Department of Microbiology Faculty of Biology Geography and

Department of Microbiology Faculty of Biology, Geography and Oceanology University of Gdańsk www. microbiology. univ. gda. pl e-mail: kaczorow@biotech. univ. gda. pl

Main fields of interest: 1. Biology of bacterial DNA restriction and modification systems. 2. New approaches for DNA sequencing and analysis.
Restriction-modification systems. What are they for?
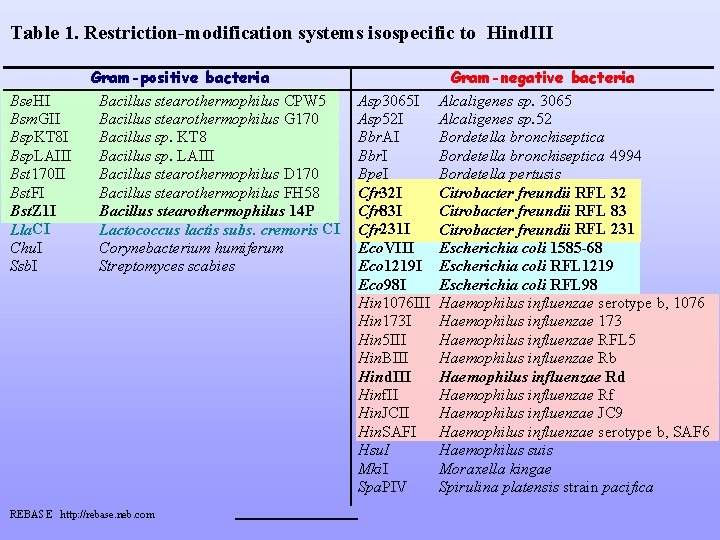
Table 1. Restriction-modification systems isospecific to Hind. III Bse. HI Bsm. GII Bsp. KT 8 I Bsp. LAIII Bst 170 II Bst. FI Bst. Z 1 I Lla. CI Chu. I Ssb. I Gram-positive bacteria Bacillus stearothermophilus CPW 5 Bacillus stearothermophilus G 170 Bacillus sp. KT 8 Bacillus sp. LAIII Bacillus stearothermophilus D 170 Bacillus stearothermophilus FH 58 Bacillus stearothermophilus 14 P Lactococcus lactis subs. cremoris CI Corynebacterium humiferum Streptomyces scabies REBASE http: //rebase. neb. com Gram-negative bacteria Asp 3065 I Alcaligenes sp. 3065 Asp 52 I Alcaligenes sp. 52 Bbr. AI Bordetella bronchiseptica Bbr. I Bordetella bronchiseptica 4994 Bpe. I Bordetella pertusis Cfr 32 I Citrobacter freundii RFL 32 Cfr 83 I Citrobacter freundii RFL 83 Cfr 231 I Citrobacter freundii RFL 231 Eco. VIII Escherichia coli 1585 -68 Eco 1219 I Escherichia coli RFL 1219 Eco 98 I Escherichia coli RFL 98 Hin 1076 III Haemophilus influenzae serotype b, 1076 Hin 173 I Haemophilus influenzae 173 Hin 5 III Haemophilus influenzae RFL 5 Hin. BIII Haemophilus influenzae Rb Hind. III Haemophilus influenzae Rd Hinf. II Haemophilus influenzae Rf Hin. JCII Haemophilus influenzae JC 9 Hin. SAFI Haemophilus influenzae serotype b, SAF 6 Hsu. I Haemophilus suis Mki. I Moraxella kingae Spa. PIV Spirulina platensis strain pacifica
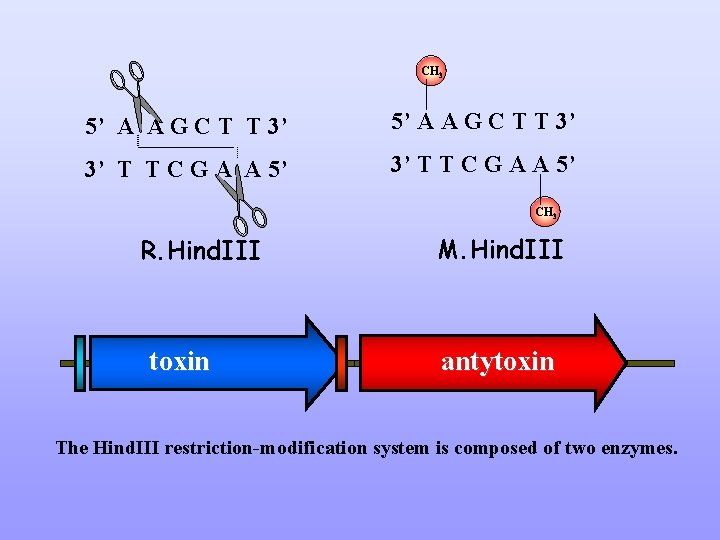
CH 3 5’ A A G C T T 3’ 3’ T T C G A A 5’ CH 3 R. Hind. III toxin M. Hind. III antytoxin The Hind. III restriction-modification system is composed of two enzymes.

Objectives: 1. how similar are genes encoding isospecific enzymes ? 2. is it possible to map their functional domains ? 3. do they recognize cognate sequence in the same way ? 4. what is their mode of action ?
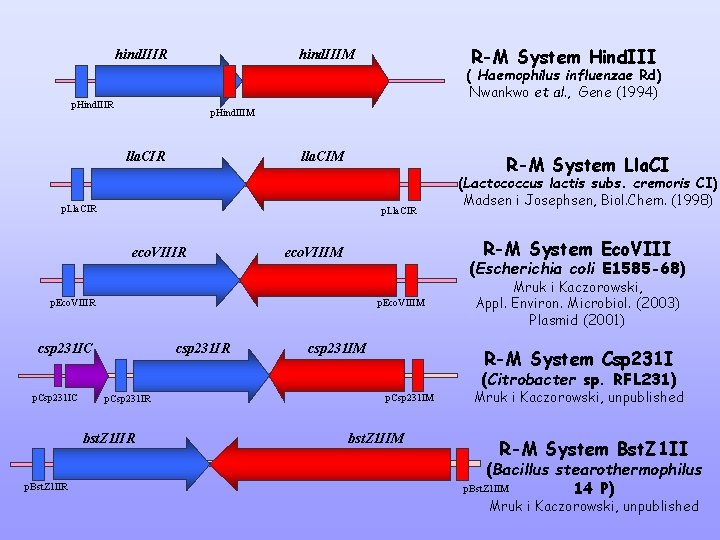
hind. IIIR R-M System Hind. III hind. IIIM ( Haemophilus influenzae Rd) Nwankwo et al. , Gene (1994) p. Hind. IIIR p. Hind. IIIM lla. CIR lla. CIM R-M System Lla. CI p. Lla. CIR eco. VIIIR R-M System Eco. VIII eco. VIIIM (Escherichia coli E 1585 -68) p. Eco. VIIIR p. Eco. VIIIM csp 231 IC csp 231 IR (Lactococcus lactis subs. cremoris CI) Madsen i Josephsen, Biol. Chem. (1998) csp 231 IM Mruk i Kaczorowski, Appl. Environ. Microbiol. (2003) Plasmid (2001) R-M System Csp 231 I (Citrobacter sp. RFL 231) p. Csp 231 IC p. Csp 231 IR bst. Z 1 IIR p. Bst. Z 1 IIR p. Csp 231 IM bst. Z 1 IIM Mruk i Kaczorowski, unpublished R-M System Bst. Z 1 II (Bacillus stearothermophilus p. Bst. Z 1 IIM 14 P) Mruk i Kaczorowski, unpublished
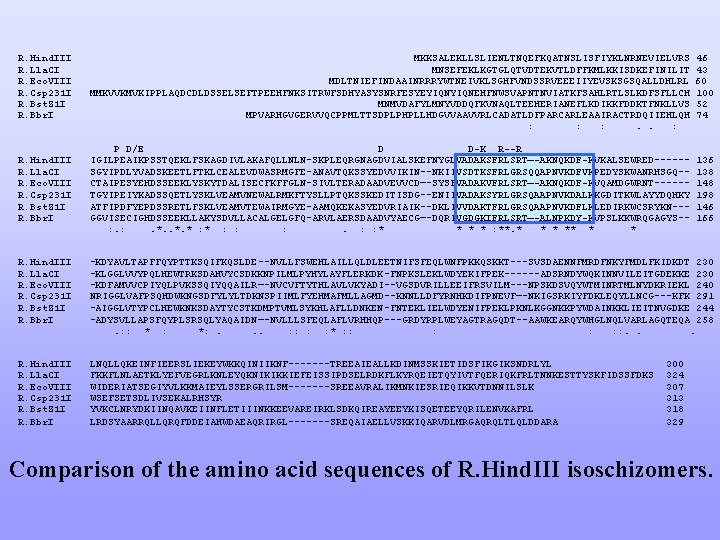
R. Hind. III R. Lla. CI R. Eco. VIII R. Csp 231 I R. Bst. Z 1 I R. Bbr. I MKKSALEKLLSLIENLTNQEFKQATNSLISFIYKLNRNEVIELVRS MNSEFEKLKGTGLQTVDTEKVTLDFFKMLKKISDKEFINILIT MDLTNIEFINDAAINRRRYWTNEIVKLSGHFVNDSSRVEEEIIYEVSKSGSQALLDHLRL MMKVVKMVKIPPLAQDCDLDSSELSEFTPEEHFNKSITRWFSDHYASYSNRFESYEYIQNEHFNWSVAPNTNVIATKFSAHLRTLSLKDFSFLLCH MNMVDAFYLMNYVDDQFKVNAQLTEEHERIANEFLKDIKKFDDKTFNKLLVS MPVARHGVGERVVQCPPMLTTSDPLPHPLLHDGVVAAVVRLCADATLDFPARCARLEAAIRACTRDQIIEHLQH : : : . . : P D/E D D-K R--R IGILPEAIKPSSTQEKLFSKAGDIVLAKAFQLLNLN-SKPLEQRGNAGDVIALSKEFNYGLVADAKSFRLSRT—-AKNQKDF-KVKALSEWRED-----SGYIPDLYVADSKEETLFTKLCEALEVDWASRMGFE-ANAVTQKSSYEDVVIKIN--NKIIVSDTKSFRLGRSQQAPNVKDFVKPEDYSKWANRHSGQ-CTAIPESYEHDSSEEKLYSKYTDALISECFKFFGLN-SIVLTERADAADVEVVCD—-SYSFVADAKVFRLSRT—-AKNQKDF-KVQAMDGWRNT-----TGYIPEIYKADSSQETLYSKLVEAMVNEWALRMKFTYSLLPTQKSSKEDITISDG--ENIIVADAKSYRLGRSQAAPNVKDALKKGDITKWLAYYDQHKY ATFIPDFYEPDSSRETLFSKLVEAMVTEWAIRMGYE-AAMQKEKASYEDVRIAIK--DKLIVVDAKTFRLGRSQAAPNVKDFLKLEDIRKWCSRYKN--GGVISECIGHDSSEEKLLAKYSDVLLACALGELGFQ-ARVLAERSDAADVYAECG—-DQRIVGDGKIFRLSRT—-ALNPKDY-KVPSLKKWRQGAGYS-: . *. * : : : * * : **. * ** * * 46 43 60 100 52 74 136 138 148 198 146 166 R. Hind. III R. Lla. CI R. Eco. VIII R. Csp 231 I R. Bst. Z 1 I R. Bbr. I -KDYAVLTAPFFQYPTTKSQIFKQSLDE--NVLLFSWEHLAILLQLDLEETNIFSFEQLWNFPKKQSKKT---SVSDAENNFMRDFNKYFMDLFKIDKDT 230 -KLGGLVVYPQLHEWTRKSDAHVYCSDKKNPILMLPYHYLAYFLERKDK-FNPKSLEKLWDYEKIFPEK------ADSRNDYWQKINNVILEITGDEKKE 230 -KDFAMVVCPIYQLPVKSSQIYQQAILR—-NVCVFTYTHLAVLVKYADI--VGSDVRILLEEIFRSVILM---NPSKDSVQYWTMINRTMLNYDKRIEKL 240 NRIGGLVAFPSQHDWKNGSDFYLYLTDKNSPIIMLFYEHMAFMLLAGMD--KNNLLDFYRNHKDIFPNEVF—-NKIGSRKIYFDKLEQYLLNCG---KFK 291 -AIGGLVTYPCLHEWKNKSDAYTYCSTKDMPTVMLSYKHLAFLLDNKEN-FNTEKLIELWDYENIFPEKLPKNLKGGNKKPYWDAINKKLIEITNVGDKE 244 -ADYSVLLAPSFQYPLSRSQLYAQAIDN—-NVLLLSFEQLAFLVRHHQP---GRDYRPLWEYAGTRAGQDT--AAWKEARQYWHGLNQLVARLAGQTEQA 258. : : *: . . . : : * : : : . . . R. Hind. III R. Lla. CI R. Eco. VIII R. Csp 231 I R. Bst. Z 1 I R. Bbr. I LNQLLQKEINFIEERSLIEKEYWKKQINIIKNF-------TREEAIEALLKDINMSSKIETIDSFIKGIKSNDRLYL FKKFLNLAETKLYEFVEGRLKNLEYQKNIKIKKIEFEISSIPDSELRDKFLKYRQEIETQYIVTFQERIQKFRLTNNKESTTYSKFIDSSFDKS WIDERIATSEGIYVLKKMAIEYLSSERGRILSM-------SREEAVRALIKMNKIESRIEQIKKVTDNNILSLK WSEFSETSDLIVSEKALRHSYR YVKCLNRYDKIINQAVKEIINFLETIIINKKEEVAREIRKLSDKQIREAYEEYKISQETEEYQRILENVKAFRL LRDSYAARRQLLQRQFDDEIAHWDAEAQRIRGL-------SREQAIAELLVSKKIQARVDLMRGAQRQLTLQLDDARA 300 324 307 313 318 329 Comparison of the amino acid sequences of R. Hind. III isoschizomers.
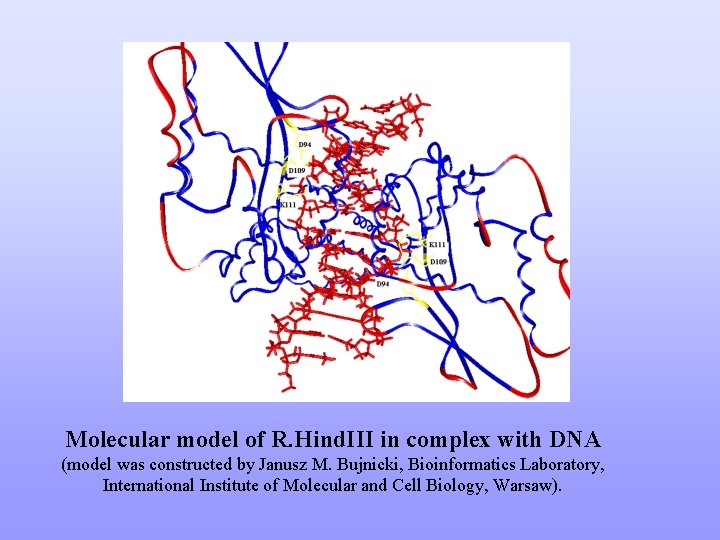
Molecular model of R. Hind. III in complex with DNA (model was constructed by Janusz M. Bujnicki, Bioinformatics Laboratory, International Institute of Molecular and Cell Biology, Warsaw).
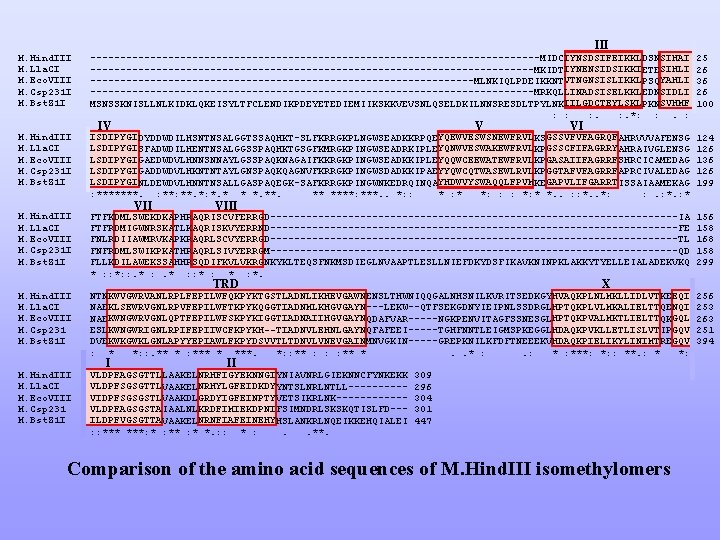
III M. Hind. III M. Lla. CI M. Eco. VIII M. Csp 231 I M. Bst. Z 1 I --------------------------------------MIDCIYNSDSIFEIKKLDSNSIHAI -------------------------------------MKIDTIYNENSIDSIKKIETESIHLI --------------------------------MLNKIQLPDEIKKNTVTNGNSISLIKKLPSQYAHLI -------------------------------------MRKQLLINADSISELKKLEDNSIDLI MSNSSKNISLLNLKIDKLQKEISYLTFCLENDIKPDEYETEDIEMIIKSKKVEVSNLQSELDKILNNSRESDLTPYLNKIILGDCTEYLSKLPKNSVHMF : : : . *: : . : 25 26 36 26 100 M. Hind. III M. Lla. CI M. Eco. VIII M. Csp 231 I M. Bst. Z 1 I ISDIPYGIDYDDWDILHSNTNSALGGTSSAQHKT-SLFKRRGKPLNGWSEADKKRPQEYQEWVESWSNEWFRVLKSGSSVFVFAGRQFAHRVVVAFENSG LSDIPYGISFADWDILHENTNSALGGSSPAQHKTGSGFKMRGKPINGWSEADRKIPLEYQNWVESWAKEWFRVLKPGSSCFIFAGRRYAHRAIVGLENSG LSDIPYGIGAEDWDVLHNNSNNAYLGSSPAQKNAGAIFKKRGKPINGWSEADKKIPLEYQQWCEEWATEWFRVLKPGASAIIFAGRRFSHRCICAMEDAG LSDIPYGIGADDWDVLHKNTNTAYLGNSPAQKQAGNVFKRRGKPINGWSDADKKIPAEYYQWCQTWASEWLRVLKPGGTAFVFAGRRFAPRCIVALEDAG LSDIPYGINLDEWDVLHNNTNSALLGASPAQEGK-SAFKRRGKPINGWNKEDRQINQAYHDWVYSWAQQLFPVMKEGAPVLIFGARRTISSAIAAMEKAG : *******. : **. *: *. * * *. ** ****: ***. . *: : * *: : : *: * *. . : : *. . *: : . : * 124 126 136 126 199 M. Hind. III M. Lla. CI M. Eco. VIII M. Csp 231 I M. Bst. Z 1 I FTFKDMLSWEKDKAPHRAQRISCVFERRGD----------------------------------IA FTFRDMIGWNRSKATLKAQRISKVYERRND----------------------------------FE FNLRDIIAWMRVKAPKRAQRLSCVYERRGD----------------------------------TL FNFRDMLSWIKPKATHRAQRLSIVYERRGM----------------------------------QD FLLKDILAWEKSSAHHRSQDIFKVLVKRGNKYKLTEQSFNKMSDIEGLNVAAPTLESLLNIEFDKYDSFIKAVKNINPKLAKKYTYELLEIALADEKVKQ * : : *: : . * : : * : *. 156 158 168 158 299 M. Hind. III M. Lla. CI M. Eco. VIII M. Csp 231 M. Bst. Z 1 I NTNKWVGWRVANLRPLFEPILWFQKPYKTGSTLADNLIKHEVGAWNENSLTHWNIQQGALNHSNILKVRITSEDKGYHVAQKPLNLMKLLIDLVTKEEQI NAEKLSEWRVGNLRPVFEPILWFTKPYKQGGTIADNMLKHGVGAYN---LEKW--QTFSEKGDNYIEIPNLSSDRGLHPTQKPLVLMKALIELTTQENQI NAEKWNGWRVGNLQPTFEPILWFSKPYKIGGTIADNAIIHGVGAYNQDAFVAR-----NGKPENVITAGFSSNESGLHPTQKPVALMKTLIELTTQKGQL ESLKWNGWRIGNLRPIFEPIIWCFKPYKH--TIADNVLEHNLGAYNQFAFEEI-----TGHFNNTLEIGMSPKEGGLHDAQKPVKLLETLISLVTIPGQV DVEKWKGWKLGNLAPYYEPIAWLFKPYDSVVTLTDNVLVNEVGAINMNVGKIN-----GREPKNILKFDFTNEEEKVHDAQKPIELIKYLINIMTREGQV : * *: : . ** * : *** * ***. *: : ** : : : ** *. . * : . : ***: *: : **. : * *: 256 253 263 251 394 M. Hind. III M. Lla. CI M. Eco. VIII M. Csp 231 M. Bst. Z 1 I VLDPFAGSGTTLLAAKELNRHFIGYEKNNGIYNIAVNRLGIEKNNCFYNKEKK VLDPFSGSGTTLVAAKELNRHYLGFEIDKDYYNTSLNRLNTLL-----VIDPFSGSGSTLVAAKDLGRDYIGFEINPTYVETSIKRLNK------VLDPFAGSGSTAIAALNLKRDFIMIEKDPNIFSIMNDRLSKSKQTISLFD--ILDPFVGSGTTAVAAKELNRNFIAFEINEHYHSLANKRLNQEIKKEHQIALEI : : ***: * : * *. : : * : . . **. IV V VIII TRD I VI X II 309 296 304 301 447 Comparison of the amino acid sequences of M. Hind. III isomethylomers

M. Hind. III R. Hind. III MIDCIYN--SDSIXEIKKLDSNSIHAIISDIPYGIDYDDWDILHSNTNSALGGTSSAQHK 58 MKKSALEKLLSLIENLTNQEFKQATNSLISFIYKLNRNEVIELVRSIGILPEAIKPSSTQ 60 *. . : . * : : . : . : * : : *. . . : M. Hind. III R. Hind. III TSLFKRRGKPLNGWSEADKKRPQEYQEWVESWSNEWFRVLKSGSSVFVFAGRQFAHRVVV 118 EKLFSKAGDIV--LAKAFQLLNLNSKPLEQRGNAGDVIALSKEFNYGLVADAKSFRLSRT 118. **. : : : * : : . . . *. . . : . *. : : . M. Hind. III R. Hind. III AFENSGFTFKDMLSWEKDKAPHRAQRISCVFERRGDIANTNKWVGWRVANLRPLFEPILW 178 AKNQKDFKVKALSEWREDKD-YAVLTAPFFQYPTTKSQIFKQSLDENVLLFSWEHLAILL 177 * : : . . * : . *. : ** : . . : : : . . ** M. Hind. III R. Hind. III FQKPYKTGSTLADNLIKHEVGAWNENSLTHWNIQQGALNHSNILKVRITSEDKGYHVAQK 238 QLDLEETNIFXFEQLWNFPKKQSKKTSVSDAENNFMRDFNKYFMDLFKIDKDTLNQLLQK 237. : *. : : * : . : : . *: : . : *. : : ** M. Hind. III R. Hind. III PLNLMK---LLIDLVTKEEQIVLDPFAGSGTTLLAAKELNRHFIGYEKNNGIYNIAVNRL 295 EINFIEERSLIEKEYWKKQINIIKNFTREEAIEALLKDIN-------MSSKIETIDSFIK 290 : *: : : *: . : *: : *. * M. Hind. III R. Hind. III GIEKNNCFYNKEKK 309 GIKSNDRLYL---- 300 **: . *: : * Comparison of the amino acid sequences of M. Hind. III and R. Hind. III.
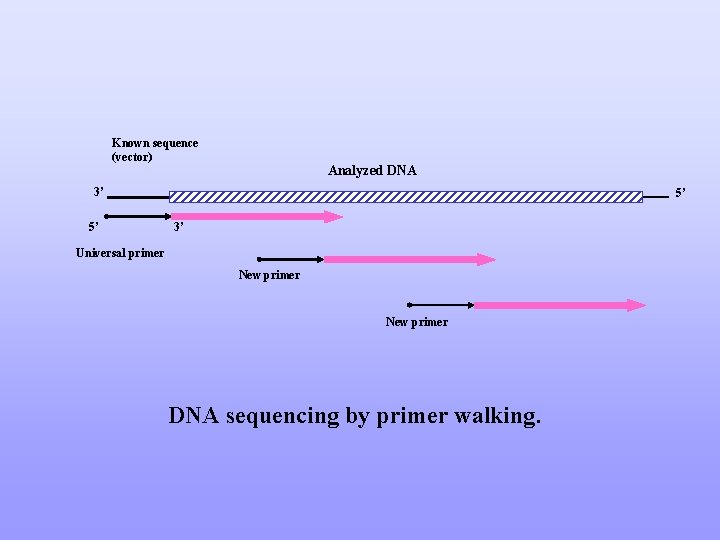
Known sequence (vector) Analyzed DNA 3’ 5’ 5’ 3’ Universal primer New primer DNA sequencing by primer walking.
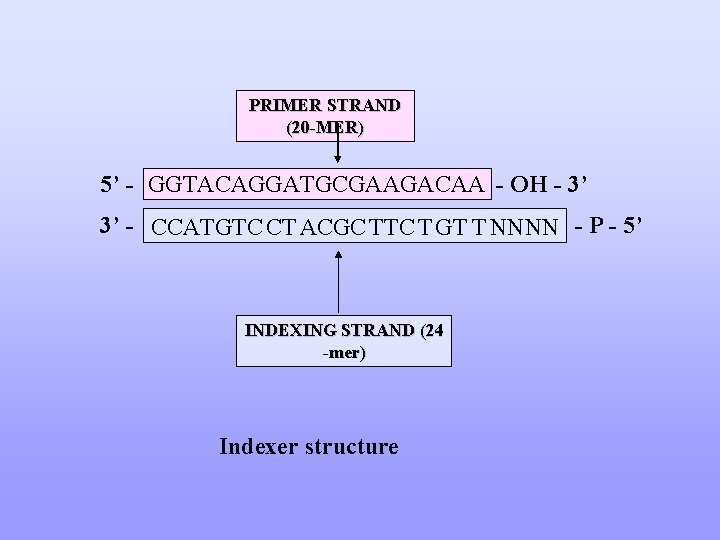
PRIMER STRAND (20 -MER) 5’ - GGTACAGGATGCGAAGACAA - OH - 3’ 3’ - CCATGTC CT ACGC TTC T GT T NNNN - P - 5’ INDEXING STRAND (24 -mer) Indexer structure
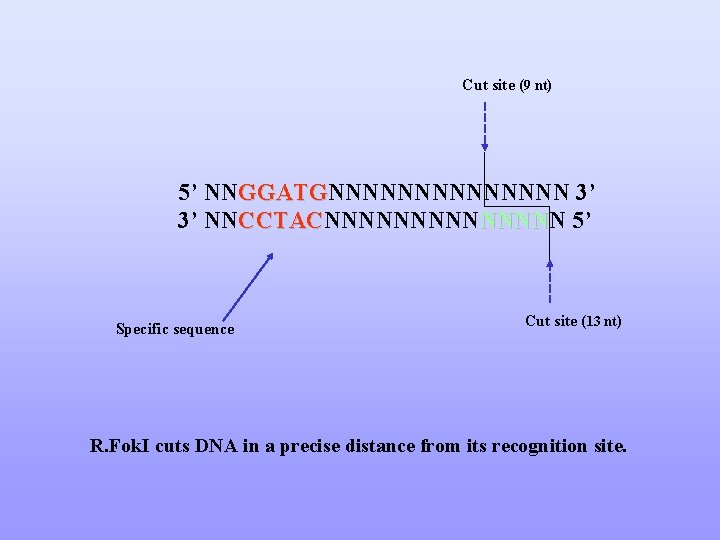
Cut site (9 nt) 5’ NNGGATGNNNNNNN 3’ GGATG 3’ NNCCTACNNNNN CCTAC NNNN 5’ Specific sequence Cut site (13 nt) R. Fok. I cuts DNA in a precise distance from its recognition site.

DNA sequencing by indexer walking Type IIS restriction endonucleases Enzyme Specific sequence Average fragment length (bp) Commercially available Aar. I Alw 26 I (Bsm. AI) Bbs. I (Bpi. I) Bbv. I (Bse. XI) Bsm. FI Bsp. MI Btg. ZI Eco 31 I Esp 3 I Fok. I Sfa. NI CACCTGC (4/8) GTCTC (1/5) GAAGAC (2/6) GCAGC (8/12) GGGAC (10/14) ACCTGC (4/8) GCGATG (10/14) GGTCTC (1/5) CGTCTC (1/5) GGATG (9/13) GCATC (5/9) 8192 512 2048 2048 512 Commercially unavailable Ace. III Bbr 7 I Bbv. II Sth 132 I Sts. I (www. rebase. neb. com/rebase) CAGCTC (7/11) GAAGAC (2/6) CCCG (4/8) GGATG (10/14) 2048 128 512
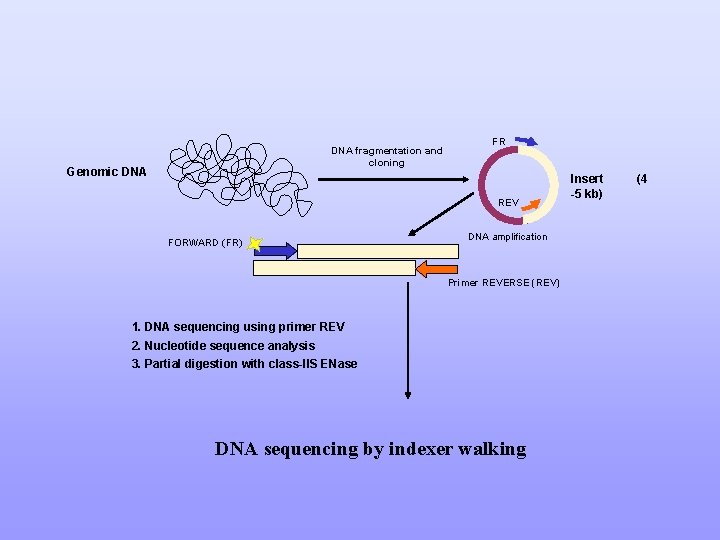
DNA fragmentation and cloning Genomic DNA FR REV FORWARD (FR) DNA amplification Primer REVERSE (REV) 1. DNA sequencing using primer REV 2. Nucleotide sequence analysis 3. Partial digestion with class-IIS ENase DNA sequencing by indexer walking Insert -5 kb) (4
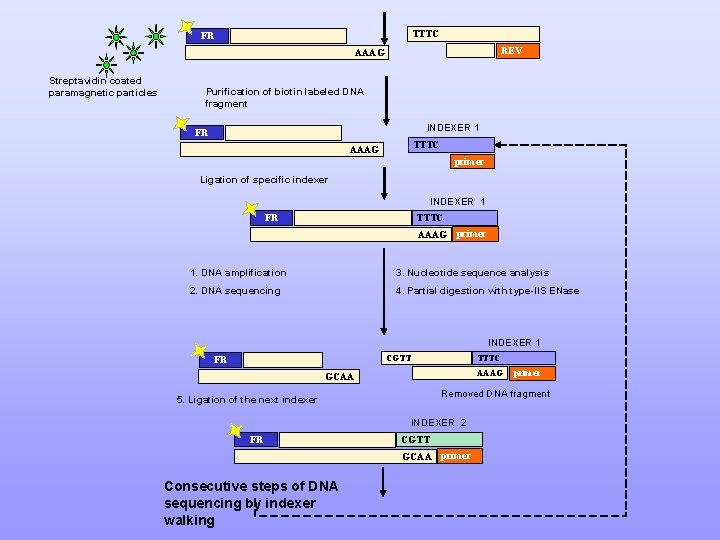
TTTC FR REV AAAG Streptavidin coated paramagnetic particles Purification of biotin labeled DNA fragment INDEXER 1 FR TTTC AAAG primer Ligation of specific indexer INDEXER 1 FR TTTC AAAG primer 1. DNA amplification 3. Nucleotide sequence analysis 2. DNA sequencing 4. Partial digestion with type-IIS ENase INDEXER 1 TTTC CGTT FR AAAG GCAA Removed DNA fragment 5. Ligation of the next indexer INDEXER 2 FR CGTT GCAA primer Consecutive steps of DNA sequencing by indexer walking primer


Financial support: Polish Committee for Scientific Research KBN 6 P 04 B-027 -18 2000 -2003 KBN 6 P 04 B-003 -12 1997 -2000 KBN IA-1433/99 1999 KBN IA-0589/2001

Lab members: Beata Furmanek-Blaszk Magdalena Cichowicz Anna Kaczorowska Agnieszka Dekowska Tadeusz Kaczorowski Katarzyna Gromek Iwona Mruk Anna Kossobucka Józef Nieradko Marian Sęktas Łucja Adamczyk Jadwiga Pomorska
- Slides: 20