Plant Ontology Consortium www plantontology org In this















- Slides: 30

Plant Ontology Consortium www. plantontology. org

In this presentation: l l l l Why develop “Plant Ontology (PO)”? What is PO? How is PO designed and structured? Annotations to PO Who is using PO? How to use PO in your research? How to request your PO term?

Why develop PO? l More and more plants genome are getting sequenced l More and more datatypes are involved l More and more databases are developed

We need controlled vocabulary A set of defined terms to describe knowledge of a specific domain l Avoid ambiguity - conducting a search in a database that use controlled vocabulary is efficient and precise l User can make a meaningful cross-species query across databases

They are all “Fruit” The seed-bearing structure in angiosperms, formed from the ovary after flowering Maize kernel Arabidopsis silique Peapod Rice grain/caryopsis Berry

What is “ Plant Ontology “ ? l PO is an arrangement of controlled vocabularies - plant anatomical, morphological structures - plant growth and developmental stages l Based on internationally published/accepted terminology and their definitions l Recognized by computers l A tool for annotation of gene expression patterns and phenotypes of germplasms across angiosperms

What is in a PO term?

Rationale on creating a term to describe plant structure l l l Botanical terms - anatomy and morphology Derivation - origin of plant parts and cell lineage; spatial/positional organization of tissues, organs and organ systems (mainly for annotation purpose) Avoids subcellular structure (GO has it) Avoids attributes (qualifiers) of anatomical terms Use synonym fields to cover various instances, types and terms with attributes

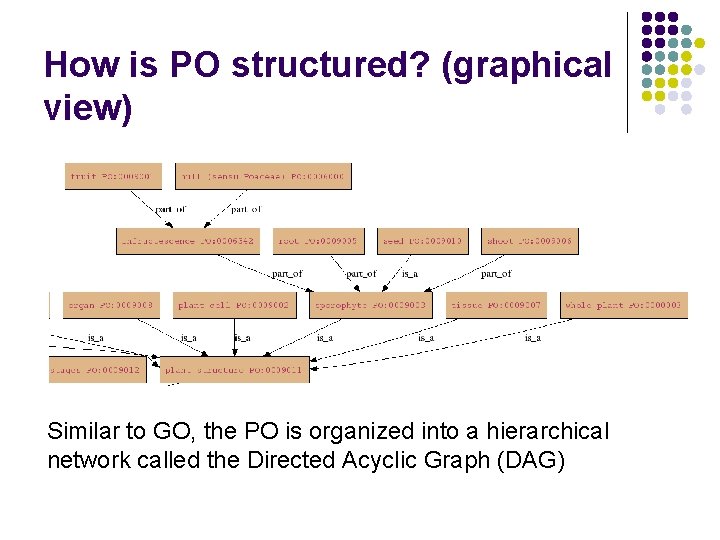
How is PO structured? (graphical view) Similar to GO, the PO is organized into a hierarchical network called the Directed Acyclic Graph (DAG)

How is PO structured? (tree view via Ami. Go)

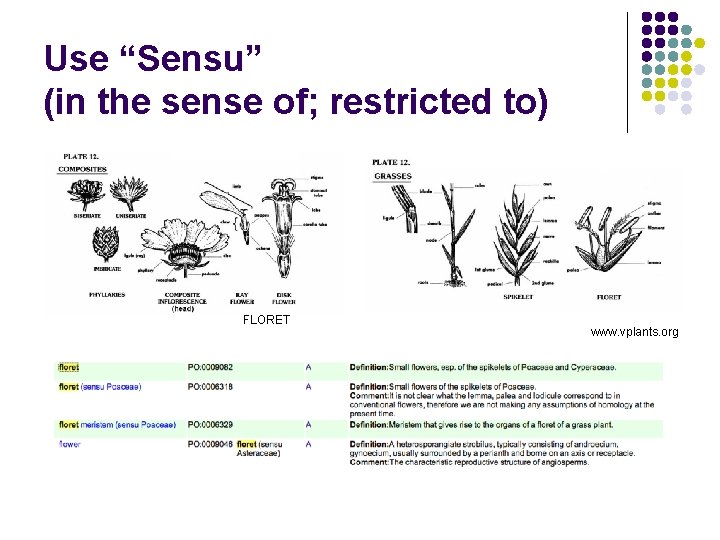
Use “Sensu” (in the sense of; restricted to) FLORET www. vplants. org

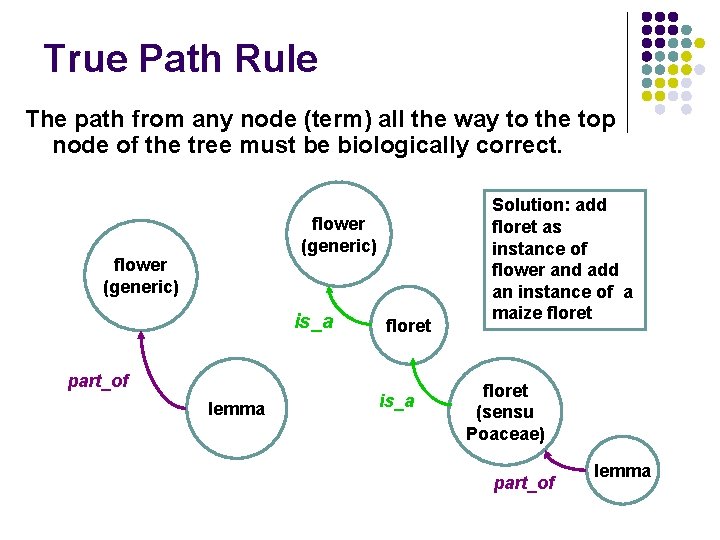
True Path Rule The path from any node (term) all the way to the top node of the tree must be biologically correct. flower (generic) is_a part_of lemma floret is_a Solution: add floret as instance of flower and add an instance of a maize floret (sensu Poaceae) part_of lemma

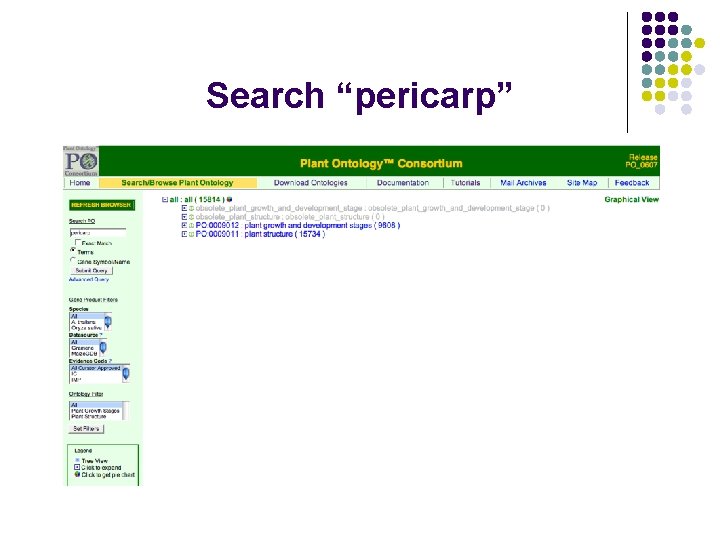
Annotation using PO Objects for annotation: Genes, Gene products, such as transcripts, proteins, ESTs, c. DNA, QTLs, mutants, germplasm, phenotypes, microarray data ……etc Example: Annotations to a plant structure term: pericarp (PO: 0009084) in the POC database

Visit www. plantontology. org

Search “pericarp”

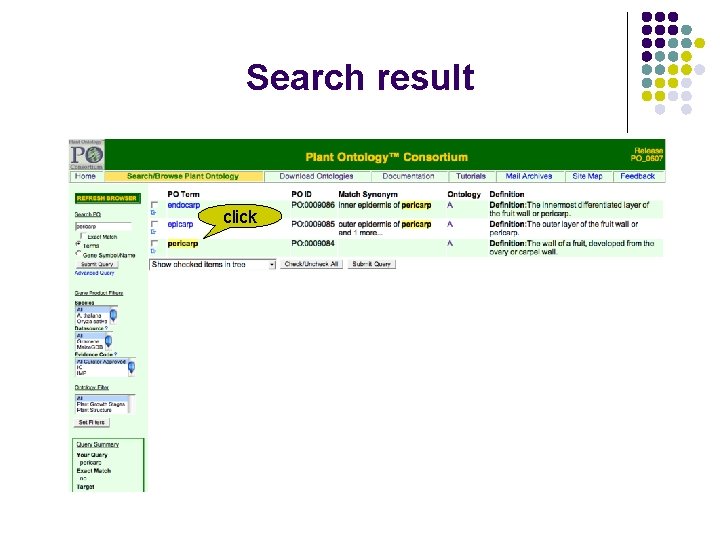
Search result click

Description of “pericarp” scroll

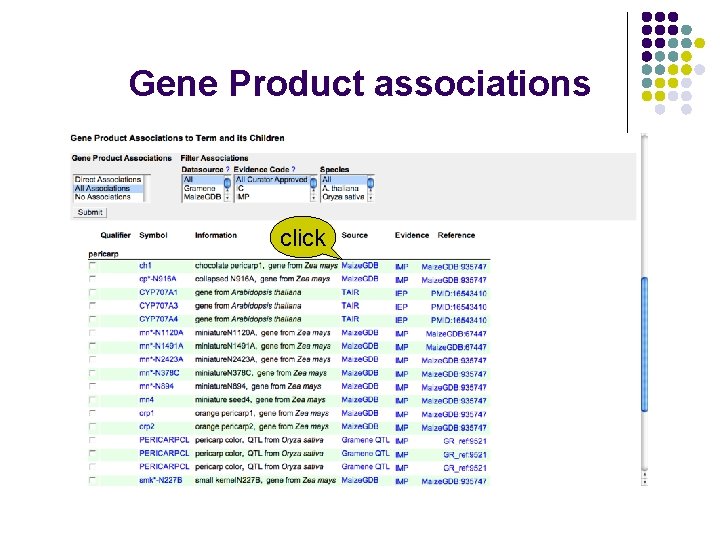
Gene Product associations click

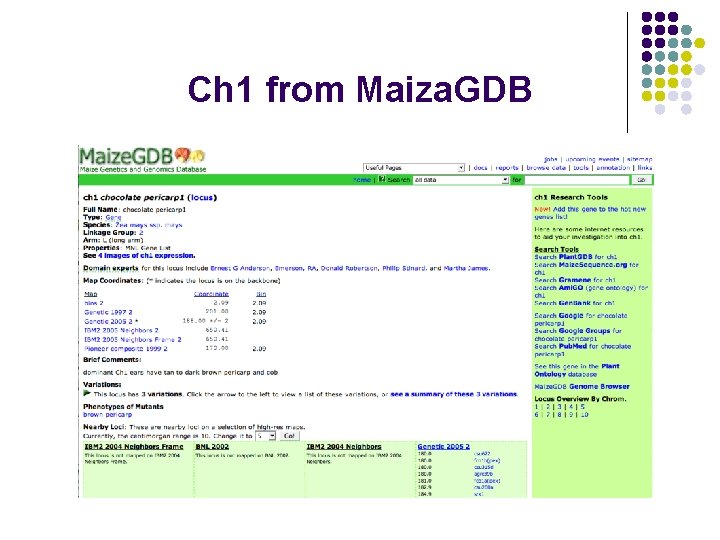
Ch 1 from Maiza. GDB

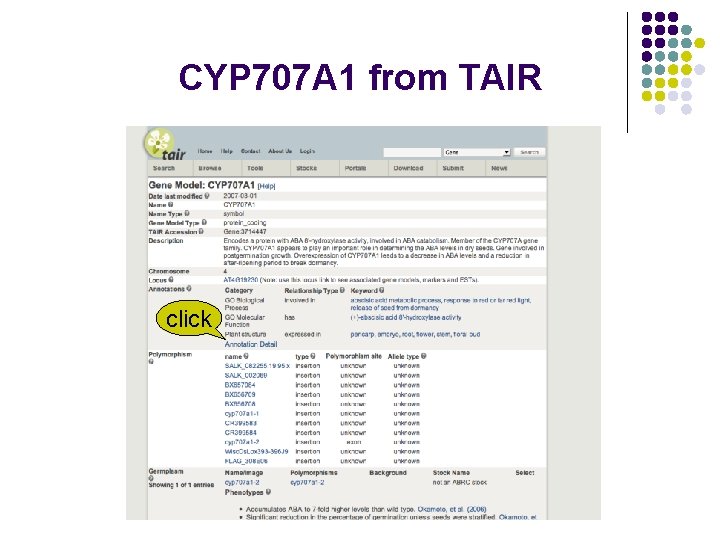
CYP 707 A 1 from TAIR click

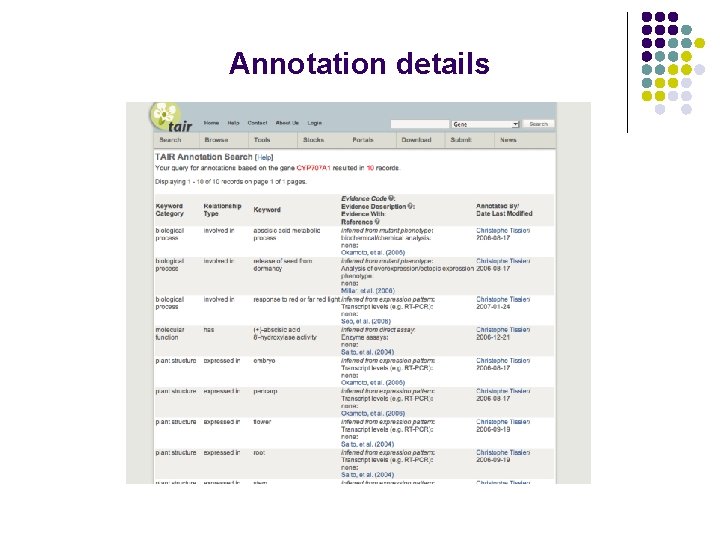
Annotation details

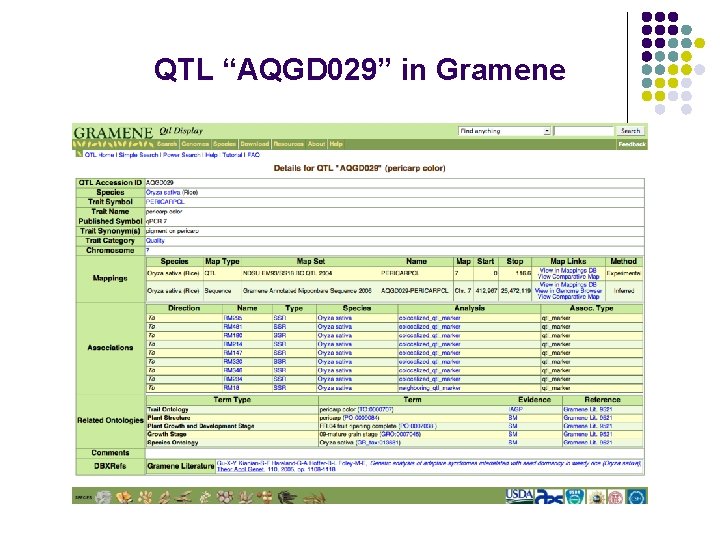
QTL “AQGD 029” in Gramene

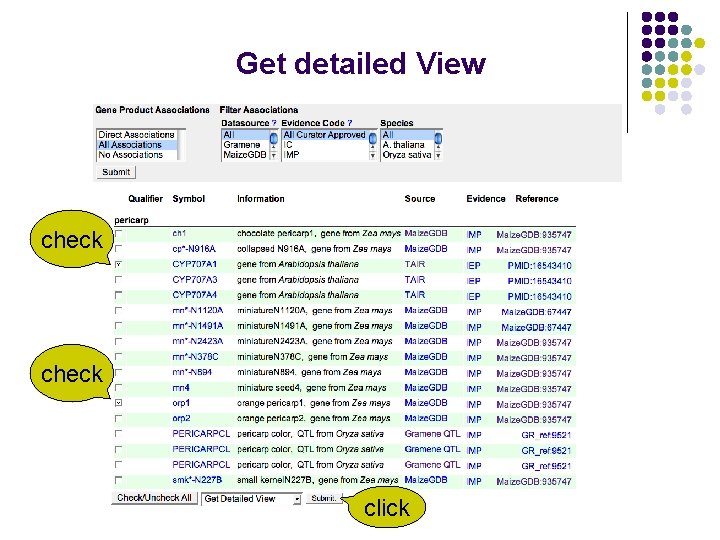
Get detailed View check click

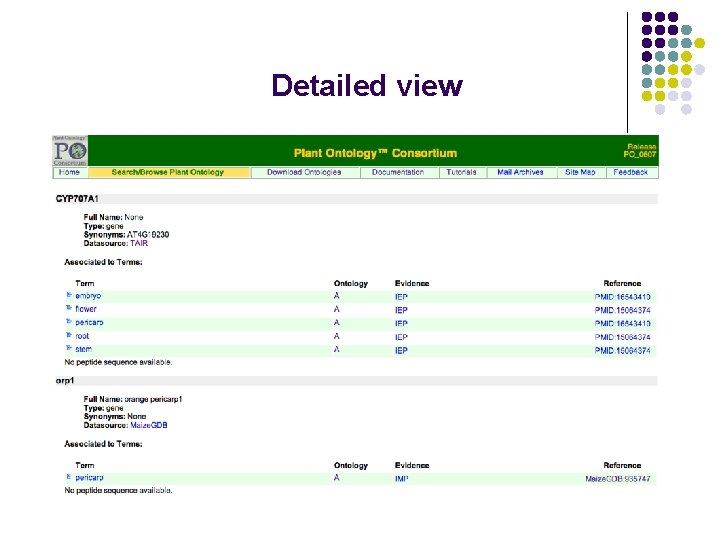
Detailed view

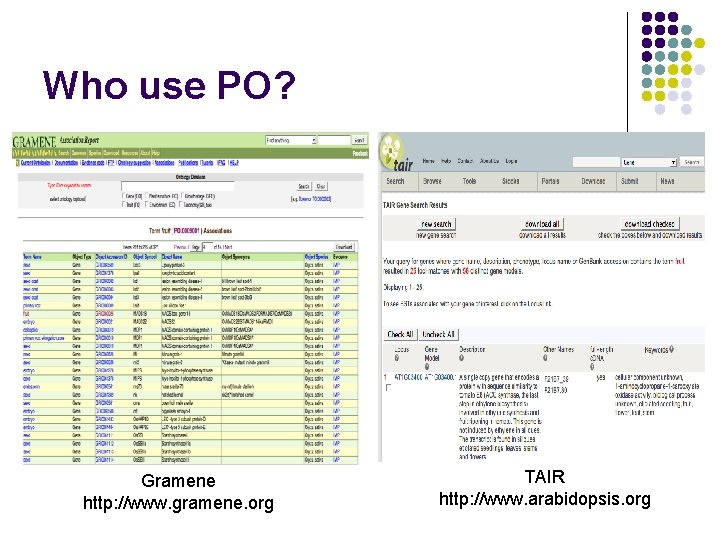
Who use PO? Gramene http: //www. gramene. org TAIR http: //www. arabidopsis. org

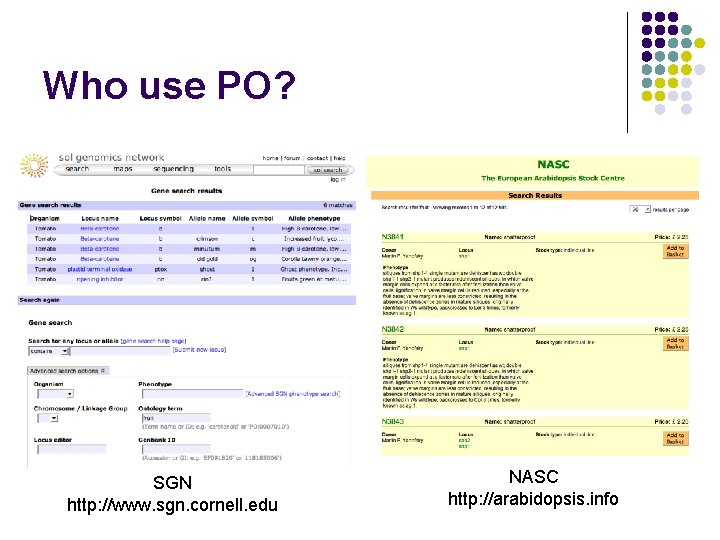
Who use PO? SGN http: //www. sgn. cornell. edu NASC http: //arabidopsis. info

Who use PO? Genevestigator https: //www. genevestigator. ethz. ch/ BRENDA http: //www. brenda. uni-koeln. de/

There are more, Yours is here!

How to apply PO in your research? l Find appropriate PO terms for your species of interest; if no term, request it! l Curate your objects using PO l Start making cross-species querying l Submit your annotation to POC or species-specific databases

Submit your PO term request!