Computational Biology Lecture 3 Mapping Bud Mishra Professor

Computational Biology Lecture #3: Mapping Bud Mishra Professor of Computer Science and Mathematics 9 ¦ 24 ¦ 2002 12/14/2021 ©Bud Mishra, 2001 1

Restriction Enzyme • Type II sequence specific restriction endonuclease – An enzyme that can “cut “a double stranded DNA by breaking the phosophodiester bonds at specific “target or restriction sites” on the DNA. • Retriction Sites: – Completely determined by their base pair decomposition – 4 » 8 long sequences of base pairs – Restriction Pattern 12/14/2021 ©Bud Mishra, 2001 2
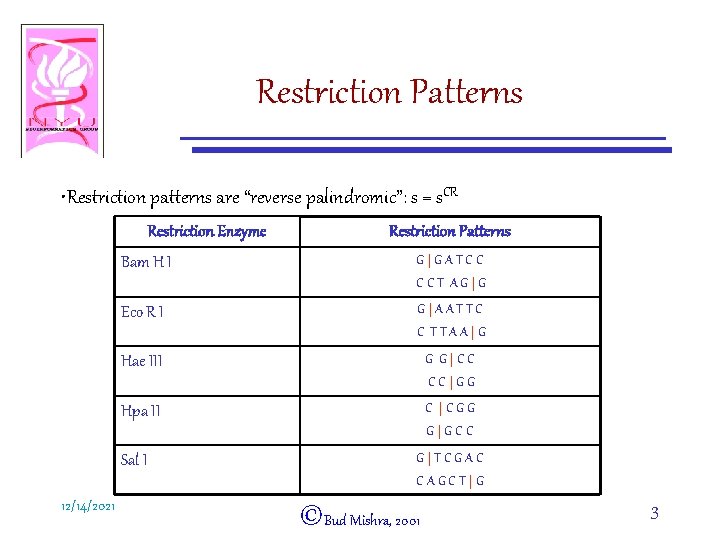
Restriction Patterns • Restriction patterns are “reverse palindromic”: s = s. CR Restriction Enzyme Bam H I G|GATCC CCT AG|G Eco R I G|AATTC C TTAA|G Hae III G G|CC CC|GG Hpa II C |CGG G|GCC Sal I 12/14/2021 Restriction Patterns G|TCGAC CAGCT|G ©Bud Mishra, 2001 3

Restriction Enzymes • Bacterial Immune Systems against Viral DNA – Bacteria use restriction enzymes by cleaving invading foreign DNA – Bacteria protect their own DNA against cleaving by a methylation process • Restriction Enzymes are very useful in biotechnology as – Biochemical Scissors – Biochemical Markers 12/14/2021 ©Bud Mishra, 2001 4

Applications of Restriction Enzyme • RFLP (Restriction Fragment Length Polymorphisms) – Polymorphisms ´ Sequence variation within a population • Restriction Maps – – Fingerprints Double Digestion Maps Multiple Complete Digestion Maps Ordered Restriction Maps • Clone Library (with Partial Digestion) • DNA Probes (Small Restriction Fragments) 12/14/2021 ©Bud Mishra, 2001 5

Digression Brun’s Sieve: Poison Approximation • Theorem: Let W be a nonnegative integer valued random variable such that E[CW, i] = li/i! Then Pr[W=M] ¼ e-l l. M/M! • Proof: Let the indicator variable IW=j be IW=j = { 1 if W=j {0 otherwise Show that IW=j = åk=01 CW, j+k Cj+k, k (-1)k 12/14/2021 ©Bud Mishra, 2001 6

Brun’s Sieve • IW=j = CW, j åk=0 W-j CW-j, k (-1)k = åk=0 W-j CW, j CW-j, k (-1)k = åk=01 CW, j+k Cj+k, k (-1)k • By Convention, CW, j = 0 if j > W. • Pr[W=m] = E[IW=m] = åk=01 E[CW, M+k CM+k, k (-1)k ] ¼ åk=01 l. M+k/(M+k)! CM+k, k (-1)k = l. M/M! åk=01 (-l)k/k! = e-l l. M/M! � 12/14/2021 ©Bud Mishra, 2001 7

Restriction Map: Resolution • G = Length of a genomic DNA • pk = Probability that an arbitrary site is a restriction site for a kcutter enzyme k = 4, 6 or 8 = Cutting Frequency • Uniform i. i. d. assumption: – “All base pairs occur at any given position with equal probability and independently: – pk = 1/(4 k) 12/14/2021 ©Bud Mishra, 2001 8

Numerical Values pk = 1/(4 k) l k = G pk 12/14/2021 Cut Numbers Cut Probability p 4 1/256 p 6 1/4, 096 p 8 1/65, 536 p 10 1/1, 048, 576 ©Bud Mishra, 2001 9

Statistics of Restriction Sites • Xj = Bernouli r. v. = Event that “there is a restriction site beginning at j. ” • W = åj=1 G Xj = Total # restriction sites in the genome = # successes in G independent trials • CW, i = # of “i-successful trials. ” • Consider the set of all “i-trials” – There are CG, i of these – Each “i-successful trial occurs with probability pki 12/14/2021 ©Bud Mishra, 2001 10
![Applying Brun’s Sieve • E[CW, i] = CG, i pki = (G(i)/i!)pki = (G Applying Brun’s Sieve • E[CW, i] = CG, i pki = (G(i)/i!)pki = (G](http://slidetodoc.com/presentation_image_h2/0962174007dd37bad617ca9dbce5e69e/image-11.jpg)
Applying Brun’s Sieve • E[CW, i] = CG, i pki = (G(i)/i!)pki = (G pk)i/i! = lki/i! • Pr[# Restriction Sites = M | G, lk] = Pr[W=M] ¼ e-lk (lk)M/M! � E[W] = lk = G pk = G/4 k s 2[W] = lk a S. D. [W] = G 1/2/2 k 12/14/2021 ©Bud Mishra, 2001 11
![Statistics of Restriction Fragments • Pr[A restriction Fragment is of length l] = (1 Statistics of Restriction Fragments • Pr[A restriction Fragment is of length l] = (1](http://slidetodoc.com/presentation_image_h2/0962174007dd37bad617ca9dbce5e69e/image-12.jpg)
Statistics of Restriction Fragments • Pr[A restriction Fragment is of length l] = (1 –pk)l pk ¼ e-l/mk/mk, where mk-1 = log (1/1 -pk) • W =r. v. with exponential distribution with mean mk : f. W(w) = e-w/mk/mk, w > 0 • Z = b W c =r. v. giving the length of a restriction fragment in base pairs E[W] = mk ¼ 1/pk= 4 k s 2[W] = mk 2 a S. D. [W] = mk ¼ 4 k 12/14/2021 ©Bud Mishra, 2001 12

Matching Rules for Restriction Fragments • Given two restriction fragments without any identifying markers, when can they be said to be the same? • We must account for small measurement errors: b = Relative Sizing Errors Matching Rule (II): Two restriction fragments are said to match if their Lengths x and y differ by less than b fraction (I. e. < 100 b % -b 5 1 – y/x 5 b 12/14/2021 ©Bud Mishra, 2001 13

False Positive Match Probability • Given: Two randomly chosen distinct restriction fragments obtained by cleaving a large genomic DNA by the same restriction enzyme, • What is the probability that the matching rule accidentally identify the fragments as the same? 12/14/2021 ©Bud Mishra, 2001 14

False Positive Probability • mk = Expected length of a restriction fragment x, y » Exponential(1/mk) f. X(x) = e-x/mk/mk • False Positive Probability = s 01 ( sx(1 -b)x(1+b) e-y/mk/mk dy ) e-x/mk/mk dx =s 01 ( sv(1 -b)v(1+b) e-udu) e-vdv =s 01 ( e-v(1 -b)-ev(1+b)) e-vdv =(1/2 -b) – (1/2+b) = 2 b/(4 -b 2) ¼ b/2 12/14/2021 ©Bud Mishra, 2001 15

Maps using Clones • Clone: – A large fragment of genomic DNA that has been preselected. – One can make faithful copies of a clone large number of times from a small number of initial clones. – All location information for a clone is assumed to be lost. For instance: it is not known: • Which chromosome a clone belongs to… • Whether two clones overlap… • What base-pair sequence the clone has… etc. 12/14/2021 ©Bud Mishra, 2001 16
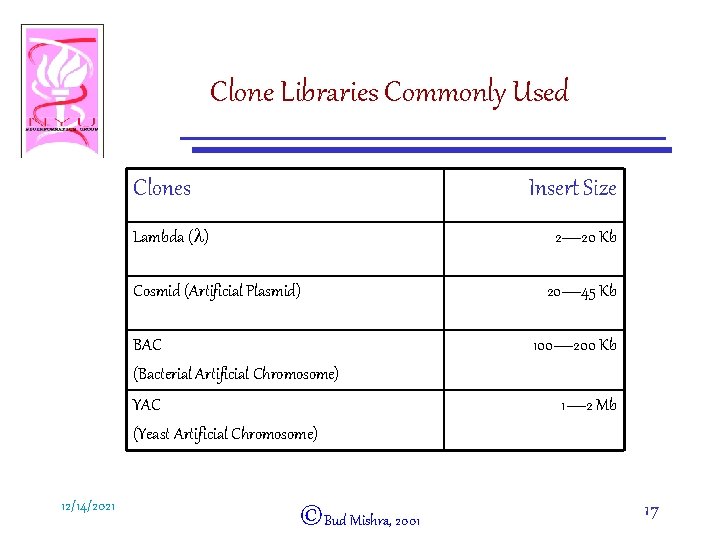
Clone Libraries Commonly Used Clones Insert Size Lambda (l) 2— 20 Kb Cosmid (Artificial Plasmid) BAC (Bacterial Artificial Chromosome) YAC (Yeast Artificial Chromosome) 12/14/2021 ©Bud Mishra, 2001 20— 45 Kb 100— 200 Kb 1— 2 Mb 17

Clone Library A preselected set of clones ´ Clone Library Locations of the clones are assumed to be uniformly random i. i. d. The size of a clone is roughly same. G = Genome length, L = Clone Length, N = # Clones in a library • Coverage = NL/G = c • • (The number of times the clones will cover the genome if the clones are concatenated end-to-end. Also, the expected number of clones covering any location of the genome. ) 12/14/2021 ©Bud Mishra, 2001 18

Example • A BAC library for human • G = 3, 300 Mb, L =180 Kb, N = 96, 000 c = NL/G = 96 £ 103 £ 180 £ 103/ (3. 3 £ 109) ¼ 6£ – 96, 000 randomly chosen BACs from the human genome provide a 6£ library. – Certain regions of the genome may be difficult to clone and hence may not be represented in the library. • A Tiling Path = A subset of clones that minimally cover the genome. – Removal of any clone from the tiling path will leave some location of the genome uncovered. – Every location of the genome is covered by no more than two clones. Every clone is overlapped by at most two other clones. – The coverage for a tiling path: 1 · c. TP · 2 12/14/2021 ©Bud Mishra, 2001 19
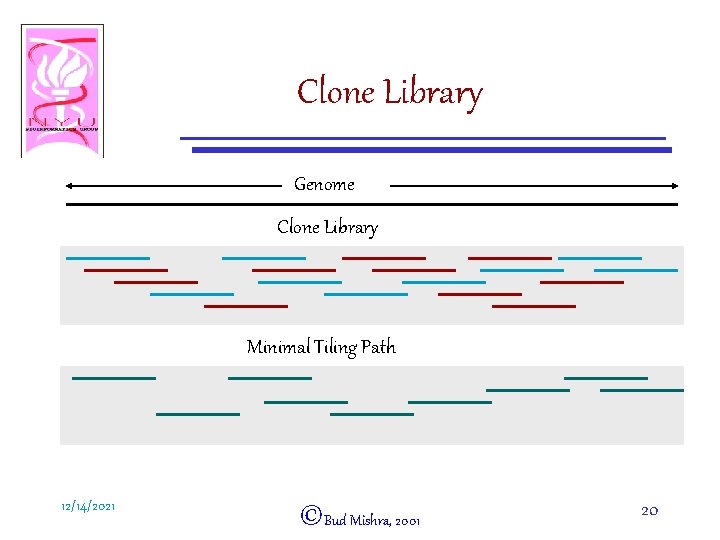
Clone Library Genome Clone Library Minimal Tiling Path 12/14/2021 ©Bud Mishra, 2001 20

Mapping A Single Clone • Decorate a clone with additional information—E. g. , – Restriction Pattern (Ordered Restriction Map, Finger Prints) – End Sequencing (500 base pairs on each end) – Probes (PCR products, Hybridization probes, etc. ) • Restriction Pattern: – Take a clone and completely digest it into small pieces (restriction fragments) by a restriction enzyme. – The restriction fragments and their order are always the same for that clone. 12/14/2021 ©Bud Mishra, 2001 21
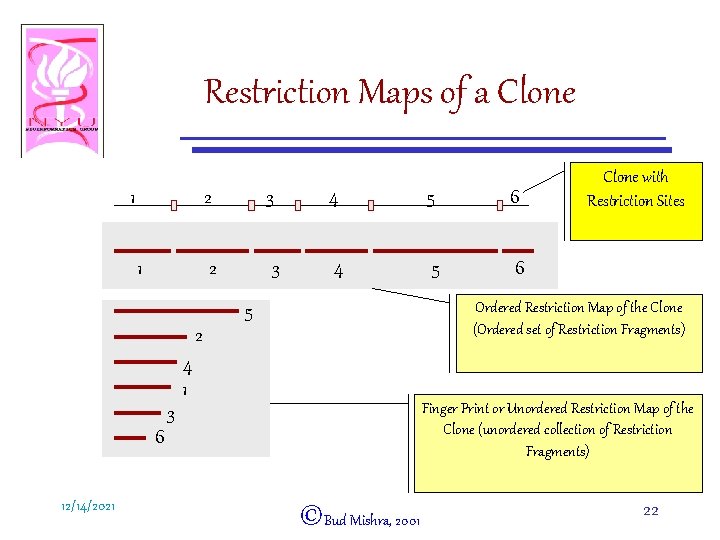
Restriction Maps of a Clone 1 1 2 3 4 5 6 Ordered Restriction Map of the Clone (Ordered set of Restriction Fragments) 5 2 4 1 Finger Print or Unordered Restriction Map of the Clone (unordered collection of Restriction Fragments) 3 6 12/14/2021 Clone with Restriction Sites ©Bud Mishra, 2001 22

A Clone Map • Key Question: • Given two clones, when can we say whether they overlap by simply examining their fingerprints or maps? • Issues: – – 12/14/2021 False positive and false negative in overlap detection Ordering all the clones using the overlap prpoerties Computing the tiling path Subcloning and sequencing (Divide-and Conquer) ©Bud Mishra, 2001 23
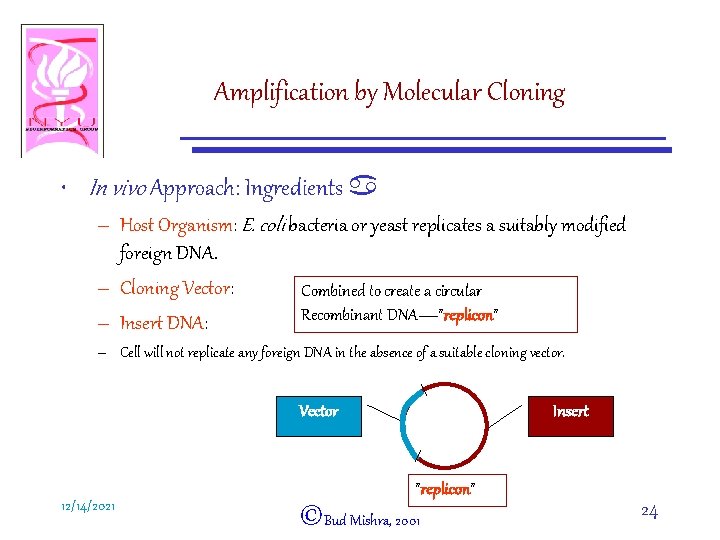
Amplification by Molecular Cloning • In vivo Approach: Ingredients a – Host Organism: E. coli bacteria or yeast replicates a suitably modified foreign DNA. – Cloning Vector: Combined to create a circular Recombinant DNA—”replicon” – Insert DNA: – Cell will not replicate any foreign DNA in the absence of a suitable cloning vector. Vector 12/14/2021 Insert ”replicon” ©Bud Mishra, 2001 24

Cloning • – Inserts and vectors with same “sticky ends” are mixed together with ligase enzyme. – This produces a circular replicon. • – Transformed host cells are transferred to culture dishes containing a solid growth medium – Cells divide making a colony containing 230 ¼ 109 inserts in 10 hours. Step 1: Step 2: – Transform the host cell by exposing a population of hosts to the ligase mixture containing the replicon – The replicons are inserted into the host cell 12/14/2021 • Step 3: – Identify the colonies of clones containing the copies of the inserts – Pick these colonies – Isolate and linearize the replicons. ©Bud Mishra, 2001 25

Sequencing A Genome • A “divide-and-conquer” approach: – Step 1: Divide…Create a “high coverage” clone library by choosing many randomly located clones (E. g. , 96, 000 BAC clones- each of length 180 Kb – from a human genome of length 3, 300 Mb. 6£ coverage BAC library. ) – Step 2: Contig…Use the clone overlap information to create the contigs (E. g. , 6 £ coverage BAC library would yield 96, 000 £ e-6 ¼ 200 contigs —About 10 contigs per chromosome each of size aout 10 Mb) 12/14/2021 ©Bud Mishra, 2001 26

Sequencing A Genome – Step 3: Prune…Remove “non-essential” clones from the contigs to form a “minimal tiling path. ” (E. g. , Minimal tiling path would consist of » 32, 000 BAC clones. ) – Step 4: Shotgun Sequencing…Subclone a BAC on the minimal tiling path into M 13’s. Generate sequence reads from M 13 subclones. Sequence reads = 300 » 1, 000 bps, 95% accuracy. – Step 5: Contig the sequence reads… – Step 6: Assemble the sequences and close the gaps… 12/14/2021 ©Bud Mishra, 2001 27

Finishing Phase • Filling the gaps between the contigs: – Synthesize a primer from the end of the contig sequence – Generate a new read from the M 13 subclone that starts with the synthesized primer. – If there is no such M 13 subclone— • Synthesize a pair of primers from the sequence at the ends of a “gap” • Amplify the DNA across the gap by performing PCR on the clone DNA • Sequence the PCR product. 12/14/2021 ©Bud Mishra, 2001 28

Sequence Assembly • Idealized Assembly: – Assuming no error in the read sequence. • Shortest Common Superstring Problem: – Given: A set {si}, where si is a string over some alphabet. – Find: The shortest string S which contains each si as a contiguous substring. • (SCSP – Shortest Common Superstring Problem – is NP-complete) 12/14/2021 ©Bud Mishra, 2001 29

Greedy Algorithm for Sequence Assembly • Find overlaps between pairs of sequence reads – – (Only consider overlaps that span at least 15 bps. ) • Sort overlaps by decreasing length • Merge read contigs according to the sorted list. 12/14/2021 ©Bud Mishra, 2001 30
- Slides: 30