1 CE 530 Molecular Simulation Lecture 4 ObjectOriented

1 CE 530 Molecular Simulation Lecture 4 Object-Oriented Programming and Etomica David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng. buffalo. edu

2 Object-Oriented Programming ¡ Programming accomplished through the actions and interactions of objects • everything is an object ¡ Forces abstract thinking about the structure and activities of a program ¡ Promotes re-use of code and extension to new applications ¡ Good design is difficult to develop • requires thorough understanding of application • conversely, its use facilitates a better understanding of application presents a good vehicle for teaching ¡ It’s fun!

What is an Object? ¡ A fancy variable • stores data • can perform operations using the data ¡ Every object has a type, or “class” • analogous to real, integer, etc. Fortran: real x, y, z Java: Atom a 1, a 2; • you define types (classes) as needed to solve your problems public class Atom { double mass; Vector r, p; } • types differ in the data they hold and the actions they can perform on it • every object is an “instance of a class” a 1 = new Atom(); 3

4 Makeup of an Object ¡ Fields (data) • primitive types (integer, float, double, boolean, etc. ) • handles to other objects complex objects are composed from simpler objects (composition) ¡ Methods (actions) • “subroutines and functions” • may take arguments and return values • have complete access to all fields of object ¡ A class has an interface • what the object presents to enable its manipulation • implementation (how it accomplishes its operations) can be hidden • object is viewed in terms of its “actions” and not its “thoughts” ¡ Inheritance • can define subclasses which inherit features of parent class • same interface, but different implementations • subclasses can be used anywhere parent class is expected • mechanism to change behavior of simulation
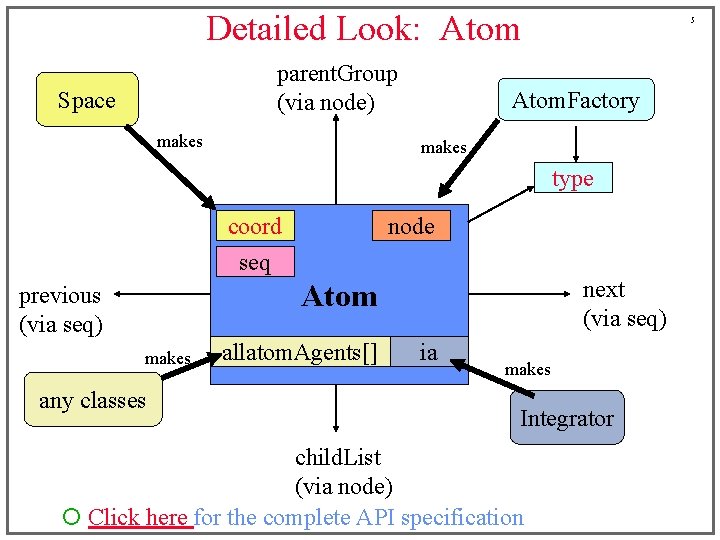
Detailed Look: Atom parent. Group (via node) Space makes 5 Atom. Factory makes type coord seq node next (via seq) Atom previous (via seq) makes any classes allatom. Agents[] ia makes Integrator child. List (via node) ¡ Click here for the complete API specification

6 Etomica ¡ Application Programming Interface (API) • Library of components used to assemble a simulation • Can be used independent of development environment Invoked in code programmed using Emacs (for example) ¡ GUI-based development environment • Simulation is constructed by piecing together elements • No programming required • Result can be exported to run stand-alone as applet or application ¡ Written in Java • Widely used and platform independent • Features of a modern programming language • Object-oriented ¡ Vehicle for presentation of molecular simulation methods
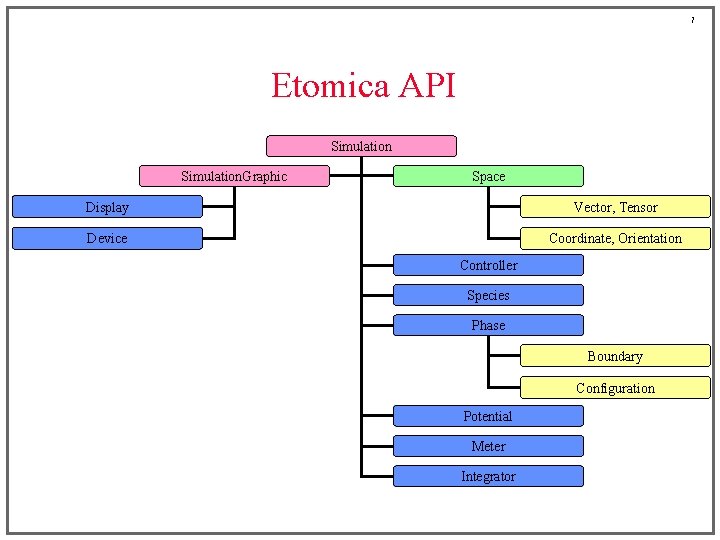
7 Etomica API Simulation. Graphic Space Display Vector, Tensor Device Coordinate, Orientation Controller Species Phase Boundary Configuration Potential Meter Integrator

8 Simulation Elements: Simulation Space ¡ Simulation. Graphic collects all the other elements of the simulation and ensures that they are connected for proper interaction Display Vector, Tensor ¡ Device There is only one instance of a Simulation in any. Coordinate, Orientation Controller applet/application Species • Applet: runs in a web page Phase • Application: runs as a program, from command line ¡ Simulation. Graphic Boundary Configuration • Subclass of Simulation used when preparing a simulation with a Potential graphical user interface (GUI) Meter Integrator

9 Simulation Elements: Space ¡ Space defines properties of the physical space in which the simulation is performed Simulation • Dimensionality (1 D, 2 D, 3 D, etc. ); continuum Simulation. Graphic Spacevs. lattice • Display Construction of vectors, tensors Vector, Tensor • Device Makes Coordinate for placement in each Atom Coordinate, Orientation position and momentum vector Controller • Coordinate. Pair defines how distances are Species computed • Constructs various types of Boundary for placement in each Phase ¡ Concrete classes • Space 1 D • Space 2 D • Space 3 D Boundary Configuration Potential Meter Integrator
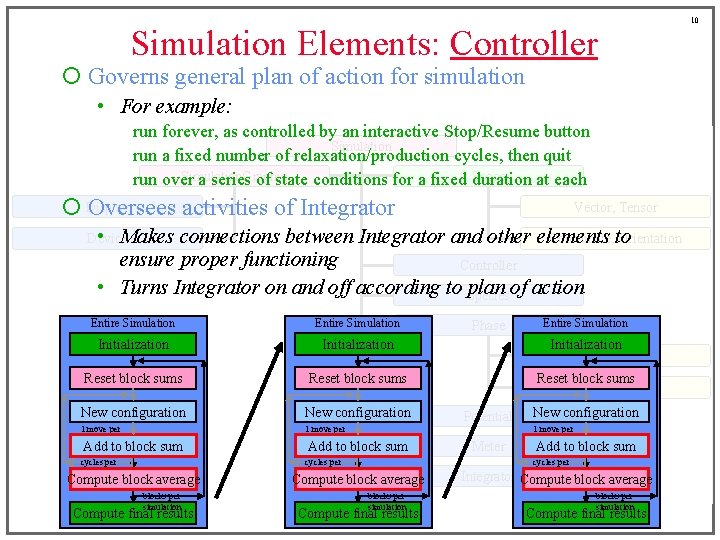
Simulation Elements: Controller ¡ Governs general plan of action for simulation • For example: run forever, as controlled by an interactive Stop/Resume button Simulation run a fixed number of relaxation/production cycles, then quit Simulation. Graphic Space run over a series of state conditions for a fixed duration at each ¡ Display Oversees activities of Integrator Vector, Tensor Device • Makes Coordinate, to Orientation connections between Integrator and other elements ensure proper functioning Controller • Turns Integrator on and off according to Species plan of action Entire Simulation Initialization Boundary Reset block sums Configuration New configuration 1 move per cycles per block Add to block sum Compute block average blocks per simulation Compute final results Phase Potential Meter Entire Simulation New configuration 1 move per cycle Add to block sum cycles per block Integrator Compute block average blocks per simulation Compute final results 10

11 Simulation Elements: Integrator ¡ Adds the physics needed Simulation to generate configurations properly • various. Simulation. Graphic integration schemes introduced by. Space developing a new integrator Display Vector, Tensor • Molecular dynamics integrators Device hard-potential dynamics various kinds of soft-potential integrator • Monte Carlo integrators more later Coordinate, Orientation Controller Species Phase ¡ Places an Agent in each atom to assist integration • for example, hard potential places Agent recording collision time collision partner Boundary Configuration Potential Meter ¡ Fires Integrator. Interval. Event to notify Integrator listeners that simulation has proceeded one step

12 Simulation Elements: Integrator Simulation ¡ run method for top-level Integrator class Simulation. Graphic Space public void run() { Display Vector, Tensor step. Count = 0; int iie. Count = interval+1; Device Coordinate, Orientation while(step. Count < max. Steps) { while(pause. Requested) do. Wait(); Controller if(reset. Requested) {do. Reset(); reset. Requested = false; } if(halt. Requested) break; Species do. Step(); //abstract method in Integrator. subclasses implement algorithms (MD/MC) if(--iie. Count == 0) { //count down to determine when a cycle is completed Phase fire. Interval. Event(interval. Event ); //notify listeners of completion of cycle iie. Count = interval; Boundary } if(do. Sleep) { //slow down simulation so display can keep up Configuration try { Thread. sleep(sleep. Period ); } catch (Interrupted. Exception e) { } Potential } step. Count++; } //end of while loop Meter fire. Interval. Event(new Interval. Event(this, Interval. Event. DONE)); } //end of run method Integrator

13 Simulation Elements: Phase ¡ Collects molecules that interact Simulation • holds root of hierarchy of atoms/molecules. Space Simulation. Graphic • handles addition/removal of molecules Display • multiple phases possible in one simulation Device • sets up base lists of all atoms, molecules, Controller species Vector, Tensor Coordinate, Orientation defines which atoms are up or down from a given atom Species ¡ Holds a Boundary object (from Space) that Phase defines the boundary conditions used in the phase Boundary ¡ Houses Configuration object that creates or saves a. Configuration configuration of molecules Potential Meter Integrator
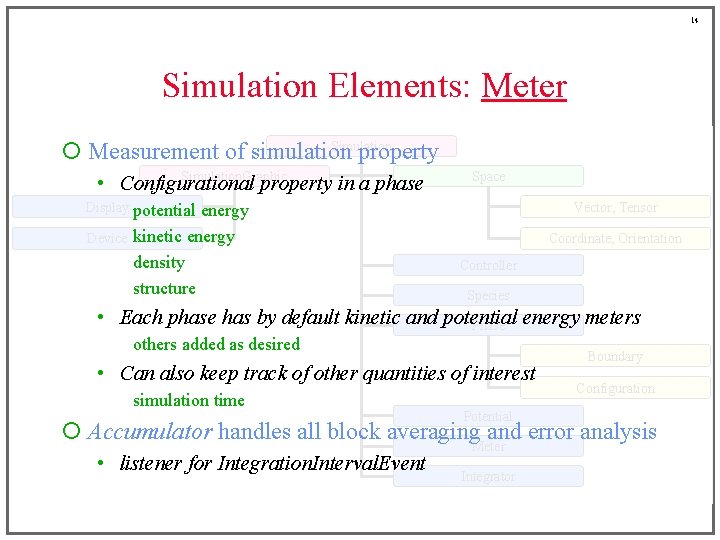
14 Simulation Elements: Meter Simulation ¡ Measurement of simulation property Simulation. Graphic • Configurational property in a phase Space Display potential Device energy kinetic energy density structure Vector, Tensor Coordinate, Orientation Controller Species • Each phase has by default kinetic and potential Phase energy meters others added as desired • Can also keep track of other quantities of interest simulation time Potential Boundary Configuration ¡ Accumulator handles all block averaging. Meter and error analysis • listener for Integration. Interval. Event Integrator

15 Simulation Elements: Species Simulation ¡ Holds Atom. Factory type of Simulation. Graphicthat defines how a particular Space molecule is constructed Display Vector, Tensor • how many atoms of which type • nominal arrangement of atoms in molecule Controller Device Coordinate, Orientation ¡ Places Species. Agent in each phase to manage molecules Species there Phase • root of atom hierarchy for species in phase • looping over molecules • addition and removal of atoms Potential Boundary Configuration ¡ Basis for defining interaction potential between molecules Meter • but does not by itself describe the pair potential Integrator
16 Simulation Elements: Potential ¡ Defines interaction between atoms (and thereby molecules) Simulation • Hard potential Simulation. Graphic Display Device Space collision time collision dynamics • Soft potential force, virial • All potentials energy Vector, Tensor Coordinate, Orientation Controller Species Phase Boundary ¡ Intermolecular potentials collect interatomic potential • Potential 1: intramolecular • Potential 2: intermolecular ¡ Hierarchical arrangement • More later Configuration Potential Meter Integrator
17 Simulation Elements: Display Simulation. Graphic from the simulation Space ¡ Presents information Display • display of configuration for animation Device • display of data table graph ¡ Connects to Meter to get information ¡ Listener of Integration. Interval. Event Vector, Tensor Coordinate, Orientation Controller Species Phase Boundary Configuration Potential Meter Integrator
18 Simulation Elements: Device Simulation ¡ Interactive. Simulation. Graphic manipulation of simulation Space Display • sliders, buttons, combo. Boxes etc. for setting properties. Vector, and. Tensor affecting course of simulation Device Coordinate, Orientation Controller Species Phase Boundary Configuration Potential Meter Integrator

Putting It Together import etomica. *; import etomica. graphics. *; public class My. Simulation extends Simulation. Graphic { Save this code in a file named My. Simulation. . java //Constructor public My. Simulation() { //Instantiate classes super(new Space 2 D()); Controller controller = new Controller (this); Integrator integrator = new Integrator. Hard(this); Potential potential = new P 2 Hard. Sphere(); Species species = new Species. Spheres. Mono(this); Phase phase = new Phase(this); Display display = new Display. Phase(this); Device device = new Device. Trio. Controller. Button(this); //Tie elements together this. element. Coordinator. go(); } //End of constructor public static void main(String[] args) { My. Simulation simulation = new My. Simulation(); simulation. make. And. Display. Frame(); } //End of main } //End of My. Simulation class 19

20 Running It: Application ¡ If needed, obtain the java runtime for your platform from Sun ¡ Get the etomica class library archive, etomica. jar. Place it in the same directory as your source file ¡ Compile your source • javac –classpath etomica. jar My. Simulation. java creates My. Simulation. class ¡ Run as an application • java -cp etomica. jar; . My. Simulation

21 Running It: Applet ¡ Add an Applet inner class to your source } //End of main public static class Applet extends javax. swing. JApplet { public void init() { get. Content. Pane(). add(new My. Simulation(). panel()); } }//end of Applet } //End of My. Simulation class ¡ Compile to create My. Simulation. class, as for application ¡ Create an html file, named (for example) Applet. html <HTML> <HEAD> <TITLE>Applet HTML</TITLE> </HEAD> <BODY bgcolor="#ffffff"> <APPLET CODE=“My. Simulation$Applet. class " archive=“etomica. jar" WIDTH=640 HEIGHT=350></APPLET> </BODY> </HTML> ¡ Run applet, either • • • Using appletviewer at command line: appletviewer Applet. html Or using your web browser, pointed at Applet. html on your machine Or deploy to a web server for the rest of the world to enjoy!
- Slides: 21