Applying Genomic Profiling to Precision Cancer Medicine in











- Slides: 14

Applying Genomic Profiling to Precision Cancer Medicine in Clinical Practice George D. Demetri MD Senior Vice-President for Experimental Therapeutics Dana-Farber Cancer Institute Professor of Medicine, Harvard Medical School Co-Director, Ludwig Center at Harvard Medical School Boston, Massachusetts USA george_demetri@dfci. harvard. edu



Challenges to Clinical Application of New Diagnostic Genomic Technologies in Cancer • Variations in testing • Lack of standards for conduct or interpretation of tumor testing • Lack of data (and overly enthusiastic hype) for value of genomic profiling of tumors • Lack of patient/payor/physician demand

How to document and define the value of genomic profiling in cancer? • What sort of profiling? – Single gene testing – “Limited” panel testing vs. “large” panel testing – Whole exome? Whole genome? • What defines “actionability”? – Results may be interesting – but do they drive important changes in patient management?

Proving the value of genomic profiling in cancers • What drives demand for genomic profiling? • What determines the value of the most accurate diagnosis? – An available therapy? A change in prognosis? • How can we make accurate, large-scale profiling a reliable, inexpensive and necessary commodity?

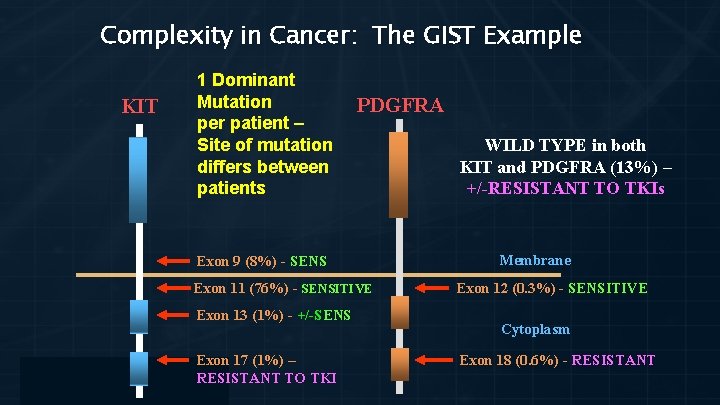
Complexity in Cancer: The GIST Example KIT 1 Dominant Mutation per patient – Site of mutation differs between patients PDGFRA Exon 9 (8%) - SENS Exon 11 (76%) - SENSITIVE Exon 13 (1%) - +/-SENS Exon 17 (1%) – RESISTANT TO TKI WILD TYPE in both KIT and PDGFRA (13%) – +/-RESISTANT TO TKIs Membrane Exon 12 (0. 3%) - SENSITIVE Cytoplasm Exon 18 (0. 6%) - RESISTANT

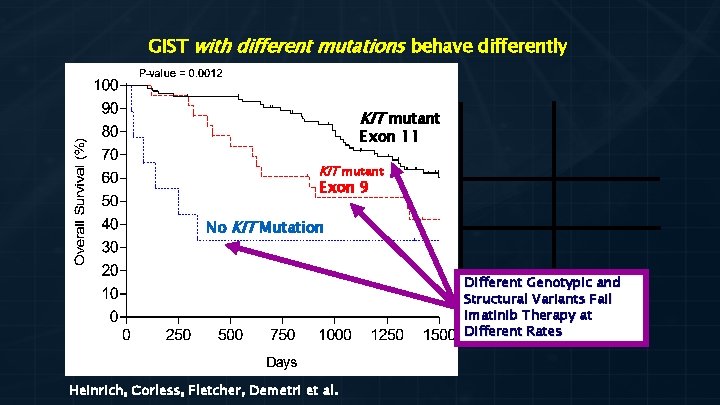
GIST with different mutations behave differently KIT mutant Exon 11 KIT mutant Exon 9 No KIT Mutation Different Genotypic and Structural Variants Fail Imatinib Therapy at Different Rates Heinrich, Corless, Fletcher, Demetri et al.

Patients Identify with Precision Medicine

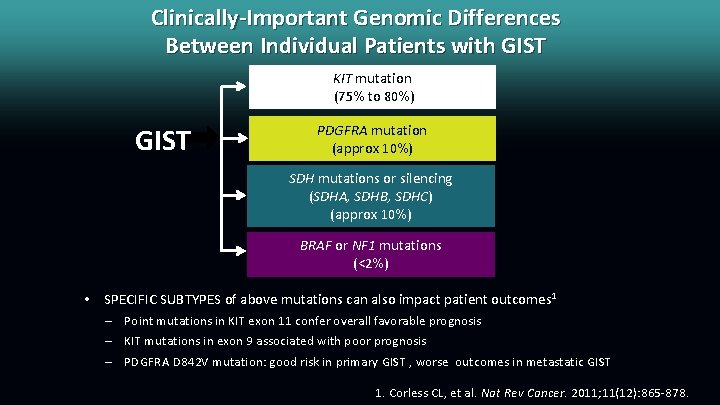
Clinically-Important Genomic Differences Between Individual Patients with GIST KIT mutation (75% to 80%) GIST PDGFRA mutation (approx 10%) SDH mutations or silencing (SDHA, SDHB, SDHC) (approx 10%) BRAF or NF 1 mutations (<2%) • SPECIFIC SUBTYPES of above mutations can also impact patient outcomes 1 – Point mutations in KIT exon 11 confer overall favorable prognosis – KIT mutations in exon 9 associated with poor prognosis – PDGFRA D 842 V mutation: good risk in primary GIST , worse outcomes in metastatic GIST 1. Corless CL, et al. Nat Rev Cancer. 2011; 11(12): 865 -878.

A Biblical Analogy • What is the minimum value for molecular profiling to become a necessary standard? • The story of Abraham negotiating with G_d about what level of ”righteousness” would save Sodom. – 50 ”righteous” people out of 100, 000? 25? 10? • 0. 0001% would be acceptable. • For 1. 7 M cancer diagnoses per year in US, need to change management in only 170 patient cases

Other Barriers to Optimal Use of Genomic Profiling • Education of physicians • Lack of (educated/rational) demand from patients • Indiscriminate interpretation of results – Not all BRAF mutations have the same therapeutic implications • Fear of legal action to obtain expensive drugs without adequate clinical justification

Summary on Status of Genomic Profiling to Enhance Meaningful Precision in Cancer Medicine • Feasibility of expansion for the best profiling technologies • Need large-scale access and rigorous analysis of impact • Fairness principles should apply (not just who is willing and able to pay for profiling) • Objective analytics on impact in clinical outcomes – Define important changes in patient care and outcomes
