Computational Biology Lecture 4 Mapping Sequencing Bud Mishra

Computational Biology Lecture #4: Mapping & Sequencing Bud Mishra Professor of Computer Science and Mathematics 10 ¦ 1 ¦ 2002 9/6/2021 ©Bud Mishra, 2001 1
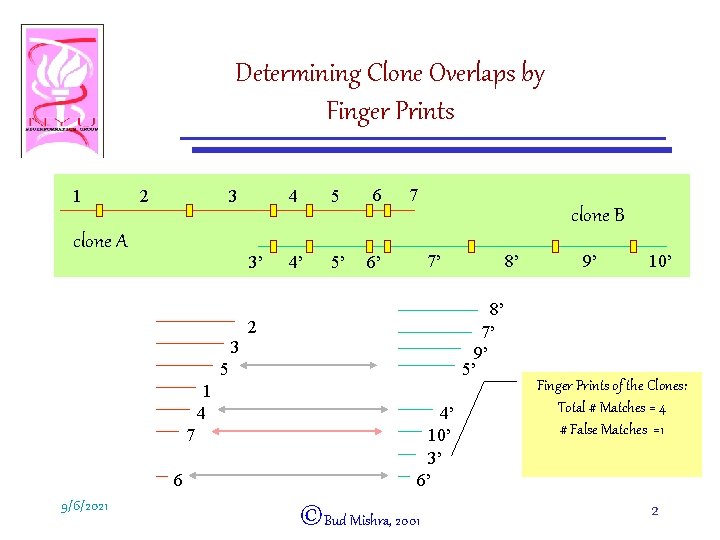
Determining Clone Overlaps by Finger Prints 1 2 3 clone A 3’ 4 5 6 4’ 5’ 6’ 7 clone B 7’ 8’ 7’ 9’ 5’ 2 3 5 1 4 7 6 9/6/2021 8’ 4’ 10’ 3’ 6’ ©Bud Mishra, 2001 9’ 10’ Finger Prints of the Clones: Total # Matches = 4 # False Matches =1 2

Matching Rule • Two clones A and B overlap if they have at least k restriction fragments in “common. ” • Question: What is the probability that two randomly chosen unrelated clones may get determined to overlap? • n = Expected # restriction fragments in a clone. • W = # restriction fragments in clone A such that they all match with fragments in clone B. 9/6/2021 ©Bud Mishra, 2001 3

False Positives Accidental Matching • Consider the set of “Clone. A i-fragments” (meaning i many restriction fragments from clone A’s finger print) • There are Cn, i such sets. • Define an indicator variable, Xj (1 · j · Cn, i): Xj = {1, if the jth set all match with restriction fragments of Clone B {0, otherwise. • CW, i = åj=1 Cn, i Xj 9/6/2021 ©Bud Mishra, 2001 4
![Applying Brun’s Sieve • E[CW, i] = Cn, i [n (b/2)][(n-1)(b/2)] L [(n-i+1)(b/2)] ¼ Applying Brun’s Sieve • E[CW, i] = Cn, i [n (b/2)][(n-1)(b/2)] L [(n-i+1)(b/2)] ¼](http://slidetodoc.com/presentation_image_h2/8bbe73d145c83316dd08c592cb3121a6/image-5.jpg)
Applying Brun’s Sieve • E[CW, i] = Cn, i [n (b/2)][(n-1)(b/2)] L [(n-i+1)(b/2)] ¼ (b n 2/2)i/i! • By Brun’s Sieve Pr[W=i] = (1/i!) (b n 2/2)i exp(-b n 2/2) • W » Poisson(b n 2/2) • Thus the false positive probability can be made negligibly small by choosing the parameter k ¸ 3 b n 2/4 9/6/2021 ©Bud Mishra, 2001 5

Efficacy of Finger Prints • The relative sizing error in finger prints should be small: b · 2/(3 n) • Finger print methods is applicable only when the expected number of fragments (n) is rather small: E. g. , if n = 10, then b · 6. 6% • For large clones (e. g. , BAC and YAC) only rare cutters can be used. 9/6/2021 ©Bud Mishra, 2001 6

A Graph Representation for the Overlap Information • Let: C = {c 1, c 2, …, c. N} be a set of clones (all of equal length L) in a clone library covering a genome of length G. • G = G/L = Normalized Genome Length • Assume perfect overlap information. That is – No false positive: ci Å cj = ; ! : overlap(ci, cj) – No false negative: ci Å cj ¹ ; ! overlap(ci, cj) 9/6/2021 ©Bud Mishra, 2001 7

Overlap Graph • Overlap Graph: G = (V, E) = Undirected graph V = {v 1, v 2, …, v. N} EµV£V [vi, vj] 2 E , Overlap(ci, cj) • Interval Graph Problem: – Given: An undirected Graph G = (V, E) – Decide: If G is an interval graph. Can the vertices of G be represented by N unit intervals {I 1, I 2, …, IN} 2 R 1 such that [vi, vj] 2 E iff Ii Å Ij ¹ ; . � 9/6/2021 ©Bud Mishra, 2001 8
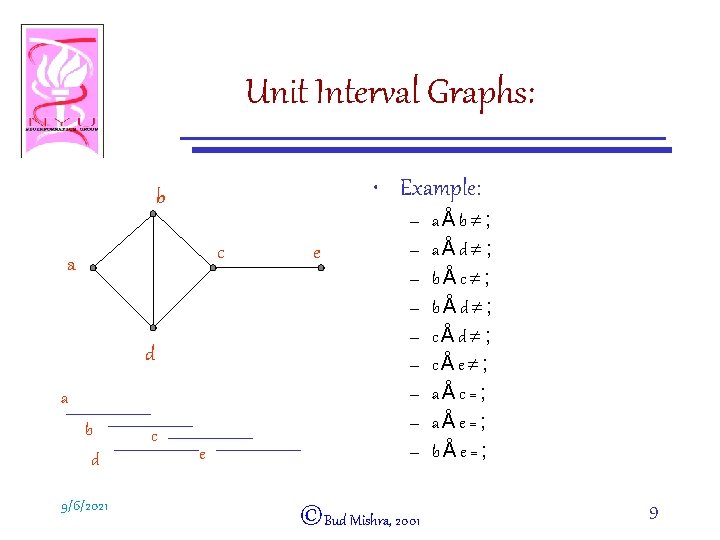
Unit Interval Graphs: • Example: b c a d a b d 9/6/2021 c e e – – – – – ©Bud Mishra, 2001 aÅb¹; aÅd¹; bÅc¹; bÅd¹; cÅe¹; aÅc=; aÅe=; bÅe=; 9
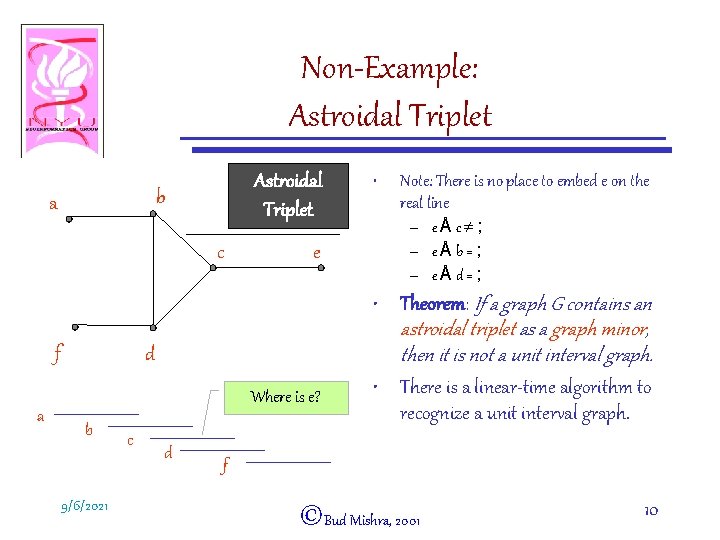
Non-Example: Astroidal Triplet b a c f a e d Where is e? b 9/6/2021 c d • Note: There is no place to embed e on the real line – eÅc¹; – eÅb=; – eÅd=; • Theorem: If a graph G contains an astroidal triplet as a graph minor, then it is not a unit interval graph. • There is a linear-time algorithm to recognize a unit interval graph. f ©Bud Mishra, 2001 10
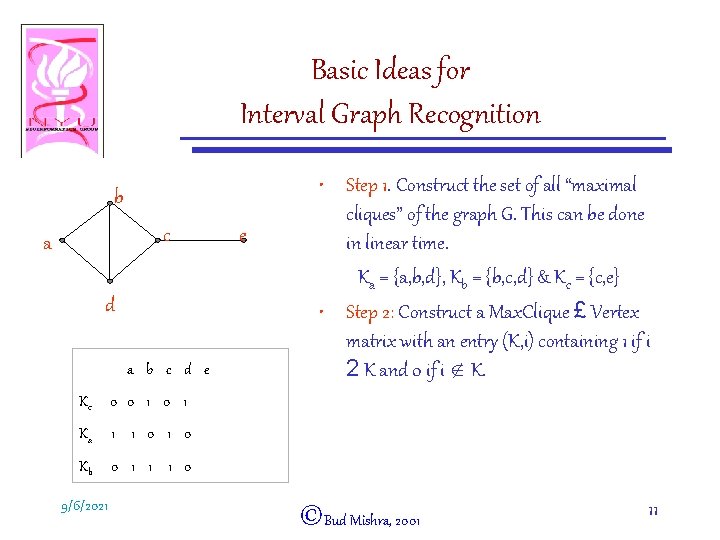
Basic Ideas for Interval Graph Recognition b c a d a b c d e Kc 0 0 1 Ka 1 1 0 Kb 0 1 1 1 0 9/6/2021 e • Step 1. Construct the set of all “maximal cliques” of the graph G. This can be done in linear time. Ka = {a, b, d}, Kb = {b, c, d} & Kc = {c, e} • Step 2: Construct a Max. Clique £ Vertex matrix with an entry (K, i) containing 1 if i 2 K and 0 if i Ï K. ©Bud Mishra, 2001 11
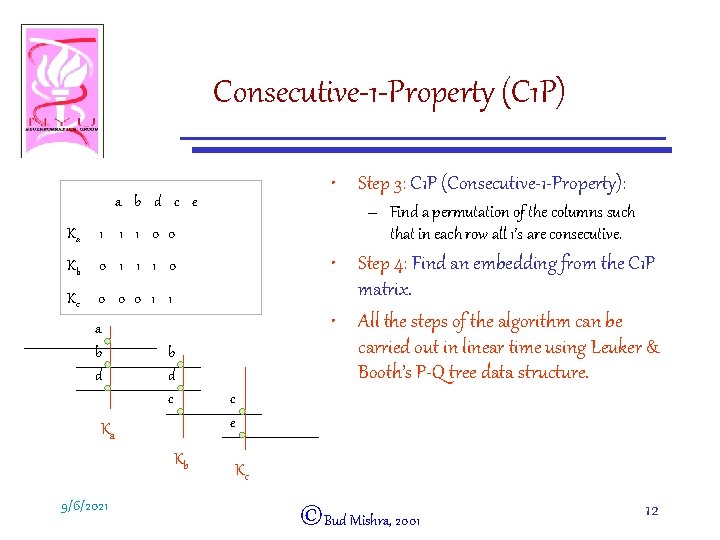
Consecutive-1 -Property (C 1 P) • Step 3: C 1 P (Consecutive-1 -Property): a b d c e Ka 1 1 1 0 0 Kb 0 1 1 1 0 Kc 0 0 0 1 1 a b d c Ka Kb 9/6/2021 – Find a permutation of the columns such that in each row all 1’s are consecutive. • Step 4: Find an embedding from the C 1 P matrix. • All the steps of the algorithm can be carried out in linear time using Leuker & Booth’s P-Q tree data structure. c e Kc ©Bud Mishra, 2001 12

A More Realistic Model • Errors: – Some of the overlaps are false positive… – Some of the non-overlaps are false negative… • Model: Two undirected graphs on the same vertices, V – Gsure = (V, Esure) a If [vi, vj] 2 Esure then ci Å cj ¹ ; surely… – Gunsure = (V, Eunsure) a If [vi, vj] 2 Eunsure then ci Å cj ¹ ; with some uncertainty… • Interval Graph Sandwich Problem (IGSP) – Does there exist an interval graph G = (V, E) such that Esure µ Eunsure? – ISGP is NP-complete. 9/6/2021 ©Bud Mishra, 2001 13

Mapping with Approximate Overlaps • Overlap Parameter: T – T= # basepairs 2 clones must have in common to ensure overlap. – | ci Å cj | ¸ T , overlap(ci, cj) • L = Length of the clone – q = (T/L) = Overlap Threshold Ratio – s = (1 -q) L L 9/6/2021 ©Bud Mishra, 2001 =T 14

Islands and Contigs – G = Genome length – L = Length of a clone – N = Number of clones • • • a = (N/G) = Probability of a clone starting at a given site. c = LN/G = coverage q = overlap threshold ratio, s = (1 -q) L-T = L(1 -q)=cs/a = “overhang” g = c s = “effective coverage” 9/6/2021 ©Bud Mishra, 2001 15
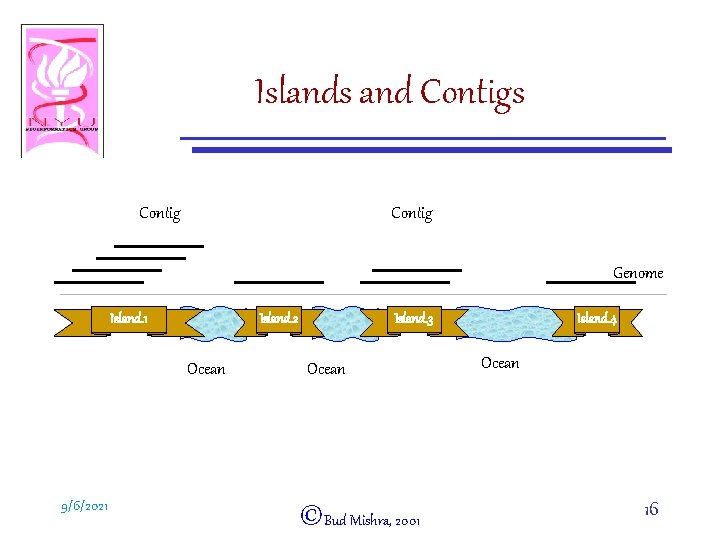
Islands and Contigs Contig Genome Island. 1 Island. 2 Ocean 9/6/2021 Island. 3 Ocean ©Bud Mishra, 2001 Island. 4 Ocean 16

Some Definitions • Island: A maximal set of clones, closed under the reflexive and transitive closure of the relation induced by the overlap rule. • Trivial Contig = Singleton Island: – An island containing exactly one clone • (Nontrivial) Contig: An island containing at least two clones. • Ocean: A region of the genome between two neighboring islands. 9/6/2021 ©Bud Mishra, 2001 17

Sij = Indicator Variable • Sij = {1, if clone Ci starts at position j; {0, otherwise. 2 k 1 a L-T 9/6/2021 T • Va = åi=1 N åa-L+Ta-1 Sij =Random Variable denoting # clones covering position a from the left by an amount no larger than L-T. ©Bud Mishra, 2001 18
![Statistics • E[CVa, k] = C(L-T)N, k (1/G)k ¼ ((L-T)N/G)k/k! = (c-q c)k/k! = Statistics • E[CVa, k] = C(L-T)N, k (1/G)k ¼ ((L-T)N/G)k/k! = (c-q c)k/k! =](http://slidetodoc.com/presentation_image_h2/8bbe73d145c83316dd08c592cb3121a6/image-19.jpg)
Statistics • E[CVa, k] = C(L-T)N, k (1/G)k ¼ ((L-T)N/G)k/k! = (c-q c)k/k! = (c s)k/k! = gk/k! • By Brun’s Sieve: Pr[Va = k] = e- g gk/k! • Pr[Va = 0] = e-cs = e-g ( Probability that a belongs to an apparent ocean. • Pr[Va ¹ 0] = 1 – e-c s= 1 - e-g ( Probability that a belongs to an apparent island. 9/6/2021 ©Bud Mishra, 2001 19
![Number of Islands • Ia = Pr[(Va = 0) Æ 9 ci (sia=1)] = Number of Islands • Ia = Pr[(Va = 0) Æ 9 ci (sia=1)] =](http://slidetodoc.com/presentation_image_h2/8bbe73d145c83316dd08c592cb3121a6/image-20.jpg)
Number of Islands • Ia = Pr[(Va = 0) Æ 9 ci (sia=1)] = a e-g ( Probability that an island begins at a. • Expected # Islands = Expected # Oceans = åa=1 G Ia= G a e-c s = N e-g • Expected # Singleton Islands = N e-2 g • Expected # Contigs = N e-g - N e-2 g 9/6/2021 ©Bud Mishra, 2001 20

Normalized Values • Expected # Islands £ (L/G) = c e-c(1 -q) • Expected # Singleton Islands £ (L/G) = c e-2 c(1 -q) • Expected # Contigs £ (L/G) = c [e-c(1 -q) - e-2 c(1 -q) ] 9/6/2021 ©Bud Mishra, 2001 21
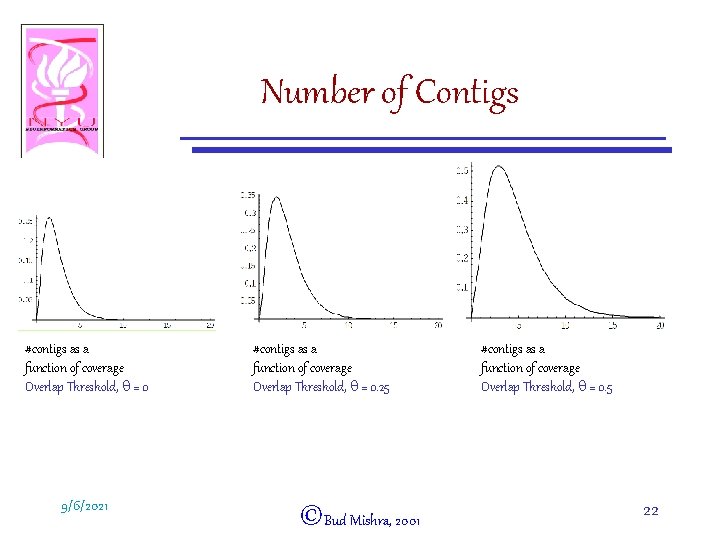
Number of Contigs #contigs as a function of coverage Overlap Threshold, q = 0 9/6/2021 #contigs as a function of coverage Overlap Threshold, q = 0. 25 ©Bud Mishra, 2001 #contigs as a function of coverage Overlap Threshold, q = 0. 5 22

Contigs (Dependence on coverage and overlap threshold) • X-axis a c = coverage • Y-axis a q = Overlap threshold • z-axis a # of contigs 9/6/2021 ©Bud Mishra, 2001 23

The Single Molecule Approach • New approaches to manipulate single macromolecules (proteins & oligonucleotides) – Optical Mapping (NYU/Courant/Wisconsin) – Molecular Combing (Pasteur) – Optical Tweezers/ Optical Traps (Stanford) – Bacterial Ion Channel (Harvard) – Fluorescent Flow Cytometry (Caltech) 9/6/2021 ©Bud Mishra, 2001 24

Optical Approaches • Single molecule on a surface can be explored by nanometer scale with tunneling electrons, forces from sharp tips or magnetic resonance: – STM (scanning tunneling microscope) – AFM (atomic force microscopy) – MRFM (magnetic resonance force microscopy) • Optical Approaches: non-invasive, avoids synchronization, need not be real-time. 9/6/2021 ©Bud Mishra, 2001 25

Optical Approaches are Inherently Noisy! • Since many biological macromolecules are smaller than the Raleigh limit, the optical approaches involve attaching single fluorescent probes to specific macromolecules. – – – 9/6/2021 Controlling Noise: Magnitude of Stoke-shift Steric hinderance Absorption cross-section Point spread function (PSF) Image Processing ©Bud Mishra, 2001 26

Error Sources • Sizing Error – (Bernoulli labeling, absorption crosssection, PSF) • • 9/6/2021 Partial Digestion False Optical Sites Orientation Spurious molecules, Optical chimerism, Calibration ©Bud Mishra, 2001 27

Shotgun Mapping • Large fragments of genomic DNA of length from 2 Mb to 12 Mb are optically mapped • The resulting ordered restriction maps are automatically contiged by “Gentig” • The consensus map computed by Gentig is free of errors due to partial digestion, sizing error and false cuts 9/6/2021 ©Bud Mishra, 2001 28
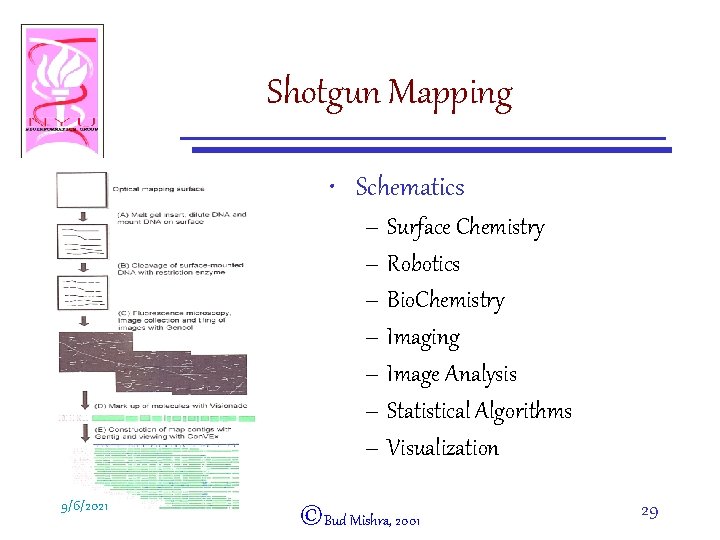
Shotgun Mapping • Schematics – Surface Chemistry – Robotics – Bio. Chemistry – Imaging – Image Analysis – Statistical Algorithms – Visualization 9/6/2021 ©Bud Mishra, 2001 29

Overlap Rule • Comparing Two Genomic Restriction Maps: Given two maps A and B, we say that they overlap, if --1. k or more of the restriction fragments align positionally (subject to sizing error) 2. Number of unmatched fragments in either prefix is bounded by r 9/6/2021 ©Bud Mishra, 2001 30

Comparing Maps: Effect of Partial Digestion • Parameters: – Partial digestion probability, p – Relative sizing error, b – # Restriction fragments, n – Overlap threshold ratio, q • m = n p = Expected # detected restriction fragments. • Controlling False Negative: K 5 np 4 q/2 and r = k 1/p 4, k 1 ¼ 2 If in fact the clones A and B overlap then we will it detect with a probability, at least (1 -exp(-k 1)) (1 – exp(-n p 4 q/8)) 9/6/2021 ©Bud Mishra, 2001 31

Overlap Rule • Controlling False Positive: Consider an arbitrary alignment: Let the random variable W denote the number of fragments in clone A that positionally match with the fragments of clone B. E[CW, i] = Cm, i (b/2)i ¼ (1/i!) (np b/2)i • By Brun’s sieve Pr[W = i] = (1/i!) (b n p/2)i exp(-b n p /2) Poisson » b n p /2 • and the false positive probability is 4 r å 1 i=k (1/i!) (b n p)I e-b n p/2 Make r as small & k as large as possible 9/6/2021 ©Bud Mishra, 2001 32

Experimental Design • Relation among the error parameters: 3 b n p /4 5 k 5 n p 4 q/2 ) p = (3 b/2 q)1/3 • Parameter choice for shotgun-mapping. Make the partial digestion probability rather high (close to 1) or the relative sizing error as low … for instance by using a rare cutter. 9/6/2021 ©Bud Mishra, 2001 33

Gentig (GENomic con. TIG) Algorithm • Scoring Function - An upper bound estimate of the false positive overlap probability - A Bayesian probability estimate for the proposed placement Maximize the Bayesian Probability Density subject to the False Positive Probability Constraint *GREEDY ALGORITHM* 9/6/2021 ©Bud Mishra, 2001 34

Concluding Remarks • The study of genetics relies on complete nucleotide sequences of the organism together with a description of the transcription units. • While this information at its finest level is not often available, or when available, suffers from various errors due to sequencing or assembly, one can garner much information from significantly coarser descriptions that are easily available in genomic maps. • Such maps with high resolution and accuracy as well as partially assembled sequences at various degrees of completion exist for many of the microbial organisms, yeasts, worms, flies and now humans. 9/6/2021 ©Bud Mishra, 2001 35

Concluding Remarks • In general, genetically or physically mapped collections of objects derived from the genome under study are still of immense utility, and require robust algorithmic tools to validate their mutual consistency and integration. • The integrated genomic databases derived from all available sources are likely to prove useful even at an early stage for – – 9/6/2021 annotation, gap detection (in sequences) and targeted gap closing, sequence contig phasing and map assisted sequence assembly. ©Bud Mishra, 2001 36
- Slides: 36