Using Traveling Salesman Problem Algorithms to Determine Multiple

Using Traveling Salesman Problem Algorithms to Determine Multiple Sequence Alignment Orders Weiwei Zhong

Topics • Background • Algorithm Design • Test Results

Background Definitions

What is a Sequence Alignment? Given • 2 or more sequences • a scoring scheme • match score • mismatch score • gap penalty Insert gaps in each sequence, so that • all sequences have the same length • maximum pairing score

Scoring Matrix Simplified Scoring • match = 2 • mismatch = -1 • gap penalty = -2 In Practice Scoring matrix

Global vs. Local Alignments Global: entire lengths of sequences F G K – G K G F G K G Local: regions of sequences - - - F G K G - -

Pairwise Alignment vs. Multiple Sequence Alignment (MSA) Pairwise: 2 sequences F G K G MSA: more than 2 sequences F F - G G G - K K F Q F G G K K G G

Background Basic Dynamic Programming
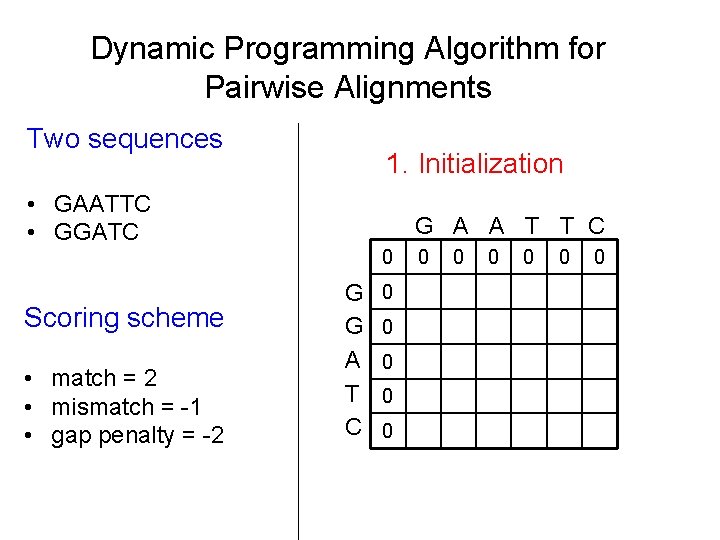
Dynamic Programming Algorithm for Pairwise Alignments Two sequences 1. Initialization • GAATTC • GGATC G A A T T C 0 Scoring scheme • match = 2 • mismatch = -1 • gap penalty = -2 G G A T C 0 0 0
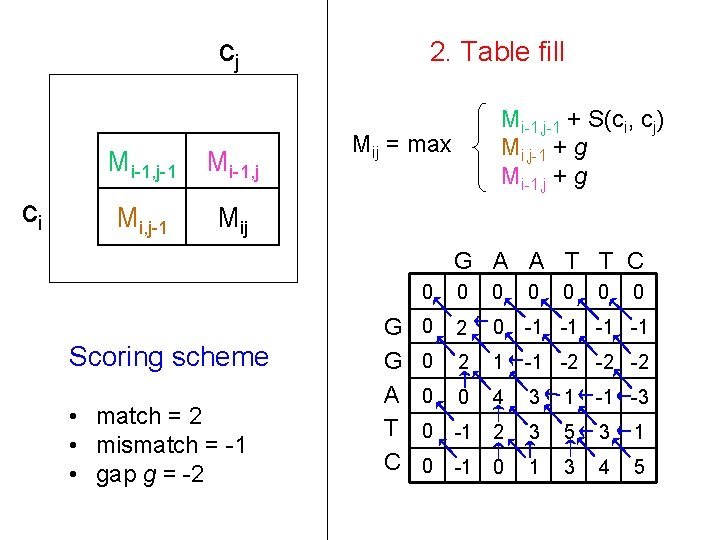
cj ci Mi-1, j-1 Mi-1, j Mi, j-1 Mij 2. Table fill Mi-1, j-1 + S(ci, cj) Mi, j-1 + g Mi-1, j + g Mij = max G A A T T C Scoring scheme • match = 2 • mismatch = -1 • gap g = -2 G G A T C 0 0 0 0 2 0 -1 -1 0 2 1 -1 -2 -2 -2 0 0 4 3 1 -1 -3 0 -1 2 3 5 3 1 0 -1 0 1 3 4 5
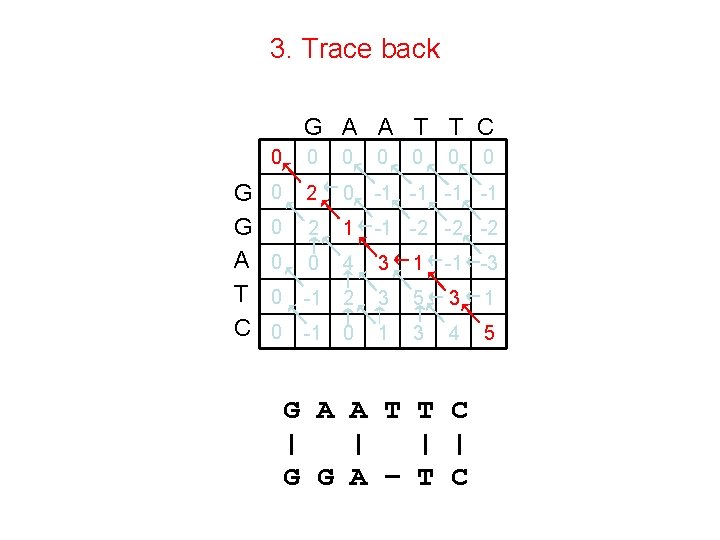
3. Trace back G A A T T C G G A T C 0 0 0 0 2 0 -1 -1 0 2 1 -1 -2 -2 -2 0 0 4 3 1 -1 -3 0 -1 2 3 5 3 1 0 -1 0 1 3 4 5 G A A T T C | | G G A – T C 0
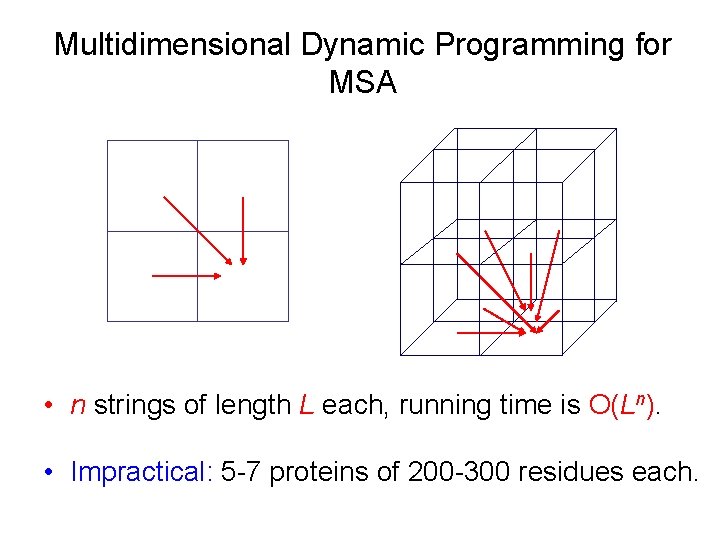
Multidimensional Dynamic Programming for MSA • n strings of length L each, running time is O(Ln). • Impractical: 5 -7 proteins of 200 -300 residues each.

Topics • Background • Algorithm Design • Test Results

Algorithm Design An MSA Heuristic
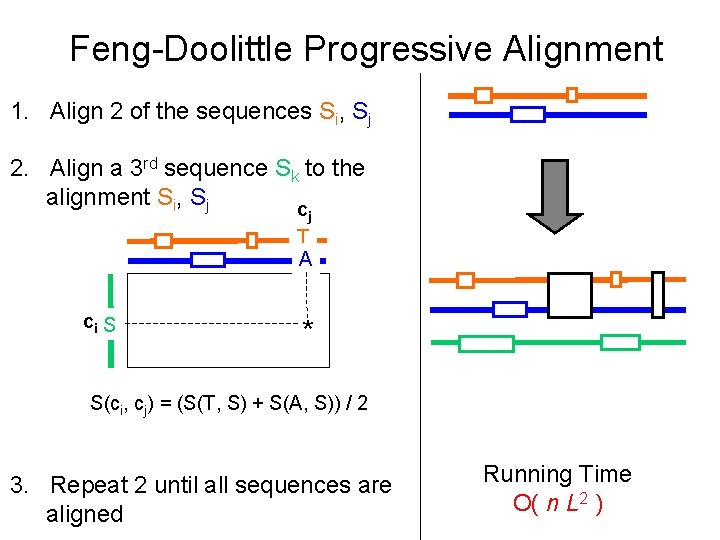
Feng-Doolittle Progressive Alignment 1. Align 2 of the sequences Si, Sj 2. Align a 3 rd sequence Sk to the alignment Si, Sj c j T A ci S * S(ci, cj) = (S(T, S) + S(A, S)) / 2 3. Repeat 2 until all sequences are aligned Running Time O( n L 2 )
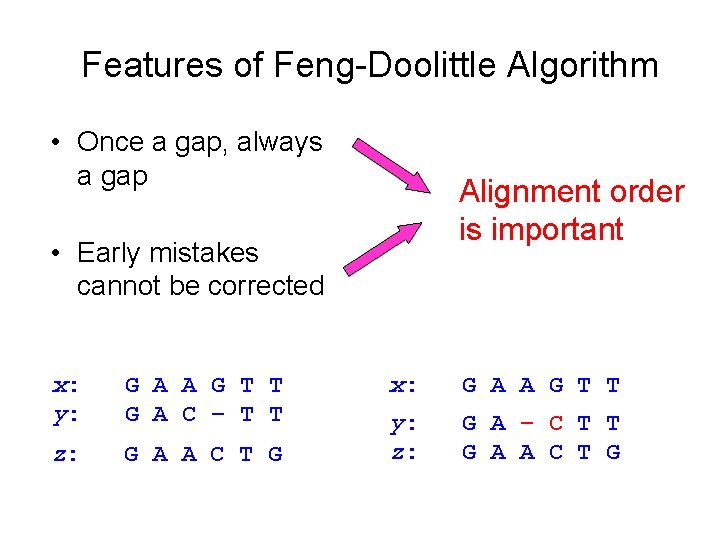
Features of Feng-Doolittle Algorithm • Once a gap, always a gap Alignment order is important • Early mistakes cannot be corrected x: y: G A A G T T G A C – T T z: G A A C T G x: G A A G T T y: z: G A – C T T G A A C T G

Algorithm Design Tsp. Msa: First Version
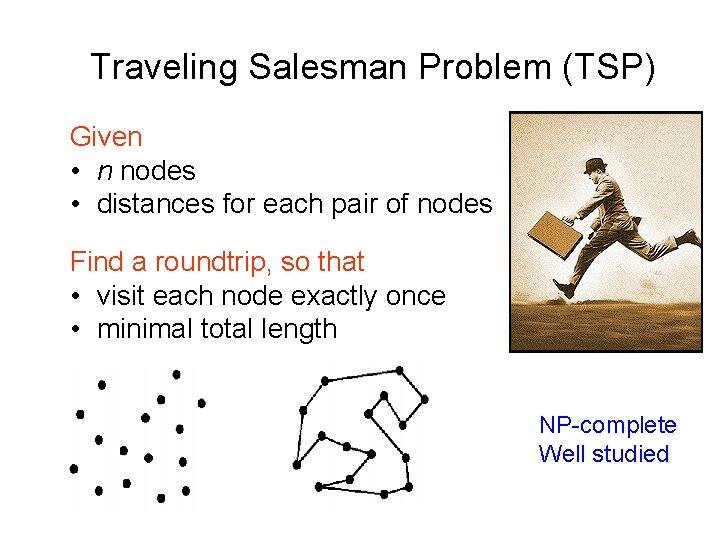
Traveling Salesman Problem (TSP) Given • n nodes • distances for each pair of nodes Find a roundtrip, so that • visit each node exactly once • minimal total length NP-complete Well studied
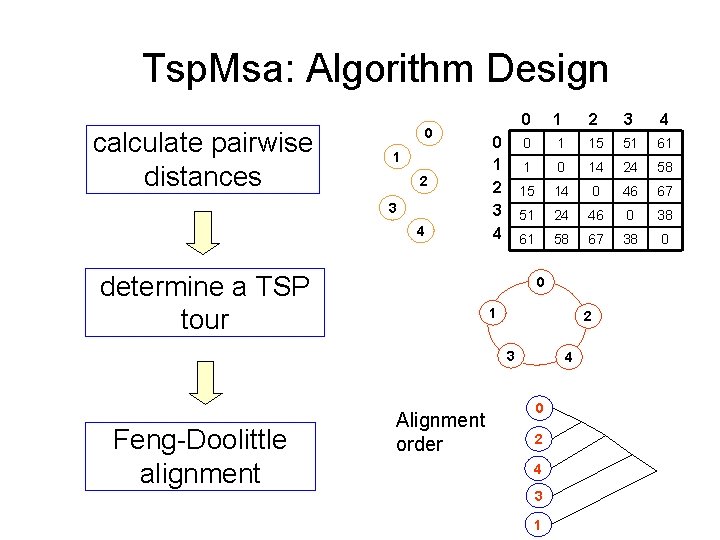
Tsp. Msa: Algorithm Design calculate pairwise distances 0 1 2 3 4 determine a TSP tour 0 1 2 3 4 0 1 15 51 61 1 0 14 24 58 15 14 0 46 67 51 24 46 0 38 61 58 67 38 0 0 1 2 3 Feng-Doolittle alignment Alignment order 4 0 2 4 3 1
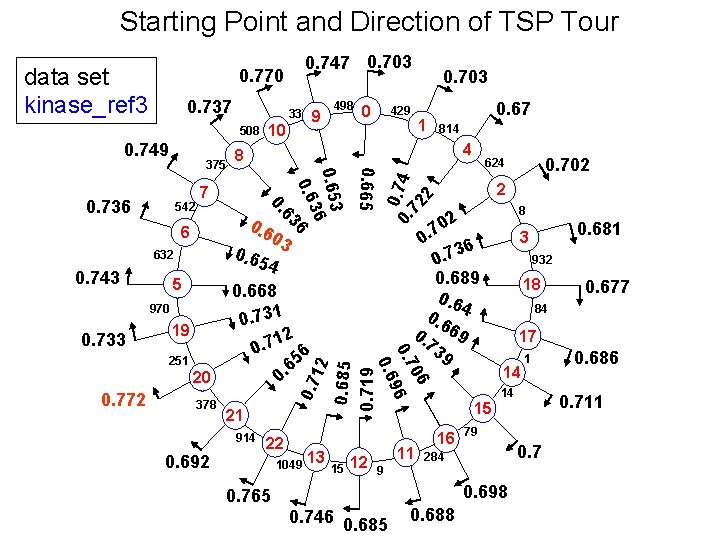
Starting Point and Direction of TSP Tour 0. 737 508 7 0. 6 03 0. 65 4 5 970 19 251 20 0. 772 378 21 914 0. 692 1 0. 67 814 22 1049 13 15 12 02 7. 0 36 0. 7 0. 689 0. 6 4 0. 69 73 9 11 8 932 18 17 14 1 14 79 0. 7 284 0. 688 0. 686 0. 711 15 16 0. 677 84 0. 698 0. 746 0. 685 0. 681 3 9 0. 765 0. 702 624 2 06 0. 7 96 0. 733 0. 668 0. 731 12 0. 7 6 5 6 0. 12 632 0. 7 6 0. 743 0. 665 542 429 4 0. 736 0 8 0. 653 36 0. 6 6 63 375 498 0. 703 0. 0. 74 72 2 0. 749 10 337 9 0. 685 0. 719 data set kinase_ref 3 0. 747 0. 703 0. 770

Algorithm Design Tsp. Msa: Modified Design
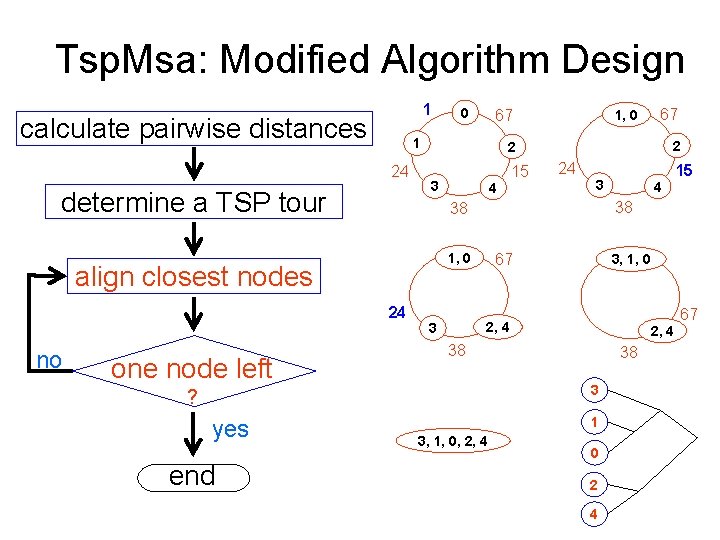
Tsp. Msa: Modified Algorithm Design 1 calculate pairwise distances determine a TSP tour 3 4 15 24 3 4 1, 0 67 3, 1, 0 2, 4 3 15 38 2, 4 38 38 3 ? end 67 2 38 24 yes 1, 0 2 align closest nodes one node left 67 1 24 no 0 1 3, 1, 0, 2, 4 0 2 4 67

Modified Algorithm is Better Alignment order for Kinase_ref 3 5 6 7 8 10 9 0 1 4 2 3 18 17 14 15 16 11 12 13 22 21 20 19 Original Tsp. Msa : 0. 603 (worst) - 0. 772 (best) Modified Tsp. Msa : 0. 836
Topics • Background • Algorithm Design • Test Results

Test Results What to Compare With?
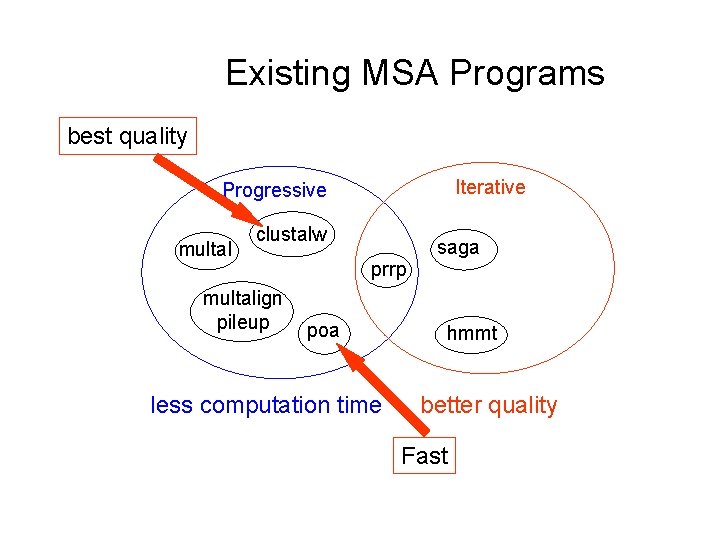
Existing MSA Programs best quality Iterative Progressive multal clustalw multalign pileup prrp poa less computation time saga hmmt better quality Fast
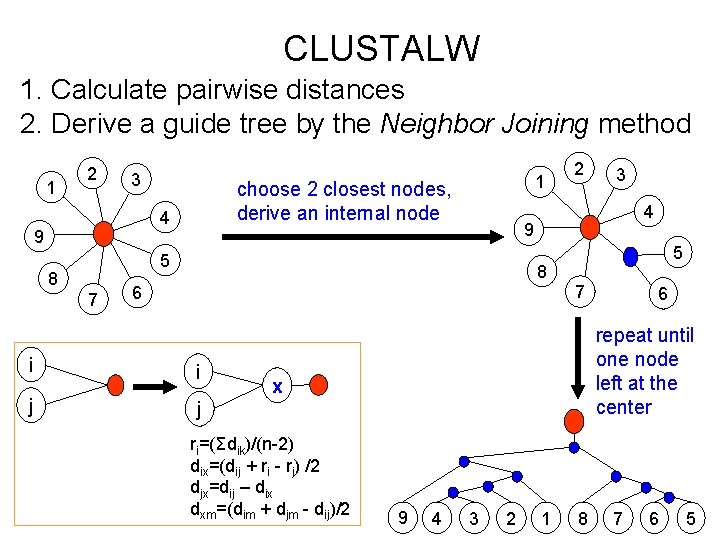
CLUSTALW 1. Calculate pairwise distances 2. Derive a guide tree by the Neighbor Joining method 1 2 3 choose 2 closest nodes, derive an internal node 4 9 1 7 3 4 9 5 8 2 5 8 7 6 i i j j repeat until one node left at the center x ri=(Σdik)/(n-2) dix=(dij + ri - rj) /2 djx=dij – dix dxm=(dim + djm - dij)/2 6 9 4 3 2 1 8 7 6 5

CLUSTALW 3. Progressively align all sequences following the guide tree • Weighted sequences 1 p e e k s a v t a l 2 g e e k a a v l a l 3 e g e w q l v l h v Without weights Score = [S(t, v) + S(l, v)] / 2 With weights Score = [S(t, v)*w 1*w 3 + S(l, v)*w 2*w 3] / 2 • 2 gap penalty values: opening, extension • Dynamically changes the gap penalty and the scoring matrix
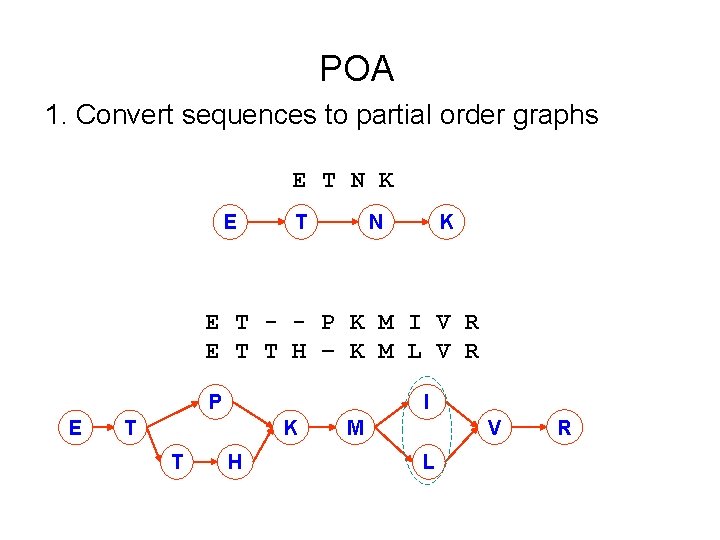
POA 1. Convert sequences to partial order graphs E T N K E T - - P K M I V R E T T H – K M L V R P E I T K T H M V L R
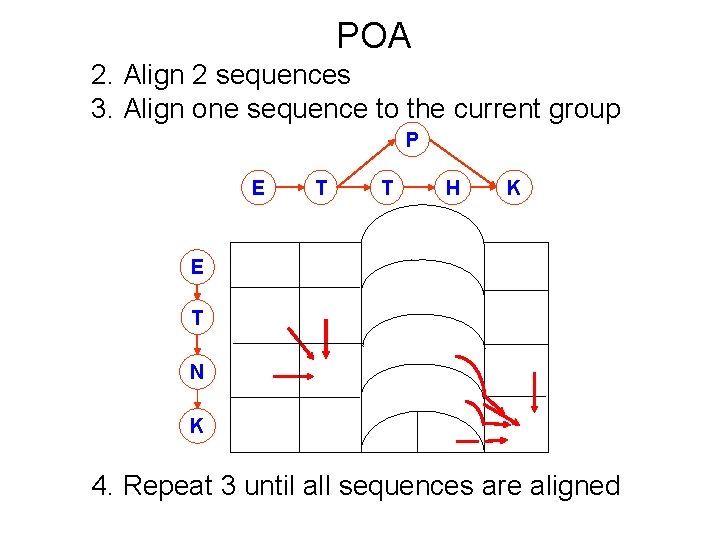
POA 2. Align 2 sequences 3. Align one sequence to the current group P E T T H K E T N K 4. Repeat 3 until all sequences are aligned

Test Results Quality Evaluation

BAli. BASE Benchmark • Reference 1: equidistance sequences with various levels of similarity. • < 25% sequence identity • 20 -40% sequence identity • > 35% sequence identity • Reference 2: closely related sequences with a highly divergent “orphan” sequence. • Reference 3: subgroups with <25% identity between groups. • Reference 4: sequences with N/C-terminal extensions. • Reference 5: sequences with internal insertions.
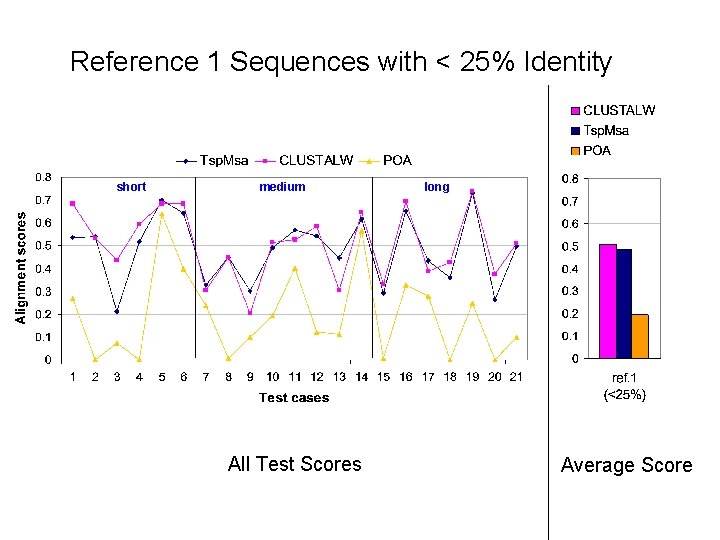
Reference 1 Sequences with < 25% Identity short medium All Test Scores long Average Score
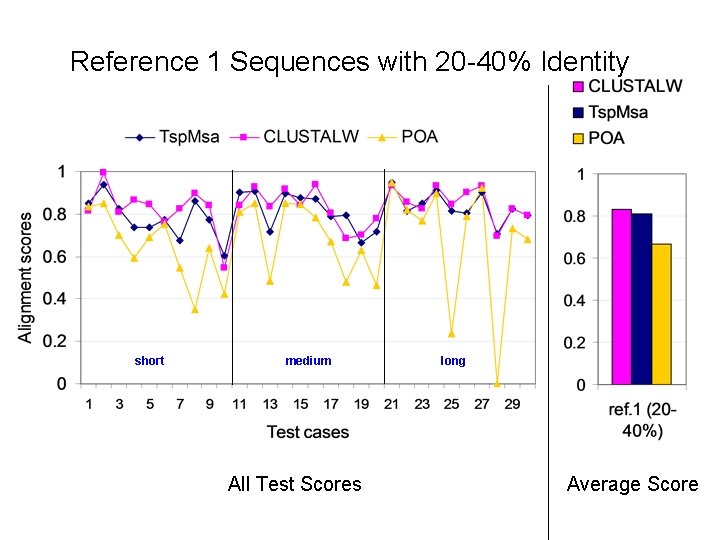
Reference 1 Sequences with 20 -40% Identity short medium All Test Scores long Average Score
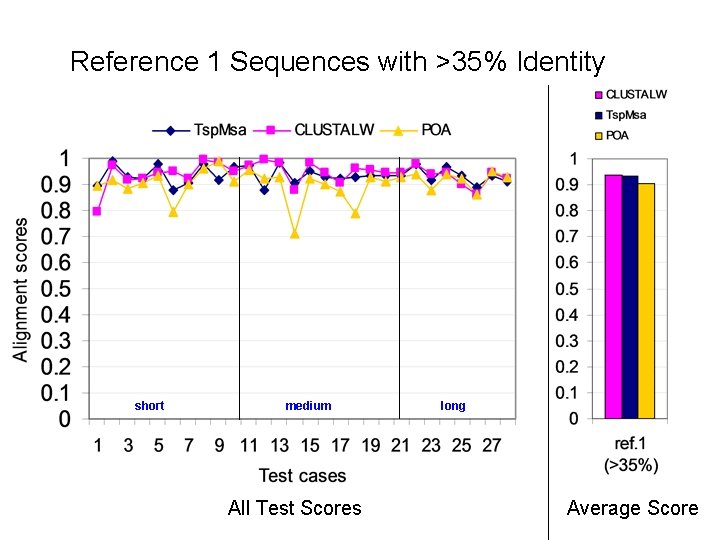
Reference 1 Sequences with >35% Identity short medium All Test Scores long Average Score
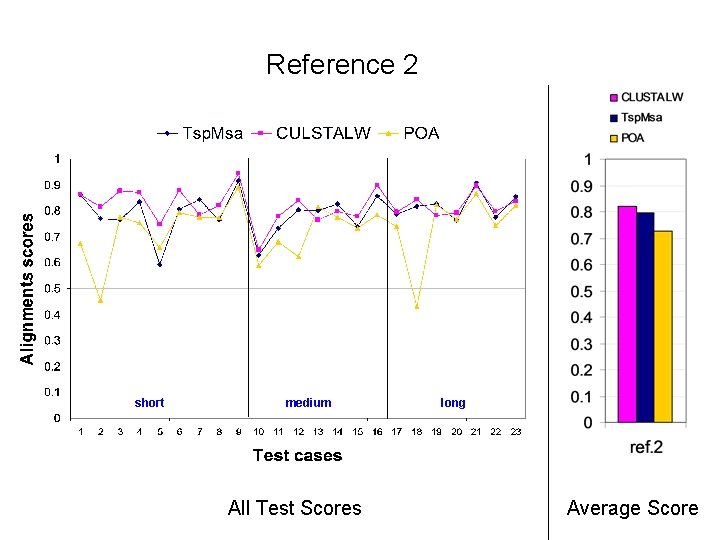
Reference 2 short medium All Test Scores long Average Score
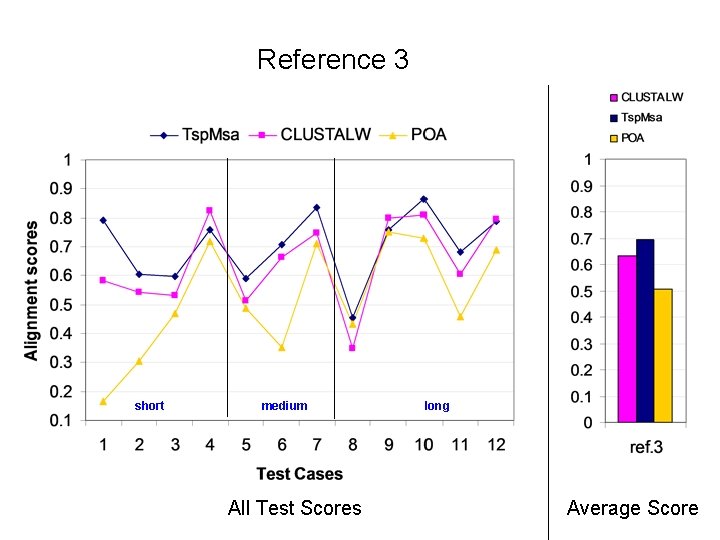
Reference 3 short medium All Test Scores long Average Score
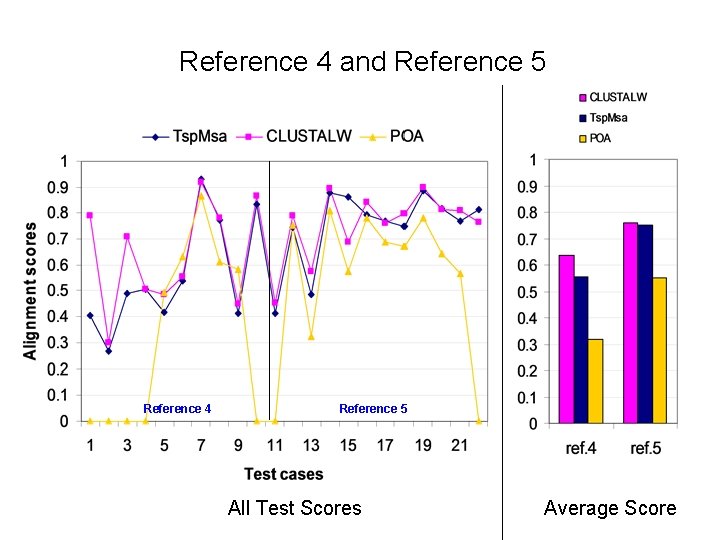
Reference 4 and Reference 5 Reference 4 Reference 5 All Test Scores Average Score

Alignment Quality Comparison Tsp. Msa and POA: Tsp. Msa better Tsp. Msa and CLUSTALW: comparable Reference 1: <25% identity: Similar * 20 -40% identity: Similar * > 35% identity: Similar Reference 2: Similar * Reference 3: Tsp. Msa better Reference 4: CLUSTALW better Reference 5: Similar * CLUSTALW slightly better for short sequences.

Test Results Execution Time Evaluation

Fast Mode Tsp. Msa Most time consuming step: Pairwise distance calculations • Slow mode: full dynamic programming (accurate) • Fast mode: a fast approximate method (heuristic)
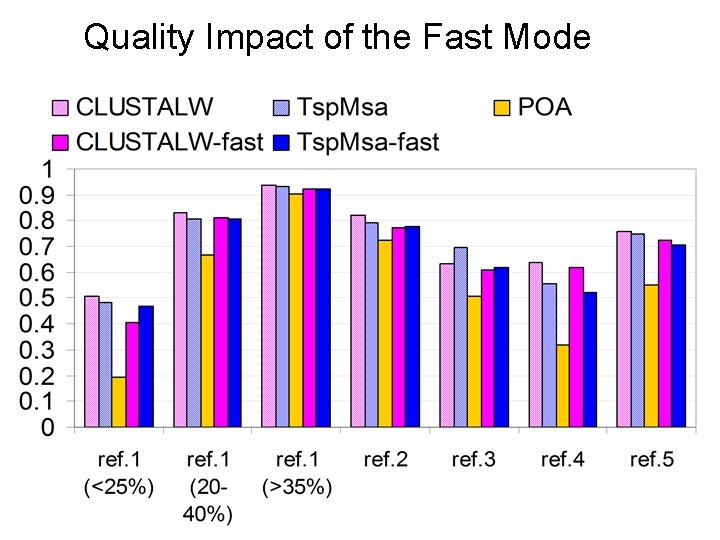
Quality Impact of the Fast Mode
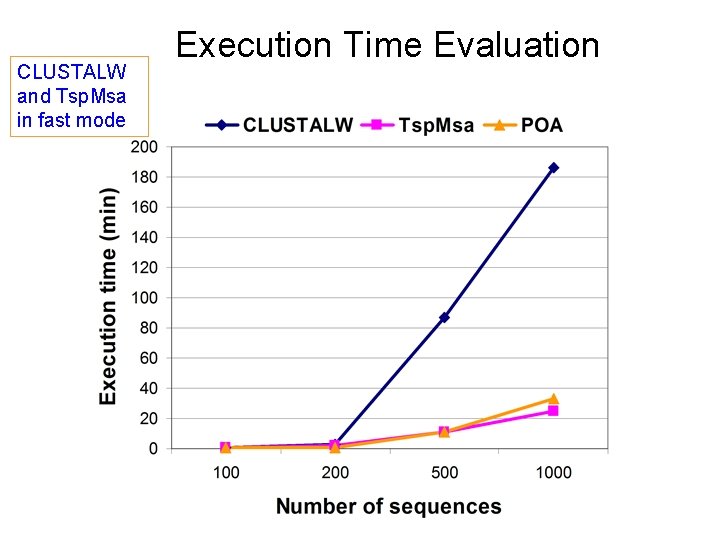
CLUSTALW and Tsp. Msa in fast mode Execution Time Evaluation

Conclusions QUALITY Slow mode • close to CLUSTALW (slow mode) • better than POA Fast mode (not as good as slow mode) • comparable to CLUSTALW (fast mode) • better than POA SPEED Fast mode • faster than CLUSTALW (fast mode) • comparable to POA

Acknowledgement Dr. Robert Robinson Dr. Russell Malmberg Dr. Eileen Kraemer Computer Science Department
- Slides: 45