Randomized Block Designs RBD and RCBD 15 2

Randomized Block Designs: RBD and RCBD (§ 15. 2, 15. 5) • Randomized block designs: – Randomized Complete Block Design – Randomized Block Design 1

Randomization in Blocked Designs For all one blocking classification designs: • Randomization of treatments to experimental units takes place within each block. • A separate randomization is required for each block. • The design is said to have one restriction on randomization. A completely randomized design requires only one randomization. Note: The randomized block design generalizes the paired t-test to the AOV setting. 2

Analysis of a RBD Traditional analysis approach is via the linear (regression on indicator variables) model and AOV. A RBD can occur in a number of situations: 1. A randomized block design with each treatment replicated once in each block (balanced and complete). This is a randomized complete block design (RCBD). 2. A randomized block design with each treatment replicated once in a block but with one block/treatment combination missing. (incomplete). 3. A randomized block design with each treatment replicated two or more times in each block (balanced and complete, with replication in each block). We will concentrate on 1 and discuss the others. 3
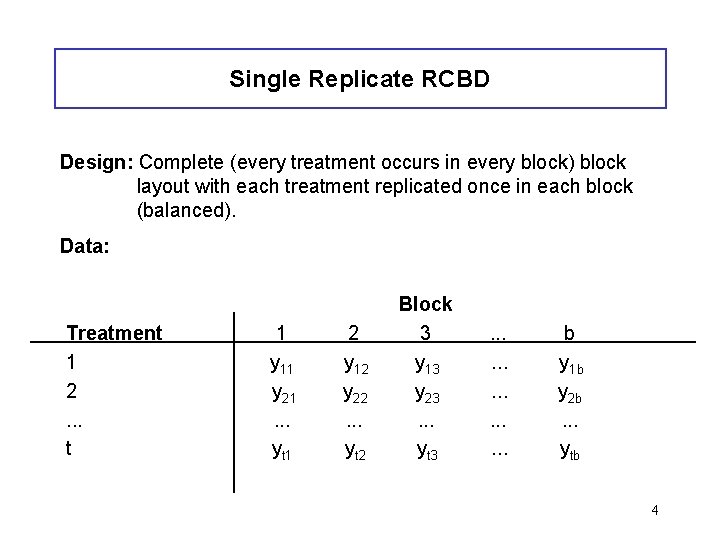
Single Replicate RCBD Design: Complete (every treatment occurs in every block) block layout with each treatment replicated once in each block (balanced). Data: Treatment 1 2. . . t 1 y 11 y 21. . . yt 1 2 y 12 y 22. . . yt 2 Block 3 y 13 y 23. . . yt 3 . . . . b y 1 b y 2 b. . . ytb 4
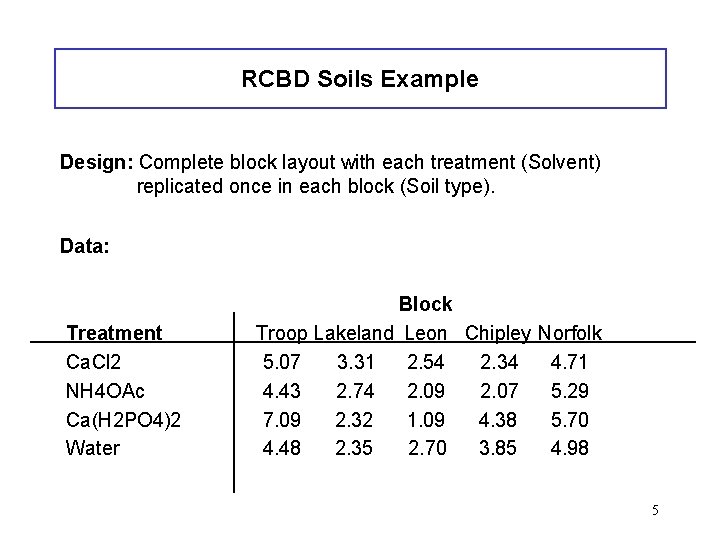
RCBD Soils Example Design: Complete block layout with each treatment (Solvent) replicated once in each block (Soil type). Data: Treatment Ca. Cl 2 NH 4 OAc Ca(H 2 PO 4)2 Water Block Troop Lakeland Leon Chipley Norfolk 5. 07 3. 31 2. 54 2. 34 4. 71 4. 43 2. 74 2. 09 2. 07 5. 29 7. 09 2. 32 1. 09 4. 38 5. 70 4. 48 2. 35 2. 70 3. 85 4. 98 5
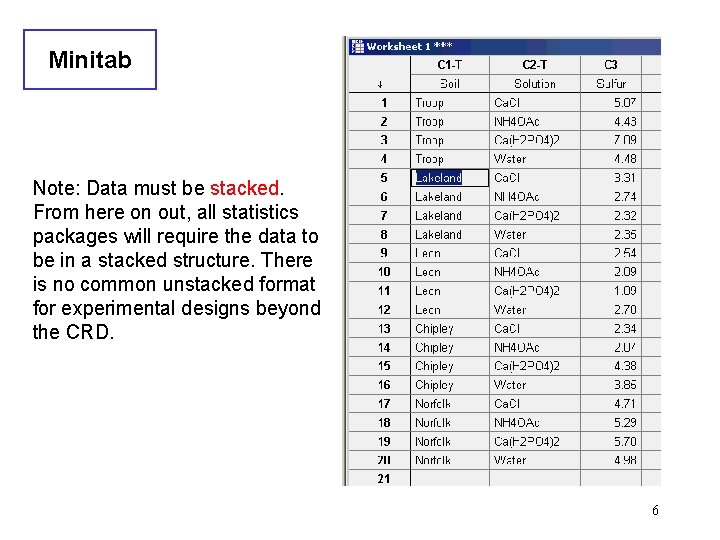
Minitab Note: Data must be stacked. From here on out, all statistics packages will require the data to be in a stacked structure. There is no common unstacked format for experimental designs beyond the CRD. 6

Linear Model: A Two-Factor (Two-Way) AOV constraints treatment i effect w. r. t. grand mean Treatment 1 2. . . t mean block j effect w. r. t. grand mean 1 m 11 m 21. . . mt 1 m + b 1 2 m 12 m 22. . . mt 2 m + b 2 Block 3 m 13 m 23. . . mt 3 m + b 3 . . . . b m 1 b m 2 b. . . mtb m + bb mean m + a 1 m + a 2 m + at 7

Model Effects Linear model Treatment effects are filtered out from block effects (show on board…) H 0 B: No block effects: b 1=b 2=b 3=. . . =bb = 0 H 0 T: No treatment effects: a 1=a 2=a 3=. . . =at = 0 SAS approach: Test with a multiple regression model with appropriate dummy variables and the F drop tests. 8
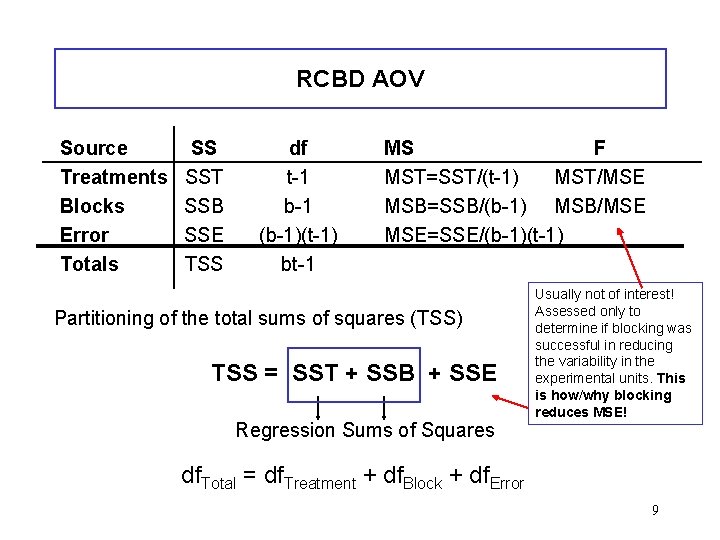
RCBD AOV Source Treatments Blocks Error Totals SS SST SSB SSE TSS df t-1 b-1 (b-1)(t-1) bt-1 MS F MST=SST/(t-1) MST/MSE MSB=SSB/(b-1) MSB/MSE MSE=SSE/(b-1)(t-1) Partitioning of the total sums of squares (TSS) TSS = SST + SSB + SSE Regression Sums of Squares Usually not of interest! Assessed only to determine if blocking was successful in reducing the variability in the experimental units. This is how/why blocking reduces MSE! df. Total = df. Treatment + df. Block + df. Error 9

Sums of Squares - RCBD Expectation under Ha. T Expectation under Ha. B Expectation of MST and MSB under respective null hypotheses is same as E(MSE) 10
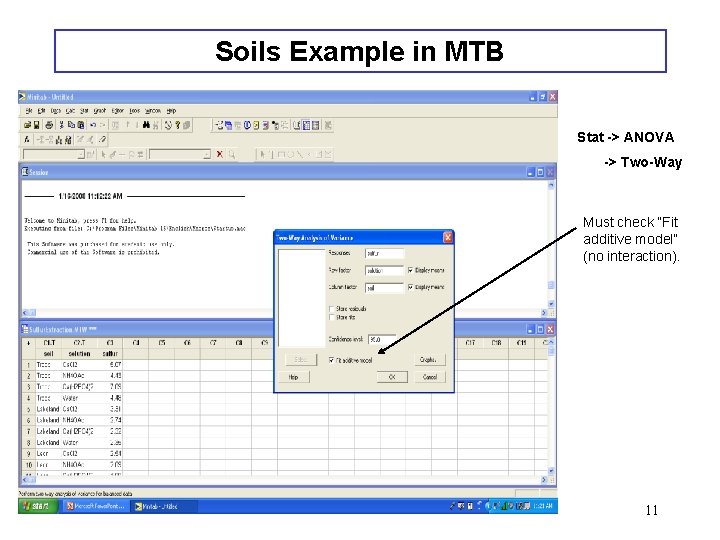
Soils Example in MTB Stat -> ANOVA -> Two-Way Must check “Fit additive model” (no interaction). 11
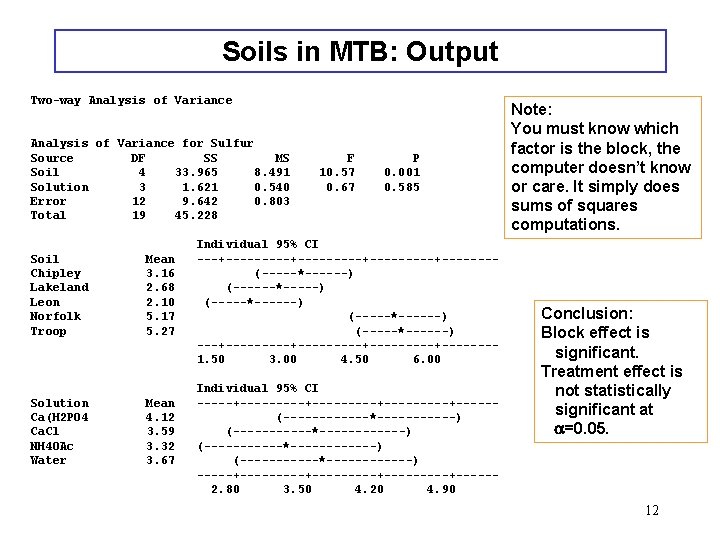
Soils in MTB: Output Two-way Analysis of Variance for Sulfur Source DF SS MS Soil 4 33. 965 8. 491 Solution 3 1. 621 0. 540 Error 12 9. 642 0. 803 Total 19 45. 228 Soil Chipley Lakeland Leon Norfolk Troop Mean 3. 16 2. 68 2. 10 5. 17 5. 27 Solution Ca(H 2 PO 4 Ca. Cl NH 4 OAc Water Mean 4. 12 3. 59 3. 32 3. 67 F 10. 57 0. 67 P 0. 001 0. 585 Individual 95% CI ---+---------+-------(-----*------) (------*-----) (-----*------) ---+---------+-------1. 50 3. 00 4. 50 6. 00 Individual 95% CI -----+---------+-----(------*-----------) (-----------*-------) (------*------) -----+---------+-----2. 80 3. 50 4. 20 4. 90 Note: You must know which factor is the block, the computer doesn’t know or care. It simply does sums of squares computations. Conclusion: Block effect is significant. Treatment effect is not statistically significant at a=0. 05. 12
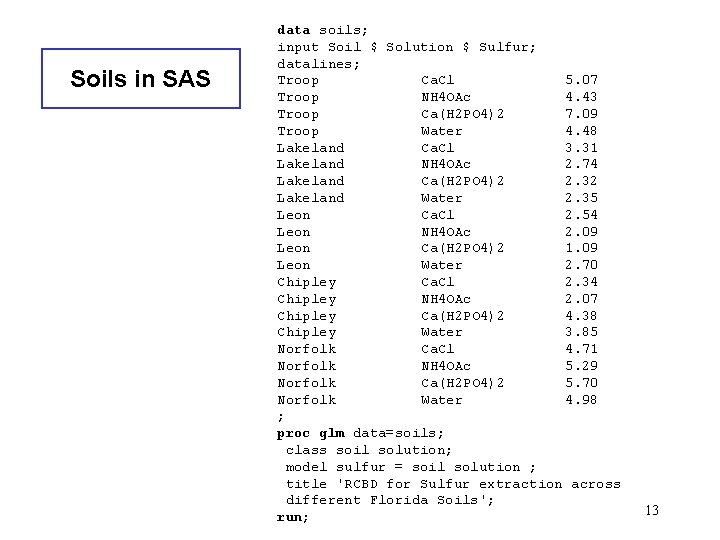
Soils in SAS data soils; input Soil $ Solution $ Sulfur; datalines; Troop Ca. Cl 5. 07 Troop NH 4 OAc 4. 43 Troop Ca(H 2 PO 4)2 7. 09 Troop Water 4. 48 Lakeland Ca. Cl 3. 31 Lakeland NH 4 OAc 2. 74 Lakeland Ca(H 2 PO 4)2 2. 32 Lakeland Water 2. 35 Leon Ca. Cl 2. 54 Leon NH 4 OAc 2. 09 Leon Ca(H 2 PO 4)2 1. 09 Leon Water 2. 70 Chipley Ca. Cl 2. 34 Chipley NH 4 OAc 2. 07 Chipley Ca(H 2 PO 4)2 4. 38 Chipley Water 3. 85 Norfolk Ca. Cl 4. 71 Norfolk NH 4 OAc 5. 29 Norfolk Ca(H 2 PO 4)2 5. 70 Norfolk Water 4. 98 ; proc glm data=soils; class soil solution; model sulfur = soil solution ; title 'RCBD for Sulfur extraction across different Florida Soils'; run; 13
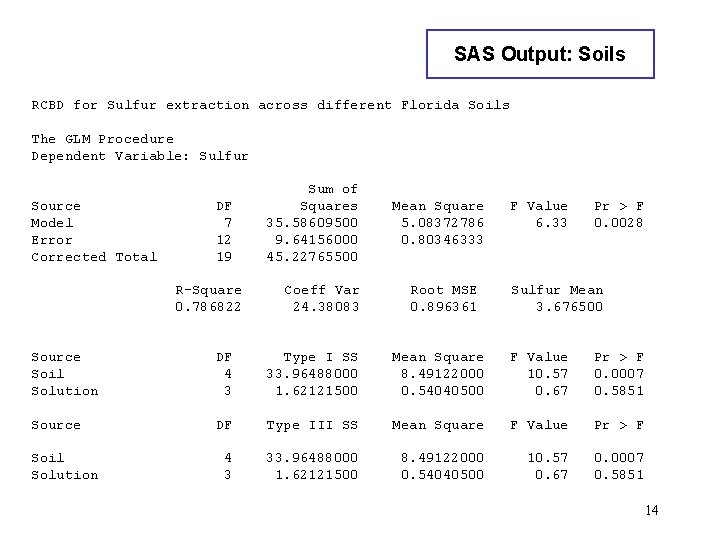
SAS Output: Soils RCBD for Sulfur extraction across different Florida Soils The GLM Procedure Dependent Variable: Sulfur Source Model Error Corrected Total DF 7 12 19 R-Square 0. 786822 Sum of Squares 35. 58609500 9. 64156000 45. 22765500 Coeff Var 24. 38083 Mean Square 5. 08372786 0. 80346333 Root MSE 0. 896361 F Value 6. 33 Pr > F 0. 0028 Sulfur Mean 3. 676500 Source Soil Solution DF 4 3 Type I SS 33. 96488000 1. 62121500 Mean Square 8. 49122000 0. 54040500 F Value 10. 57 0. 67 Pr > F 0. 0007 0. 5851 Source DF Type III SS Mean Square F Value Pr > F 4 3 33. 96488000 1. 62121500 8. 49122000 0. 54040500 10. 57 0. 67 0. 0007 0. 5851 Soil Solution 14
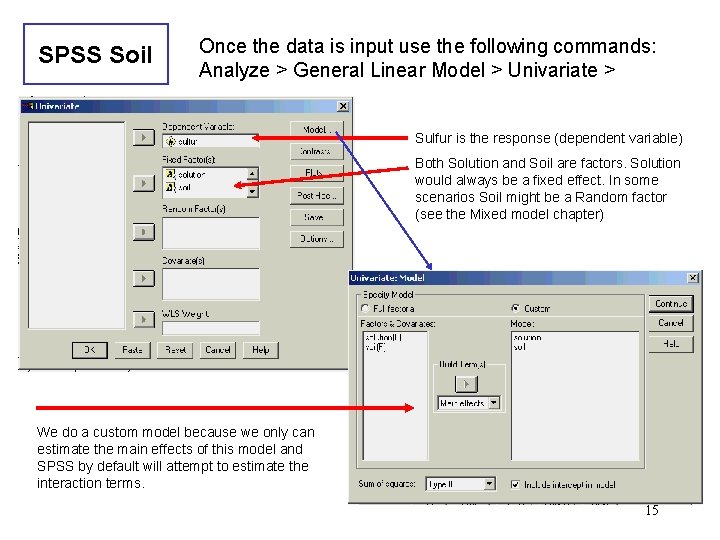
SPSS Soil Once the data is input use the following commands: Analyze > General Linear Model > Univariate > Sulfur is the response (dependent variable) Both Solution and Soil are factors. Solution would always be a fixed effect. In some scenarios Soil might be a Random factor (see the Mixed model chapter) We do a custom model because we only can estimate the main effects of this model and SPSS by default will attempt to estimate the interaction terms. 15
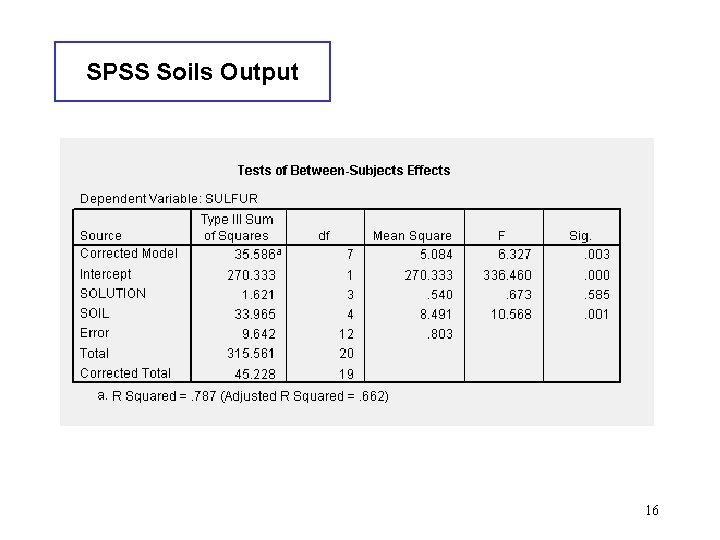
SPSS Soils Output 16

Soils RCBD in R > sulf <c(5. 07, 4. 43, 7. 09, 4. 48, 3. 31, 2. 74, 2. 32, 2. 35, 2. 54, 2. 09, 1. 09, 2. 70, 2. 34, 2. 07, 4. 38, 3. 85, 4. 71, 5. 29, 5. 70, 4. 98) > chem <- factor(rep(c("cac", "nh 4", "ca 2", "h 2 o"), 5)) > soil <factor(c(rep("Troop", 4), rep("Lake", 4), rep("Leon", 4), rep("Chip", 4), rep( "Norf", 4))) > rcbd. fit = aov(sulf~soil+chem) > # anova table > anova(rcbd. fit) Analysis of Variance Table Response: sulf Df Sum Sq Mean Sq F value Pr(>F) soil 4 33. 965 8. 491 10. 5683 0. 0006629 *** chem 3 1. 621 0. 540 0. 6726 0. 5851298 Residuals 12 9. 642 0. 803 17
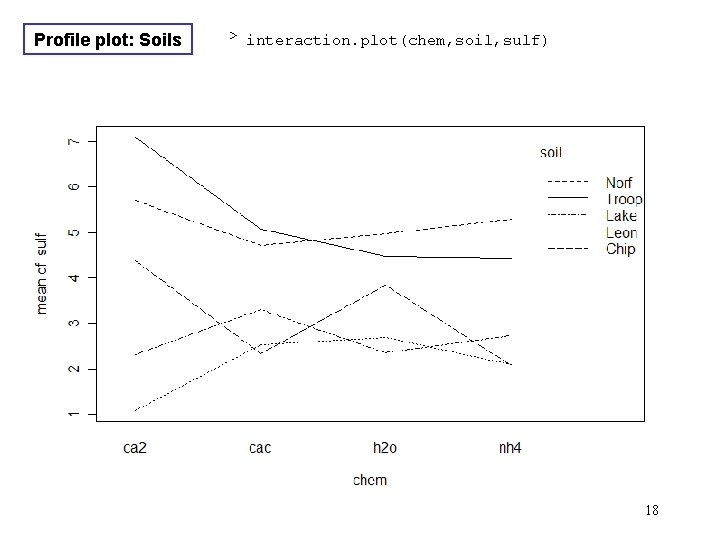
Profile plot: Soils > interaction. plot(chem, soil, sulf) 18

Nonparametric Analysis of RCBD: Friedman’s Test The RCBD, as in CRD, requires the usual AOV assumptions for the residuals: • Independence; • Homoscedasticity; • Normality. When the normality assumption fails, and transformations don’t seem to help, Friedman’s Test is a nonparametric alternative for the RCBD, just as Kruskal-Wallis was for the CRD. For example: ratings by a panel of judges (ordinal data). The procedure is based on ranks (see § 15. 5 in book), and leads to calculation of FR statistic. For large samples, we reject H 0 of equal population medians when: 19
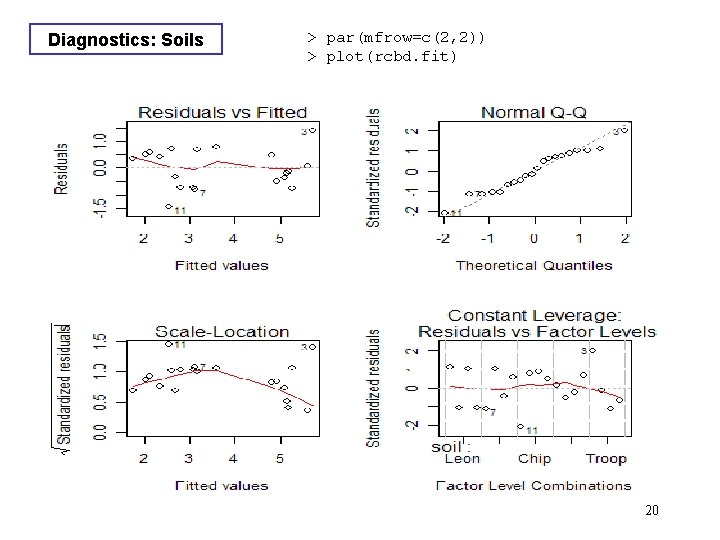
Diagnostics: Soils > par(mfrow=c(2, 2)) > plot(rcbd. fit) 20

Friedman’s Test: Soils > friedman. test(sulf, groups=chem, blocks=soil) Friedman rank sum test data: sulf, chem and soil Friedman chi-squared = 1. 08, df = 3, p-value = 0. 7819 Check group and block means: > tapply(sulf, chem, mean) ca 2 cac h 2 o nh 4 4. 116 3. 594 3. 672 3. 324 > tapply(sulf, soil, mean) Chip Lake Leon Norf Troop 3. 1600 2. 6800 2. 1050 5. 1700 5. 2675 21
- Slides: 21