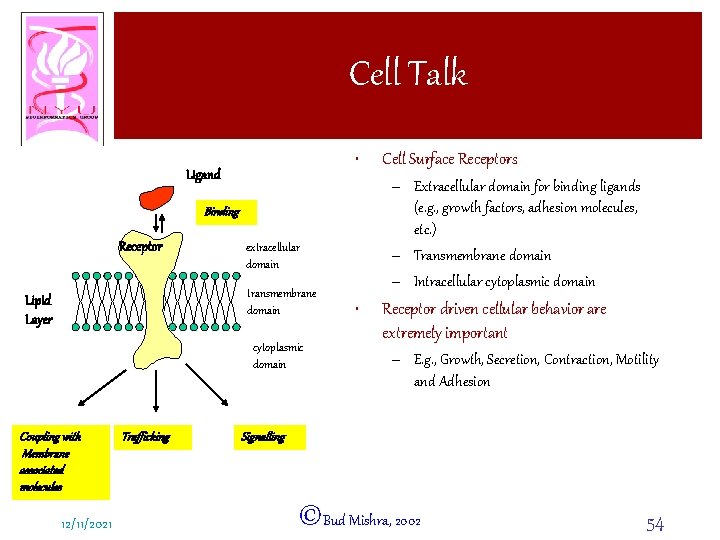
Cell Talk • Ligand – Extracellular domain for binding ligands (e. g. , growth factors, adhesion molecules, etc. ) – Transmembrane domain – Intracellular cytoplasmic domain Binding Receptor extracellular domain transmembrane domain Lipid Layer cytoplasmic domain Coupling with Membrane associated molecules 12/11/2021 Trafficking Cell Surface Receptors • Receptor driven cellular behavior are extremely important – E. g. , Growth, Secretion, Contraction, Motility and Adhesion Signalling ©Bud Mishra, 2002 54
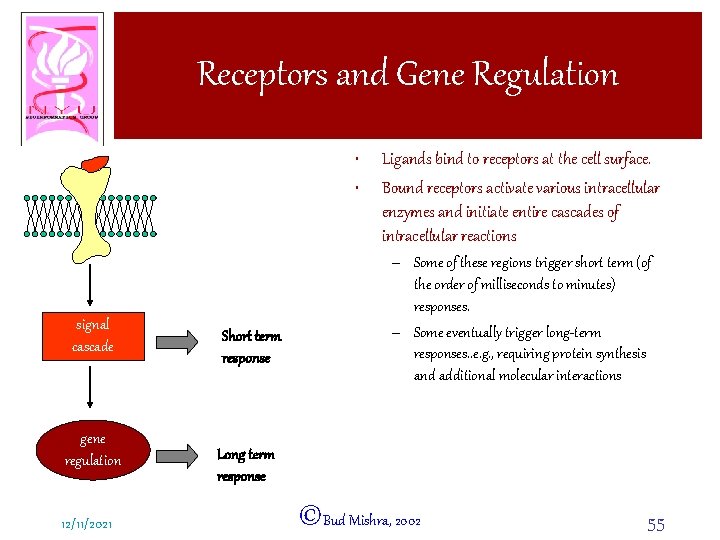
Receptors and Gene Regulation • • signal cascade gene regulation 12/11/2021 Short term response Ligands bind to receptors at the cell surface. Bound receptors activate various intracellular enzymes and initiate entire cascades of intracellular reactions – Some of these regions trigger short term (of the order of milliseconds to minutes) responses. – Some eventually trigger long-term responses. . e. g. , requiring protein synthesis and additional molecular interactions Long term response ©Bud Mishra, 2002 55
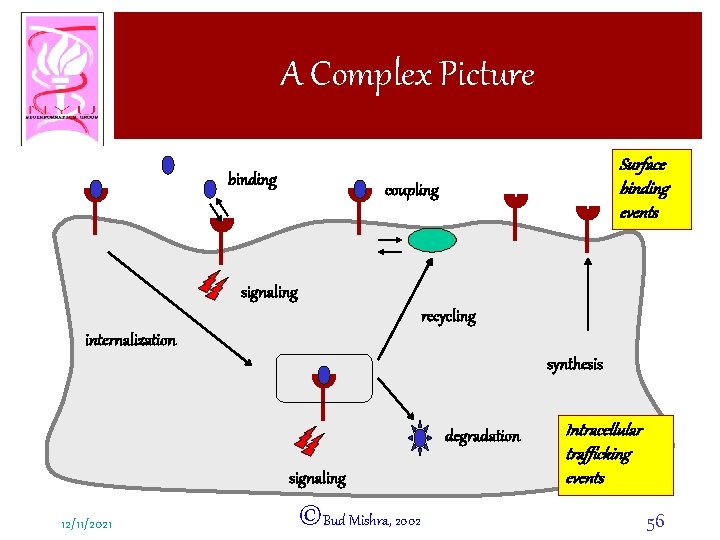
A Complex Picture binding Surface binding events coupling signaling recycling internalization synthesis degradation signaling 12/11/2021 ©Bud Mishra, 2002 Intracellular trafficking events 56

A Complex Picture • Trafficking – Receptor population undergoes many complex events of coupling with other cell surface molecules – Internalization (RME: receptor-mediated endocytosis) – Recycling – Degradation – Synthesis 12/11/2021 ©Bud Mishra, 2002 57
- Slides: 57