Minimal Recombinations Histories and Global Pedigrees Finding Minimal

Minimal Recombinations Histories and Global Pedigrees Finding Minimal Recombination Histories 1 2 3 4 1 4 3 2 Finding Common Ancestors Global Pedigrees NOW Acknowledgements Yun Song - Rune Lyngsø - Mike Steel - Carsten Wiuf
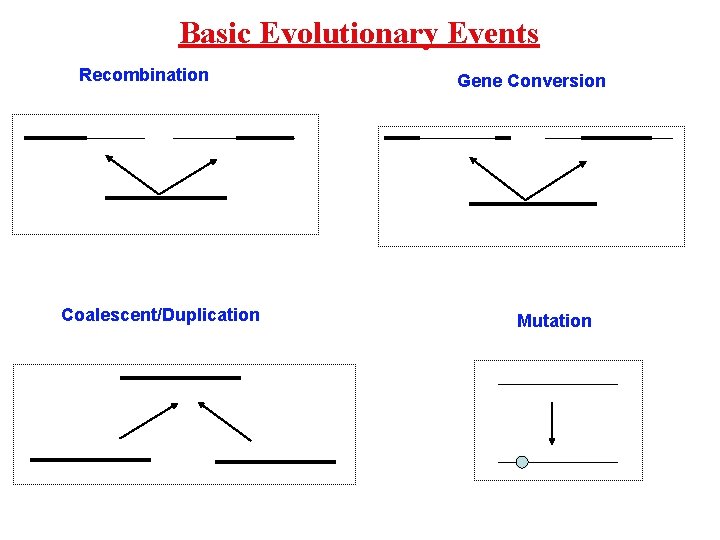
Basic Evolutionary Events Recombination Coalescent/Duplication Gene Conversion Mutation
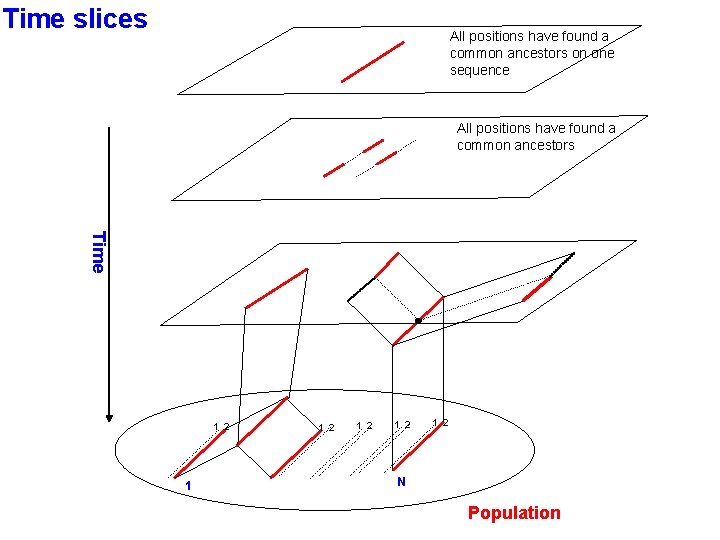
Time slices All positions have found a common ancestors on one sequence All positions have found a common ancestors Time 1 2 1 2 1 2 N Population
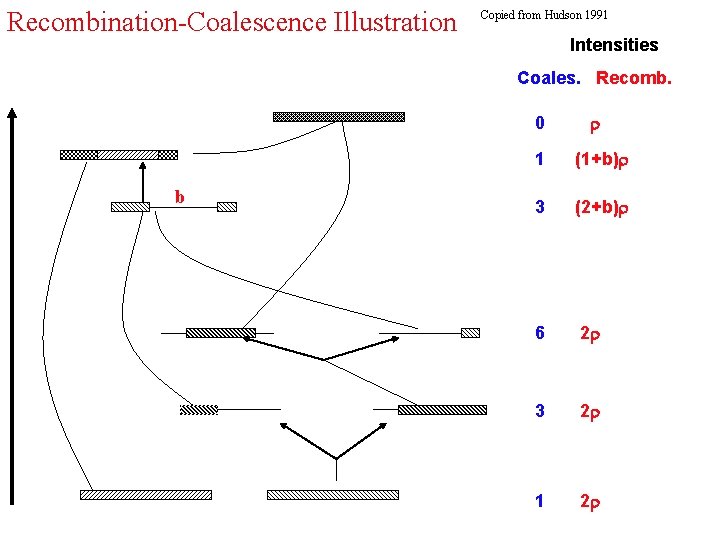
Recombination-Coalescence Illustration Copied from Hudson 1991 Intensities Coales. Recomb. 0 b 1 (1+b) 3 (2+b) 6 2 3 2 1 2
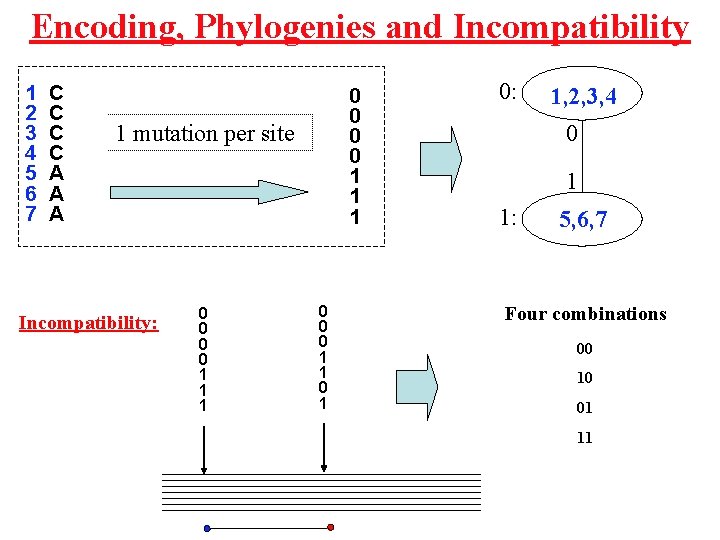
Encoding, Phylogenies and Incompatibility 1 2 3 4 5 6 7 C C A A A 0 0 1 1 mutation per site Incompatibility: 0 0 1 1 1 0 0 0 1 1 0: 1, 2, 3, 4 0 1 1: 5, 6, 7 Four combinations 00 10 01 11

The 1983 Kreitman Data & the infinite site assumption (M. Kreitman 1983 Nature) • 11 sequences of alcohol dehydrogenase gene in Drosophila melanogaster. • Can be reduced to 9 sequences (3 of 11 are identical). • 3200 bp long, 43 segregating sites, 28 of which are informative Recoded Kreitman data i. (0, 1) ancestor state known ii. Multiple copies represented by 1 sequence iii. Non-informative sites could be removed

Hudson & Kaplan’s RM 00000111110000 001000011111000000000000010 00000110000000010 0001111000000001 001000000001010111 001000000011111101 1111100000011111101 11111100100000011111101 If you equate RM with expected number of recombinations, this could be used as an estimator. Unfortunately, RM is a gross underestimate of the real number of recombinations.

Recombination Parsimony Hein, 1990, 93 & Song & Hein, 2002+ Data 1 2 3 Trees T 1 2 i-1 i L
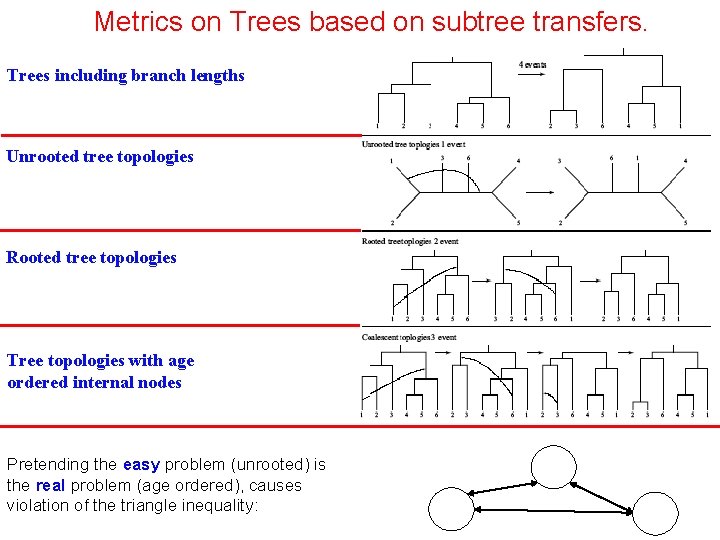
Metrics on Trees based on subtree transfers. Trees including branch lengths Unrooted tree topologies Rooted tree topologies Tree topologies with age ordered internal nodes Pretending the easy problem (unrooted) is the real problem (age ordered), causes violation of the triangle inequality:
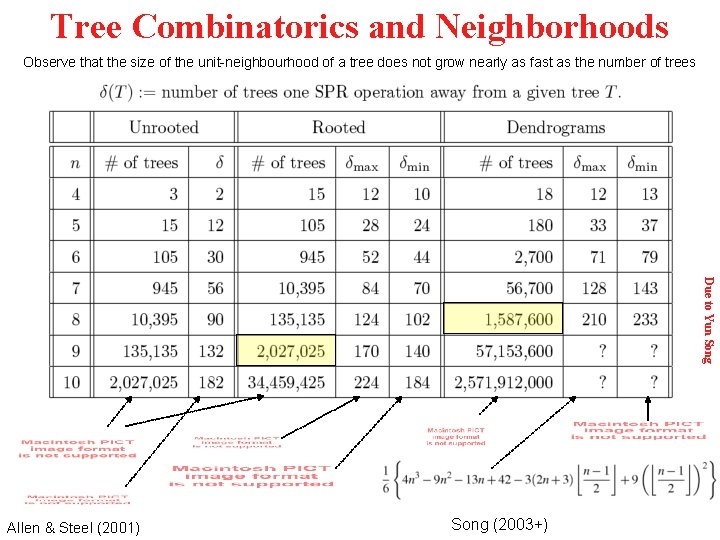
Tree Combinatorics and Neighborhoods Observe that the size of the unit-neighbourhood of a tree does not grow nearly as fast as the number of trees Due to Yun Song Allen & Steel (2001) Song (2003+)

1
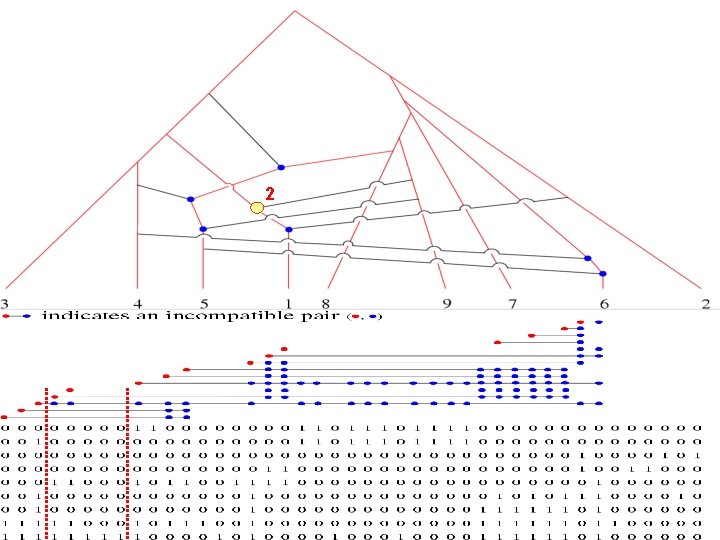
2
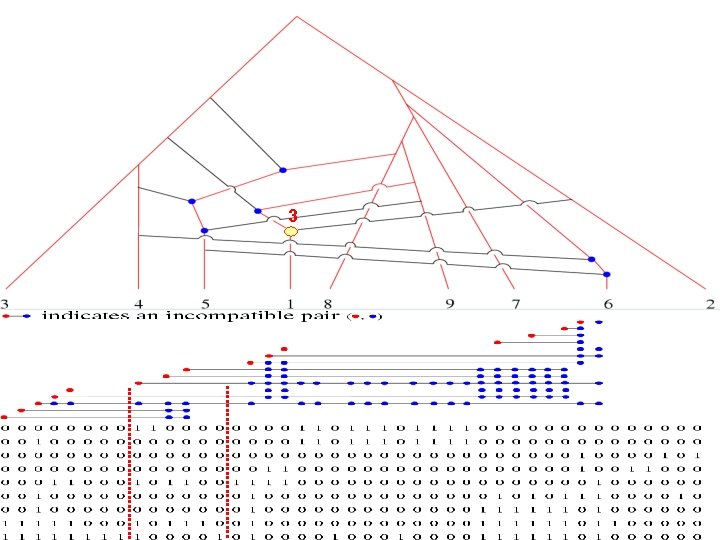
3
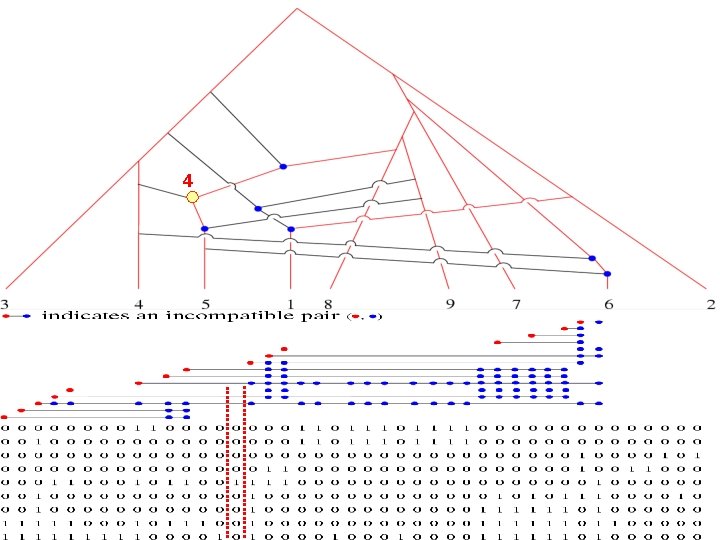
4
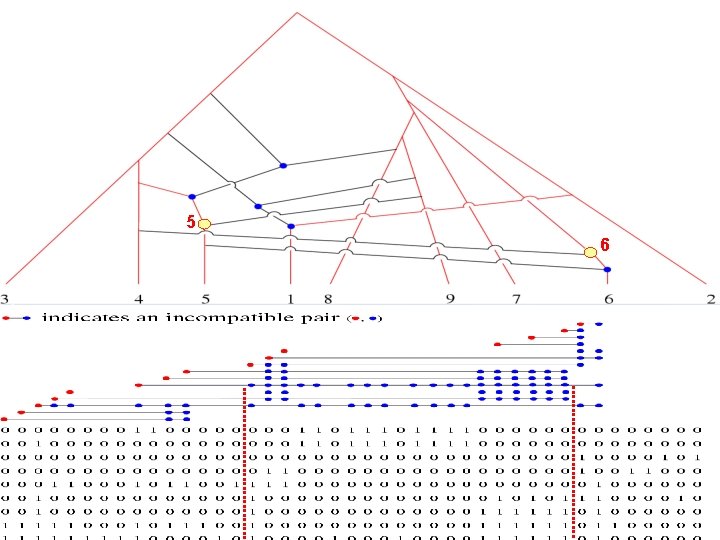
5 6

7

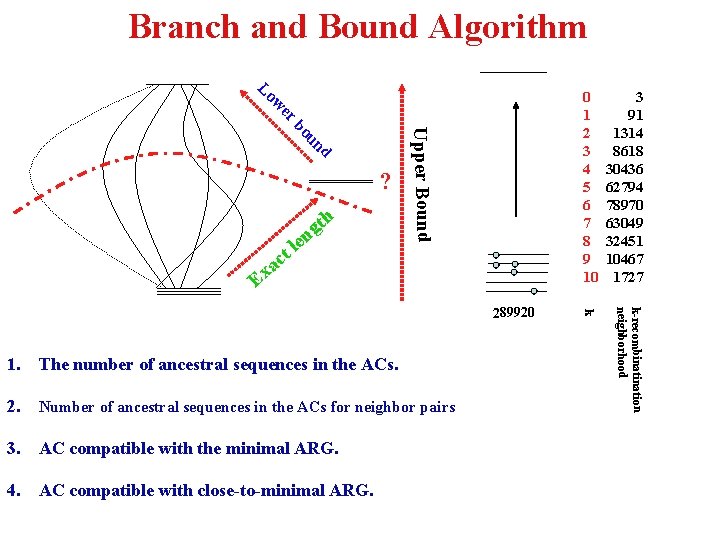
Branch and Bound Algorithm E 289920 1. The number of ancestral sequences in the ACs. 2. Number of ancestral sequences in the ACs for neighbor pairs 3. AC compatible with the minimal ARG. 4. AC compatible with close-to-minimal ARG. k-recombination neighborhood le t c xa Upper Bound d un bo h t ng 3 91 1314 8618 30436 62794 78970 63049 32451 10467 1727 k er w Lo ? 0 1 2 3 4 5 6 7 8 9 10
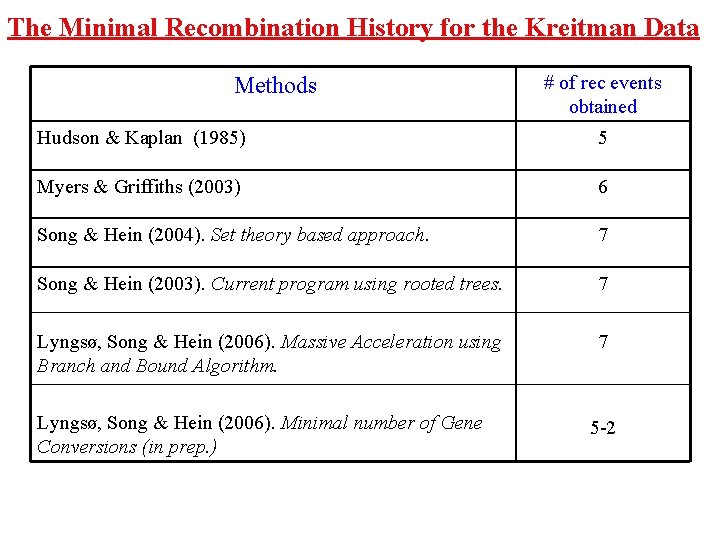
The Minimal Recombination History for the Kreitman Data Methods # of rec events obtained Hudson & Kaplan (1985) 5 Myers & Griffiths (2003) 6 Song & Hein (2004). Set theory based approach. 7 Song & Hein (2003). Current program using rooted trees. 7 Lyngsø, Song & Hein (2006). Massive Acceleration using Branch and Bound Algorithm. 7 Lyngsø, Song & Hein (2006). Minimal number of Gene Conversions (in prep. ) 5 -2
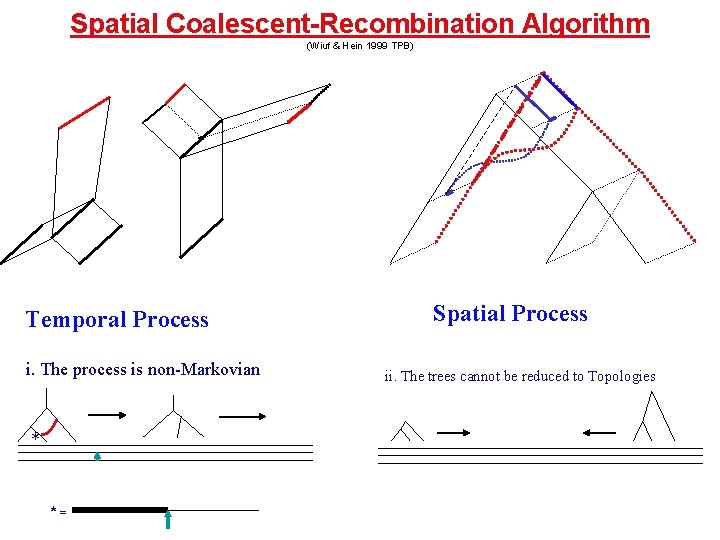
Spatial Coalescent-Recombination Algorithm (Wiuf & Hein 1999 TPB) Temporal Process i. The process is non-Markovian * *= Spatial Process ii. The trees cannot be reduced to Topologies
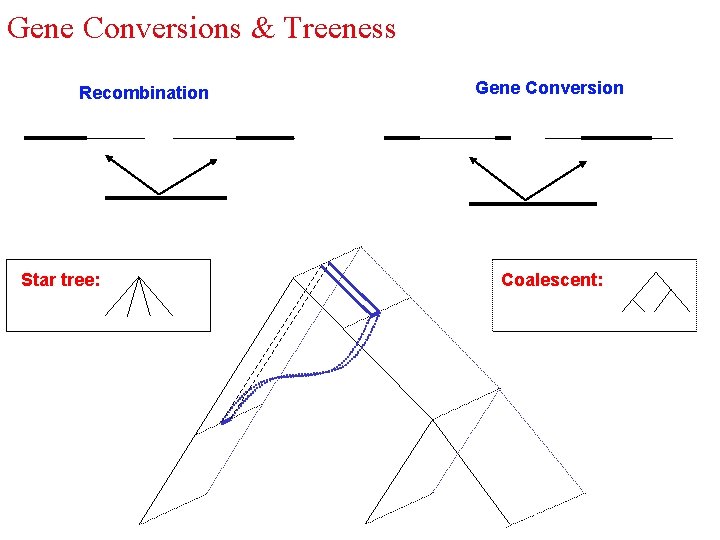
Gene Conversions & Treeness Recombination Star tree: Gene Conversion Coalescent:

The Bad News: Actual, potentially detectable and detected recombinations 1 2 3 4 Leaves 2 3 4 5 6 10 15 Root 1. 0 1. 33 1. 50 1. 66 1. 80 1. 87 500 1. 99 Edge-Length 2. 0 3. 66 4. 16 4. 57 5. 66 6. 50 Topo-Diff 0. 073 0. 134 0. 183 0. 300 0. 374 Tree-Diff. 666. 694. 714. 728. 740. 769. 790 0. 670 Minimal ARG True ARG 0 4 Mb
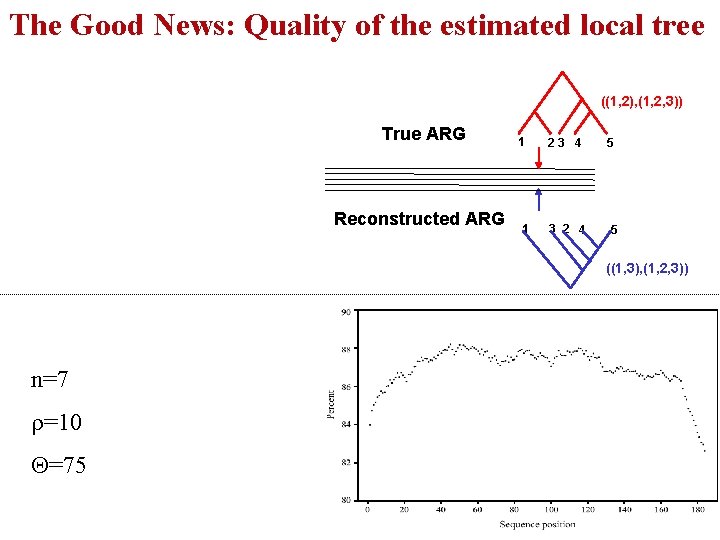
The Good News: Quality of the estimated local tree ((1, 2), (1, 2, 3)) True ARG Reconstructed ARG 1 1 23 4 3 2 4 5 5 ((1, 3), (1, 2, 3)) n=7 r=10 Q=75
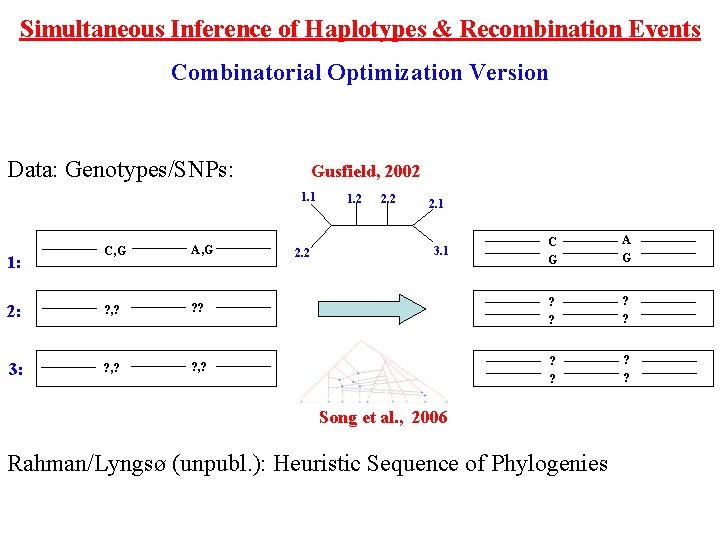
Simultaneous Inference of Haplotypes & Recombination Events Combinatorial Optimization Version Data: Genotypes/SNPs: Gusfield, 2002 1. 1 2. 2 2. 1 C G A G ? ? ? , ? ? ? C, G A, G 2: ? , ? 3: ? , ? 1: 1. 2 2. 2 3. 1 Song et al. , 2006 Rahman/Lyngsø (unpubl. ): Heuristic Sequence of Phylogenies

The Griffiths-Ethier-Tavare Recursions No recombination: Infinite Site Assumption Ancestral State Known History Graph: Recursions Exists No cycles Possible Histories without Recombination for simple data example 0 1 1 1 2 4 3 5 4 5 5 5 6 3 7 2 8 - recombination 27 ACs 8 1

Ancestral configurations to 2 sequences with 2 segregating sites: k 1 (k 2+1)*k 1 +1 possible ancestral columns. k 2 2 nd 1 st

Counting Recursion Summary statistic lumping configurations k 1(k 2+1)+1 padded with “-” + 1 k+1 k
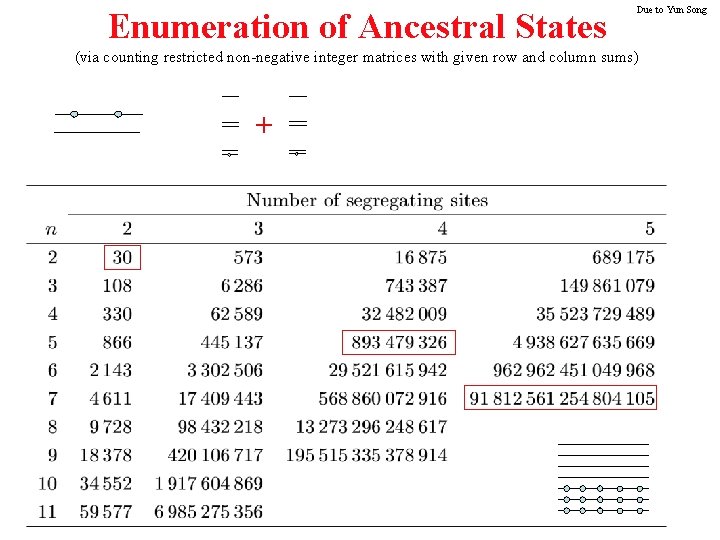
Enumeration of Ancestral States Due to Yun Song (via counting restricted non-negative integer matrices with given row and column sums) +
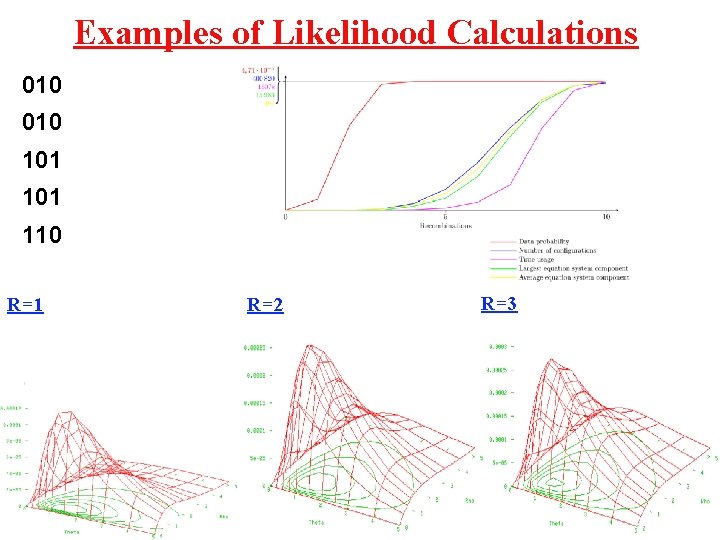
Examples of Likelihood Calculations 010 101 110 R=1 R=2 R=3
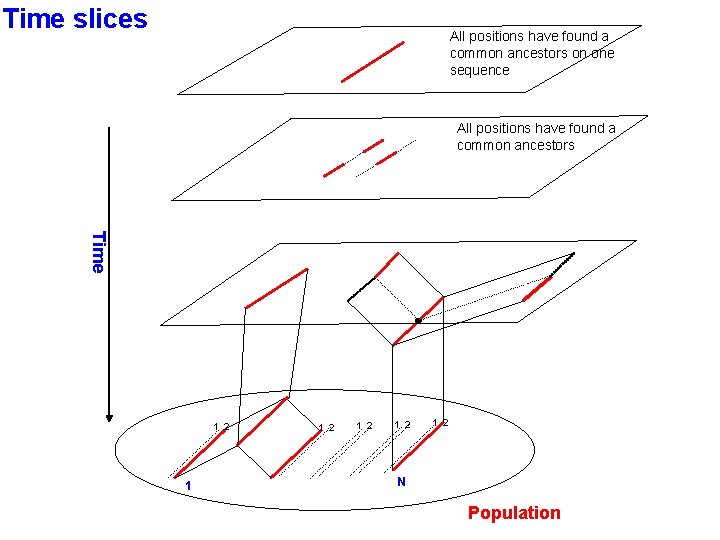
Time slices All positions have found a common ancestors on one sequence All positions have found a common ancestors Time 1 2 1 2 1 2 N Population
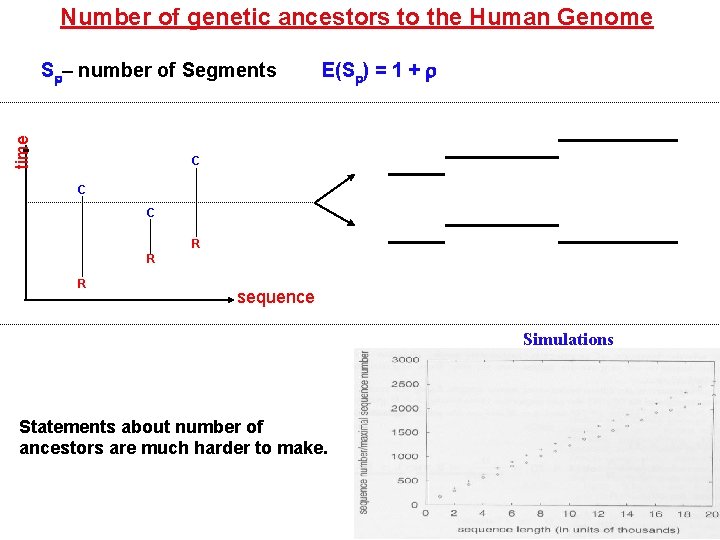
Number of genetic ancestors to the Human Genome time S – number of Segments E(S ) = 1 + C C C R R R sequence Simulations Statements about number of ancestors are much harder to make.

Applications to Human Genome (Wiuf and Hein, 97) Parameters used 4 Ne 20. 000 Chromos. 1: 263 Mb. 263 c. M Chromosome 1: Segments 52. 000 Ancestors 6. 800 All chromosomes Ancestors 86. 000 Physical Population. 1. 3 -5. 0 Mill. A randomly picked ancestor: 0 260 Mb 0 52. 000 0 7. 5 Mb 8360 6890 *250 0 (ancestral material comes in batteries!) 30 kb *35

Multiple and Simultaneous Coalescents 1. Simultaneous Events 2. Multifurcations. 3. Underestimation of Coalescent Rates
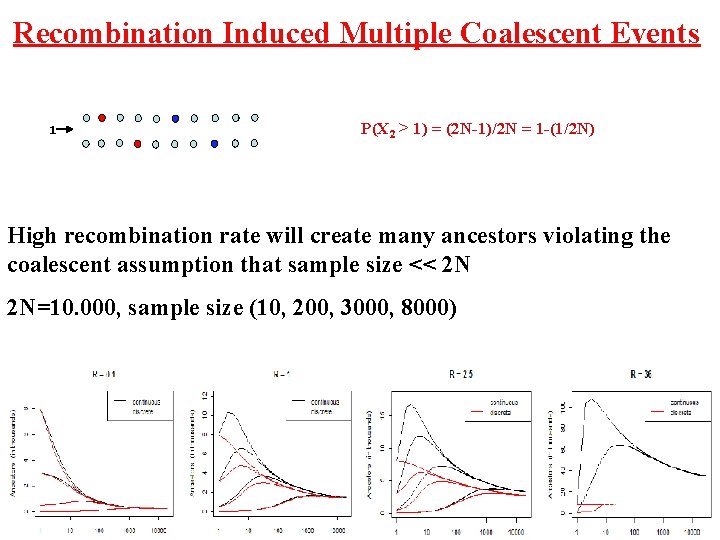
Recombination Induced Multiple Coalescent Events 1 P(X 2 > 1) = (2 N-1)/2 N = 1 -(1/2 N) High recombination rate will create many ancestors violating the coalescent assumption that sample size << 2 N 2 N=10. 000, sample size (10, 200, 3000, 8000)
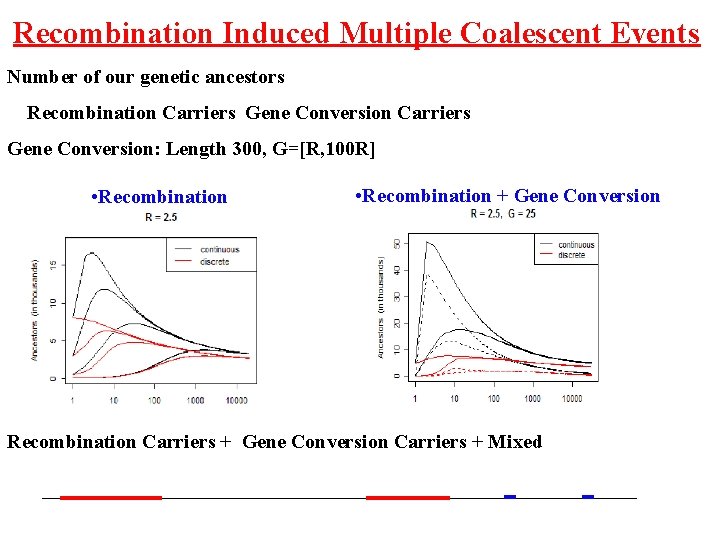
Recombination Induced Multiple Coalescent Events Number of our genetic ancestors Recombination Carriers Gene Conversion: Length 300, G=[R, 100 R] • Recombination + Gene Conversion Recombination Carriers + Gene Conversion Carriers + Mixed
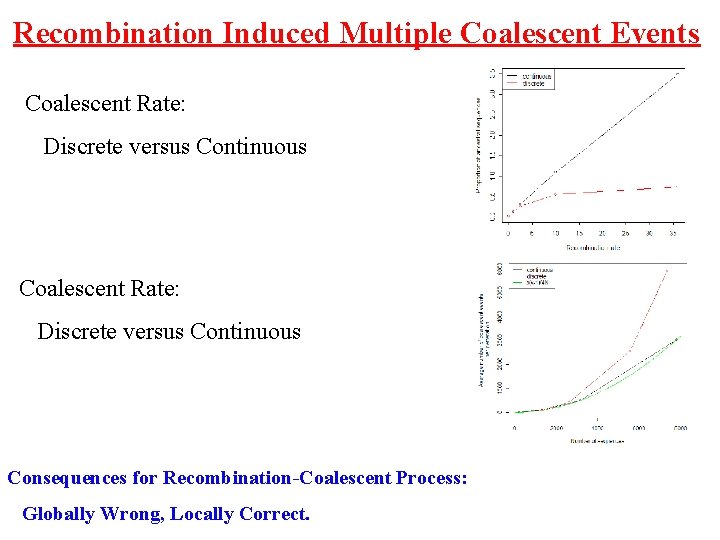
Recombination Induced Multiple Coalescent Events Coalescent Rate: Discrete versus Continuous Consequences for Recombination-Coalescent Process: Globally Wrong, Locally Correct.

Questions based on Large Data Sets Much more sequence data 1. Comparative Genomics of a Huge Scale 2. Population Genomics One issue: reconstructing population pedigrees. Extreme data: Identifiability of pedigrees 3. ”Association Mapping” on the Tree of Life 4. Somatic Genealogies and the Models of Embryology

Global Pedigrees 99 Chang and Derrida. Time to a universal common ancestor 04 Rhode tries to answer this for realistic population model • Combining the Coalescent and Pedigree Process • Super-pedigree problem • Bound on how much data is needed to infer a pedigree • Does embedded phylogenies determined the pedigree 1. Wiuf & Hein (1999) 'A contribution to the discussion of J. Chang's paper "Recent Common Ancestor of All Present Human Individuals" ' ( Adv. Appl. Prob. vol. 31. 4) 2. Hein (2004) "Pedigrees for all Humanity" Nature 431. 512 -13. Steel and Hein (2005) “Reconstructing Pedigrees: A combinatorial perspective. J. Theor. Biol.

Combining Ancestral Individuals and the Coalescent Finding Common Ancestors. Wiuf & Hein, 2000. NOW Let T be the time, when somebody was everybody’s ancestor. Changs’ result: lim T*/log 2(N) =1 prob. 1 Unify the two processes: I. Sample more individuals Result: II. Let each have 2 parents with probabilty p. A discontinuity at 1. For p<1 change log 2 logp Comment: Genetic Ancestors is a vanishing set within Genealogical Ancestors.
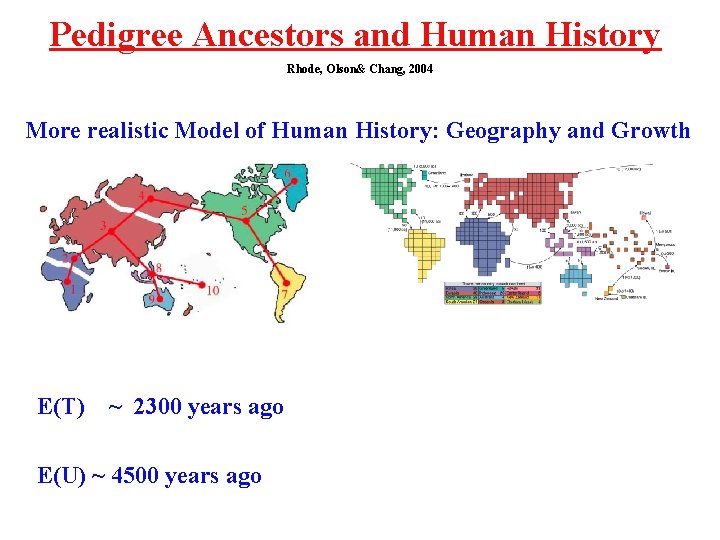
Pedigree Ancestors and Human History Rhode, Olson& Chang, 2004 More realistic Model of Human History: Geography and Growth E(T) ~ 2300 years ago E(U) ~ 4500 years ago
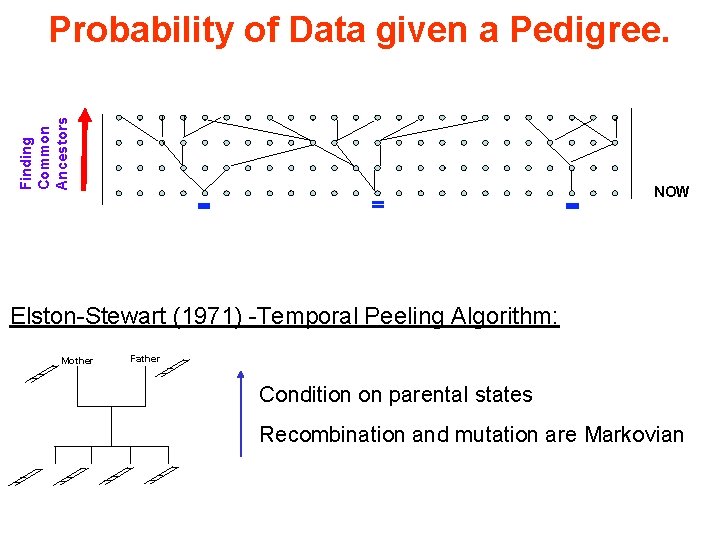
Finding Common Ancestors Probability of Data given a Pedigree. NOW Elston-Stewart (1971) -Temporal Peeling Algorithm: Mother Father Condition on parental states Recombination and mutation are Markovian
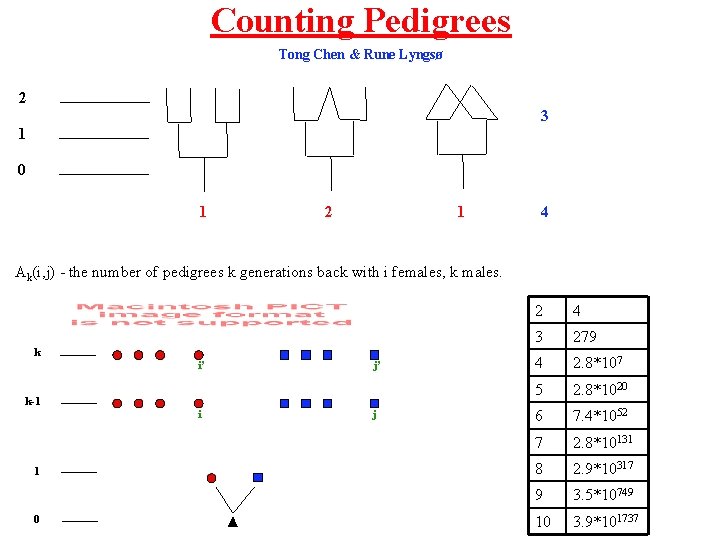
Counting Pedigrees Tong Chen & Rune Lyngsø 2 3 1 0 1 2 1 4 Ak(i, j) - the number of pedigrees k generations back with i females, k males. k i’ j’ k-1 i 1 0 j 2 4 3 279 4 2. 8*107 5 2. 8*1020 6 7. 4*1052 7 2. 8*10131 8 2. 9*10317 9 3. 5*10749 10 3. 9*101737

Pedigree Counting • Counting gender un-labelled pedigrees Much harder. • Counting gender labellings on un-labelled pedigree. gender un-labelable:
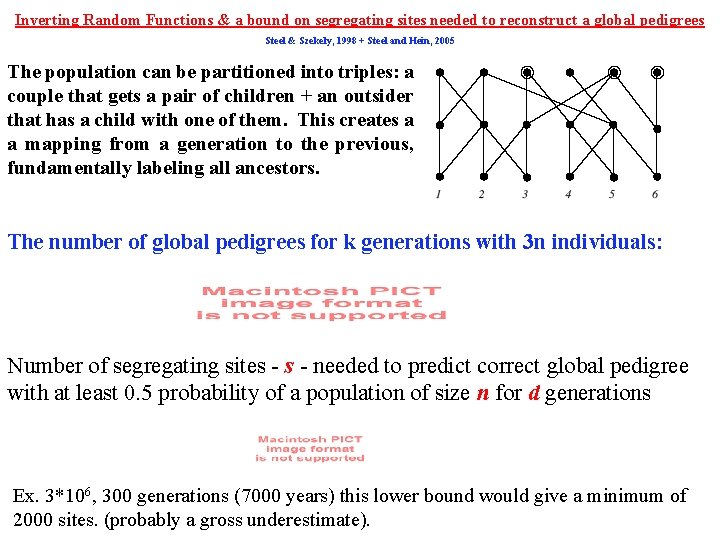
Inverting Random Functions & a bound on segregating sites needed to reconstruct a global pedigrees Steel & Szekely, 1998 + Steel and Hein, 2005 The population can be partitioned into triples: a couple that gets a pair of children + an outsider that has a child with one of them. This creates a a mapping from a generation to the previous, fundamentally labeling all ancestors. The number of global pedigrees for k generations with 3 n individuals: Number of segregating sites - needed to predict correct global pedigree with at least 0. 5 probability of a population of size n for d generations Ex. 3*106, 300 generations (7000 years) this lower bound would give a minimum of 2000 sites. (probably a gross underestimate).
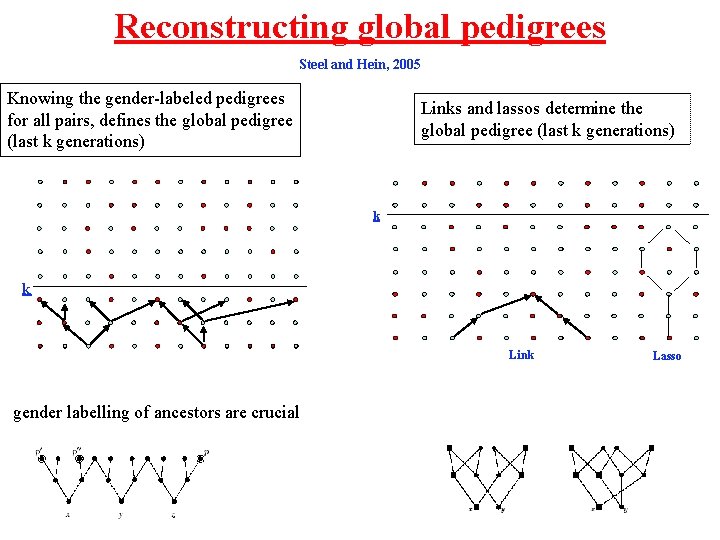
Reconstructing global pedigrees Steel and Hein, 2005 Knowing the gender-labeled pedigrees for all pairs, defines the global pedigree (last k generations) Links and lassos determine the global pedigree (last k generations) k k Link gender labelling of ancestors are crucial Lasso

Benevolent Mutation and Recombination Process Genomes with and m/ --> infinity - recombination rate, m - mutation rate • All embedded phylogenies are observable • Do they determine the pedigree? Counter example: Embedded phylogenies:

Pedigree Reconstruction Principles Distance Based Reconstructions Gender specific rates Continuous Birth Time with Perfect Clock t 3 t 1 t 2 Subtree Transfer Identification of Ancestors Recursive Definition of Ancestral Genomes

The Coalescent with Recombination Retrospective in stead of Prospective formulation of Genetical Processes (Ewens, 1979) 40 s: retrospective arguments used by both Fisher and Wright. 75: Watterson full formulation of probability of genealogical relationship of a set of alleles. 82: Three Famous articles by Kingman. 83: Hudson Includes Recombination in Genealogical Process. • Number of Ancestors to a DNA Sequence. • Reformulation of Genealogical Process. • Inclusion of Gene Conversion in Genealogical Process. 1. 2. 3. 4. Wiuf & Hein (1997): On the Number of Ancestors to a DNA Sequence Wiuf & Hein (1999): The Ancestry of a Sample of Sequences Subject to Recombination Wiuf & Hein (1999): The Coalescent with Recombination as a point process moving along sequences. Wiuf & Hein (2000): The Coalescent with Gene Conversion

Finding Minimal Recombination Histories 64 Bodmer & Edwards: Parsimony defined as reconstruction principle 85 Hudson Kaplan uses minimal recombination histories as observed recombinations • Attempts to find minimal histories of sequences • Definition of recombination as Subtree Prune Regraft operations 1. J. J. Hein: Reconstructing the history of sequences subject to Gene Conversion and Recombination. Mathematical Biosciences. (1990) 98. 185 -200. 2. 3. J. J. Hein: A Heuristic Method to Reconstruct the History of Sequences Subject to Recombination. J. Mol. Evol. 20. 402 -411. 1993 Hein, J. J. , T. Jiang, L. Wang & K. Zhang (1996): "On the complexity of comparing evolutionary trees" Discrete Applied Mathematics 71. 153 -169 Song, Y. S. (2003) “On the combinatorics of rooted binary phylogenetic trees”. Annals of Combinatorics, 7: 365– 379 4. 5. 6. Song, Y. S. & Hein, J. (2005) Constructing Minimal Ancestral Recombination Graphs. J. Comp. Biol. , 12: 147– 169 7. Song, Y. S. & Hein, J. (2003) Parsimonious reconstruction of sequence evolution and haplotype blocks: finding the minimum number of recombination events, Lecture Notes in Bioinformatics, Proceedings of WABI'03, 2812: 287– 302. Lyngsø, Song and Hein (2005) “Minimal Recombination Histories by Branch and Bound” WABI 8. Song, Y. S. & Hein, J. (2004) On the minimum number of recombination events in the evolutionary history of DNA sequences. J. Math. Biol. , 48: 160– 186.

Likelihood of Data Set 72 Ewens likelihood of allele number observations 87 Griffiths recursions for infinite site data 90 Felsenstein uses Metropolis Hastings 94 Griffiths-Tavare uses MCMC on coalescent-mutation process 96 Griffiths-Marjoram uses MCMC on coalescent-mutation-recombination process 99+ Donnelly-Matthews-Fearnhead uses IS to accellerate earlier methods 00 Hudson introduces pseuodolikelihood method • How hard is the coalescent-mutation-recombination process? 1. Song, Y. S. , Lyngsø, R. B. & Hein, J. (2005) Counting Ancestral States in Population Genetics. In Press
- Slides: 50