Computational Biology Lecture 2 Genome Organization Bud Mishra

Computational Biology Lecture #2: Genome Organization Bud Mishra Professor of Computer Science and Mathematics 9 ¦ 17 ¦ 2002 12/18/2021 ©Bud Mishra, 2001 1

Active Areas of Research(1) • Human Genome Project: (Completed? ) – Read 3 billion base pairs in 46 human chromosomes – Deemed “substantially completed on June 27, 2000. ” • Polymorphisms and Haplotyping – SNPs (Single Nucleotide Polymorphisms): Catalog the single base pair variations occurring about 1 in 800 base pairs of human genome over the entire populations – RFLP-Map: Restriction Fragment Length Polymorphisms 12/18/2021 ©Bud Mishra, 2001 2

Active Areas of Research(2) • Transcription Maps: – Identify all (about 30, 000 (? )) the genes in the human genome. – Particularly interesting are the ones involved in cancer…About 100 oncogenes and 1000 tumor suppressor genes • Linkage Analysis: – Relate genes (or polymorphic markers) to phenotypes (externally observable traits) by analyzing genomes of a family (kinship) or over a population. 12/18/2021 ©Bud Mishra, 2001 3

Active Areas of Research(3) • Functional Genomics: – Understand how an interactive network of genes affect a chain of metabolic pathways to ultimately determine the phenotypes • Comparative Genomics: – Relate genes within and across species to understand their evolutionary relationship…Phylogeny. 12/18/2021 ©Bud Mishra, 2001 4

Active Areas of Research(4) • Cell Informatics: – Interaction between proteins (membrane and soluble ones) to determine the dynamics of a cell. – Interaction among a heterogeneous population of cells. • Rational Drug Design: – Design of drugs and delivery systems to modify the dynamics of the cells. 12/18/2021 ©Bud Mishra, 2001 5

Introduction to Biology • Genome: – Hereditary information of an organism is encoded in its DNA and enclosed in a cell (unless it is a virus). All the information contained in the DNA of a single organism is its genome. • DNA molecule can be thought of as a very long sequence of nucleotides or bases: S = {A, T, C, G} 12/18/2021 ©Bud Mishra, 2001 6

Complementarity • • DNA is a double-stranded polymer and should be thought of as a pair of sequences over S. However, there is a relation of complementarity between the two sequences: – A , T, C , G – That is if there is an A (respectively, T, C, G) on one sequence at a particular position the other sequence must have a T (respectively, A, G, C) at the same position. We will measure the sequence length (or the DNA length) in terms of base pairs (bp): for instance, human (H. sapiens) DNA is 3. 3 £ 109 bp measuring about 6 ft of DNA polymer completely stretched out! 12/18/2021 ©Bud Mishra, 2001 7

The Central Dogma • • • The intermediate molecule carrying the information out of the nucleus of an eukaryotic cell is RNA, a single stranded polymer. RNA also controls the translation process in which amino acids are created making up the proteins. The central dogma(due to Francis Crick in 1958) states that these information flows are all unidirectional: “The central dogma states that once `information' has passed into protein it cannot get out again. The transfer of information from nucleic acid to nucleic acid, or from nucleic acid to protein, may be possible, but transfer from protein to protein, or from protein to nucleic acid is impossible. Information means here the precise determination of sequence, either of bases in the nucleic acid or of amino acid residues in the protein. ” 12/18/2021 ©Bud Mishra, 2001 8

Interrupted Genes: • An open reading frame (containing a gene) consists of – INTRONS: Intervening sequences a Noncoding regions – EXONS: Protein coding regions • Introns are abundant in eukaryotes and certain animal viruses. 12/18/2021 ©Bud Mishra, 2001 9
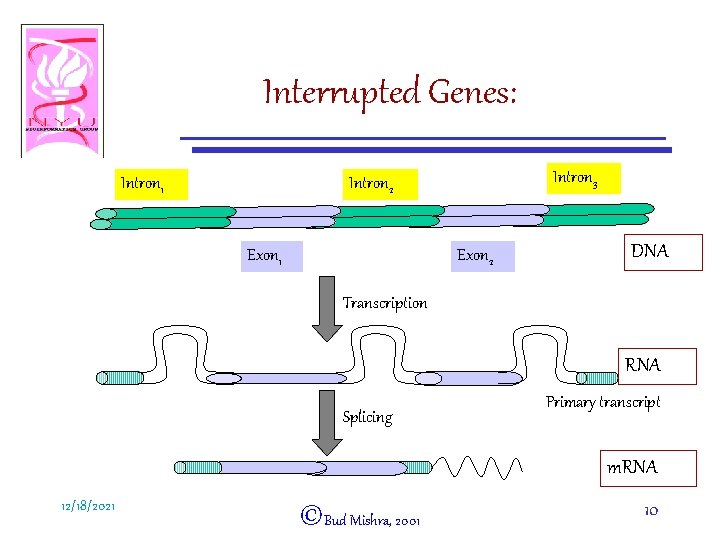
Interrupted Genes: Intron 1 Intron 3 Intron 2 Exon 1 Exon 2 DNA Transcription RNA Splicing Primary transcript m. RNA 12/18/2021 ©Bud Mishra, 2001 10
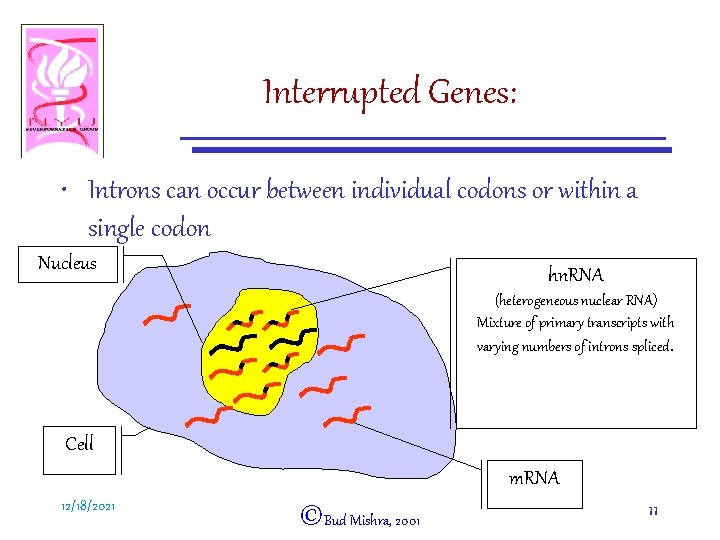
Interrupted Genes: • Introns can occur between individual codons or within a single codon Nucleus hn. RNA (heterogeneous nuclear RNA) Mixture of primary transcripts with varying numbers of introns spliced. Cell m. RNA 12/18/2021 ©Bud Mishra, 2001 11
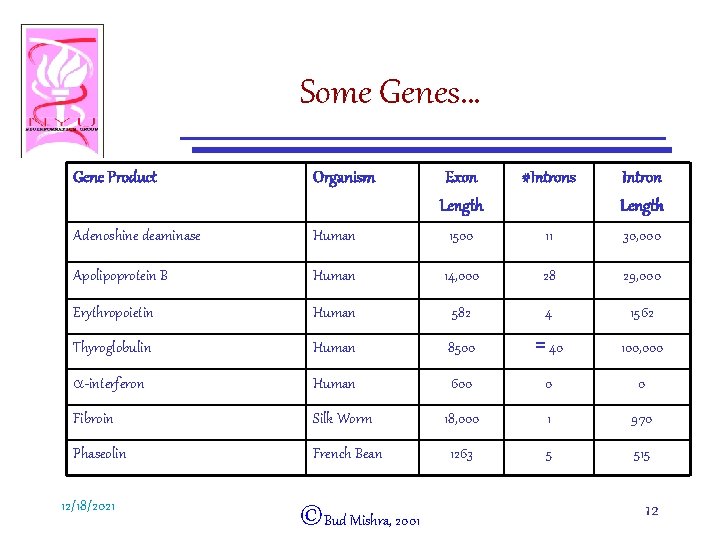
Some Genes… Gene Product Organism Exon Length #Introns Intron Length Adenoshine deaminase Human 1500 11 30, 000 Apolipoprotein B Human 14, 000 28 29, 000 Erythropoietin Human 582 4 1562 Thyroglobulin Human 8500 = 40 100, 000 a-interferon Human 600 0 0 Fibroin Silk Worm 18, 000 1 970 Phaseolin French Bean 1263 5 515 12/18/2021 ©Bud Mishra, 2001 12
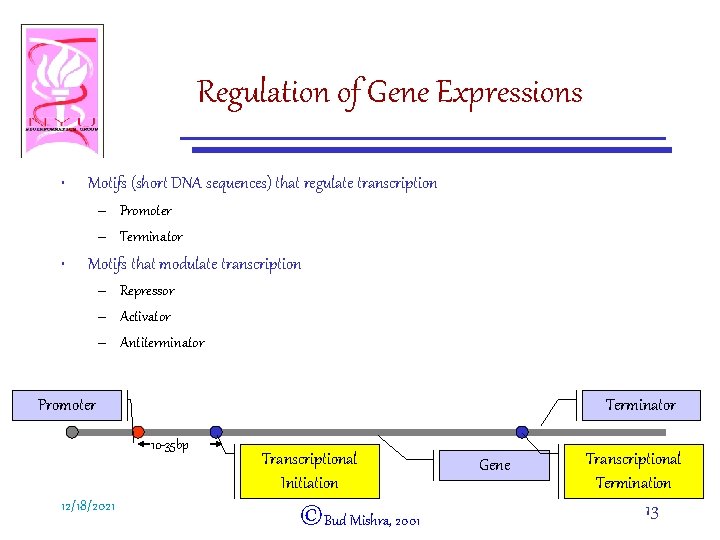
Regulation of Gene Expressions • Motifs (short DNA sequences) that regulate transcription – Promoter – Terminator • Motifs that modulate transcription – Repressor – Activator – Antiterminator Promoter Terminator 10 -35 bp 12/18/2021 Transcriptional Initiation ©Bud Mishra, 2001 Gene Transcriptional Termination 13

Promoters • pol I (RNA polymerase I) – Transcribes ribosomal RNA genes 100 » 1000 bp in front of the gene • pol II (RNA polymerase II) – Transcribes genes encoding polypeptides – Complex and variable regulatory regions • pol III (RNA polymerase III) – Transcribes transfer RNA and other small RNAs – Both up and down stream 12/18/2021 ©Bud Mishra, 2001 14

Motifs • Each motif is a binding site for a specific protein • Transcription Factor: – Transcription factors (specific to a cell/environmental conditions) bind to regulatory regions and facilitate • Assembly of RNA polymerase into a transcriptional complex • Activation of a transcriptional complex. • Termination Factor: • – Assembly of proteins for termination and modification of the end of the RNA Epigenetic Changes – Methylation of the cytosine in the 5’ region – Structural changes in cromatin 12/18/2021 ©Bud Mishra, 2001 15

Organization of Genetic Information • Bacterial Genome: – Genes are closely spaced along the DNA. – The sequences of genes may overlap. – Related genes (encoding enzymes whose functions are part of the same pathway or whose activities are related) are linked as a single transcription unit. 12/18/2021 ©Bud Mishra, 2001 16

Organization of Genetic Information • Eukaryotic Genome: – Genes are separated by long stretches of noncoding DNA sequences. – Multiple genes in a single transcription unit is extremely rare. – Multiple chromosomes – Linear – Chloroplasts and mitochondria – Circular – Genes appearing on the same chromosome are syntenic. 12/18/2021 ©Bud Mishra, 2001 17
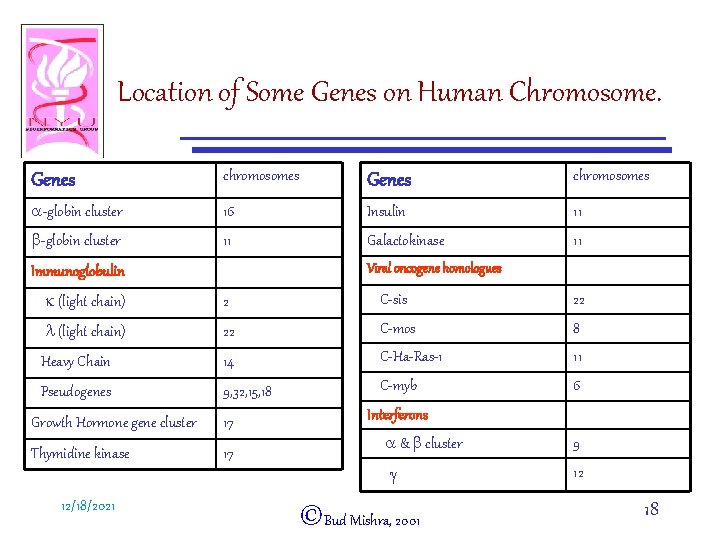
Location of Some Genes on Human Chromosome. Genes chromosomes a-globin cluster 16 Insulin 11 b-globin cluster 11 Galactokinase 11 Viral oncogene homologues Immunoglobulin k (light chain) 2 C-sis 22 l (light chain) 22 C-mos 8 Heavy Chain 14 C-Ha-Ras-1 11 Pseudogenes 9, 32, 15, 18 C-myb 6 Growth Hormone gene cluster 17 Thymidine kinase 17 12/18/2021 Interferons a & b cluster 9 g 12 ©Bud Mishra, 2001 18

Eukaryotic Genome • Multiple copies of the same gene – Solve “supply problem” – There are several hundred ribosomal RNA genes I mammals • Pseudogenes – Nonfunctional copies of genes…(Deletions or alterations in the DNA sequence) – Number of pseudo genes for a particular gene varies greatly…Different from one organism to another. 12/18/2021 ©Bud Mishra, 2001 19

Genes in Eukaryotes • A gene may appear exactly once • It may be part of a family of repeated sequence. Members of a family may be clustered or dispersed. • Members of a gene family may be related and functional (expressed at different times in development, or in different cells) or may be pseudo genes. • Chromosomal Morphology: – Nucleolar organizers (genes for ribosomal RNA) – Telomeric and Centromeric regions (Tandemly repeated sequences) 12/18/2021 ©Bud Mishra, 2001 20

The Rearrangement of DNA Sequences • Reshuffling of genes between homologous chromosomes via reciprocal crossing-over during both meiosis and mitosis. • Gene synteny and linkages are usually preserved. • Most rearrangements are random. • Some rearrangements are normal processes altering gene expressions in an orderly and programmed manner. 12/18/2021 ©Bud Mishra, 2001 21

Chromosomal Aberrations • • Breakage Translocation (Among non-homologous chromosomes. ) Formation of acentric and dicentric chromosomes. Gene Conversions Amplification and deletions Point mutations Jumping genes a Transposition of DNA segments Programmed rearrangements a E. g. , antibody responses. 12/18/2021 ©Bud Mishra, 2001 22

Repeat Structure • Copy Number: 2 » 106 • Direct Repeats “head-to-tail” – Tandem repeats or separated by other sequences • Inverted Repeats “head-to-head” – Stem-and-loop structure – Hairpin structure • Reverse Palindrome • True Palindrome 12/18/2021 ©Bud Mishra, 2001 23
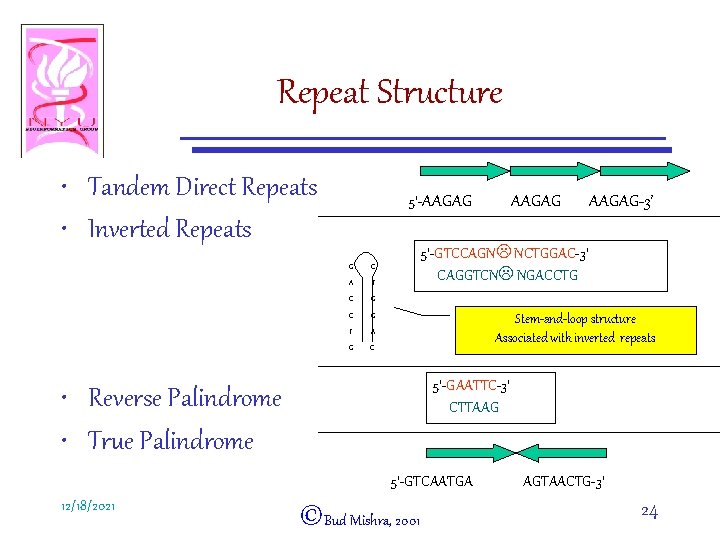
Repeat Structure • Tandem Direct Repeats • Inverted Repeats 5’-AAGAG G C A T C G T A G C AAGAG-3’ 5’-GTCCAGNL NCTGGAC-3’ CAGGTCNL NGACCTG Stem-and-loop structure Associated with inverted repeats 5’-GAATTC-3’ CTTAAG • Reverse Palindrome • True Palindrome 5’-GTCAATGA 12/18/2021 AAGAG ©Bud Mishra, 2001 AGTAACTG-3’ 24

Repeats within the Genome • Gene Family – Genes and its cognate pseudogenes • Satellite: Repeats made of noncoding units – Minisatellites: Tandem repeats…Mostly in centromeric regions – Satellite repeat units vary in length freom 2 base pairs to several thousands. 12/18/2021 ©Bud Mishra, 2001 25

Interspersed Repeats • SINES: Short Interspersed Repeats – Each repeat unit is of length 100 – 500 bps – Processed pseudogenes derived from class III genes – Example: Alu repeats…dimeric head-to-tail repeats of 130 bp • LINES: Long Interspersed Repeats – Each unit is of length > 6 Kb. 12/18/2021 ©Bud Mishra, 2001 26

A Genome Grammar • Consists of – A stochastic grammar specifying target DNA sequence together with – A description of polymorphisms and – A description of the sampling strategy for experiments • h specificationi ! h DNA-Seg i h Poly-Seg i* h Sample-Seg i+ 12/18/2021 ©Bud Mishra, 2001 27

Stochastic Grammar • h DNA-Seg i ! “. dna” h DNA-Spec i • h Poly-Seg i ! “. poly” h Weight i+ h Poly-Spec i • h Sample-Seg i ! “. sample” h Sample-spec i 12/18/2021 ©Bud Mishra, 2001 28

DNA Sequence • . dna A = 150 Ã sequence of length 150— Pr(A) = Pr(T) = Pr(C) = Pr(G) = ¼ B = A A m(. 30) Ã A followed by a mutated copy of A---Pr(Mutation) =. 30 C » 3 -7 p(. 2, . 3) Ã A string of length 3 to 7, Pr(A) =. 2, Pr(T) =. 3, Pr(C)=. 3, Pr(G) =. 2 ---C = Constant String D = C m(0. 03) n(10, 30) Ã m = mutation rate, n = copy number • S = 30, 000 B m(. 05, . 10) p(. 1, . 01) n(10) D !(500) 12/18/2021 ©Bud Mishra, 2001 29

Polymorphisms • Modify the ancestral sequence by a series of – S – D – X • Point mutation (SNPs) Deletions Translocations . poly. 8. 8 S 0. 00012 T D 1 -1. 00012 D 2 -2. 00006 D 3 -3. 00002 D 500 -1000. 00005 X 1000 -2000. 0005. poly. 4 S. 001 D 1 -2. 0005 12/18/2021 Two haplotypes of. 8 each and one haplotype of weight. 4 ©Bud Mishra, 2001 30

Sampling • • . sample 48, 000 Ã Number of Samples 400 600. 5 Ã Read Lengths. 01. 02 Ã Sequence Read Errors. 33 Ã Failure of Read. 3 1800 2200. 005 Ã Clone size. sample 12, 000 400 600. 5. 01. 03. 33. 4 9000 11000. 015 12/18/2021 ©Bud Mishra, 2001 31

Experiment • First sample generate 48, 000 end reads from inserts of average length 2 Kbp. – Sample proportions: 40% from haplotype H 1, 40% from H 2 and 20% from H 3 • Second sample generates 12, 000 end reads from inserts of average length 10 Kbp. – Sample proportions: 40% from haplotype H 1, 40% from H 2 and 20% from H 3 12/18/2021 ©Bud Mishra, 2001 32
- Slides: 32