Transport PDEs Practice Sauro Succi Problem set up

Transport PDE’s: Practice Sauro Succi

Problem set up 1 d Advection-Diffusion-Reaction Equation Spacetime domain: Initial conditions: Gaussian Boundary conditions: Periodic
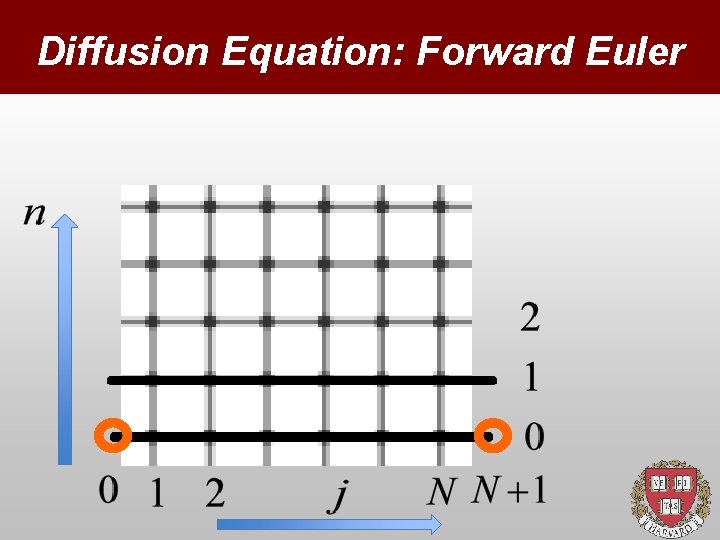
Diffusion Equation: Forward Euler
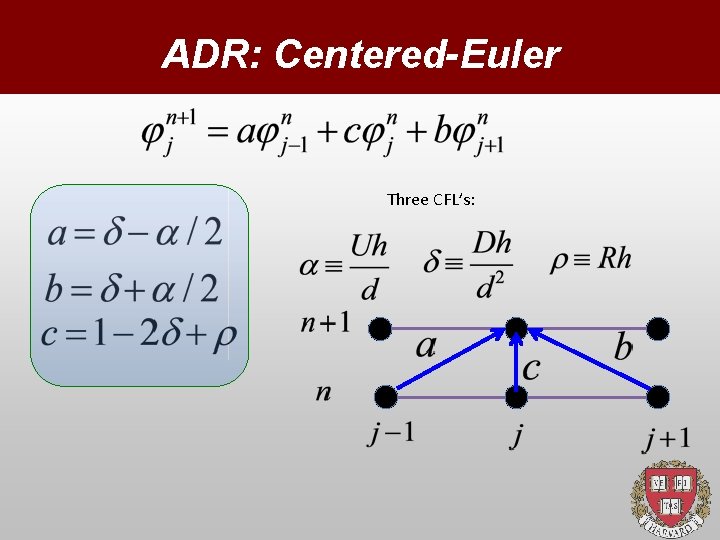
ADR: Centered-Euler Three CFL’s:

Discretization Space: Time: CFL conditions

Diagnostics-Conservations Mass conservation: Total Mass Integrating by parts: We should use as much analytical infortmation as possible for VALIDATION purposes. Guard against Computer HALLUCINATIONS! With zero-flux or periodic b. c. Analytical solution: Average position: Variance: Entropy:

Diagnostics Check against analytical solution Mass Entropy-like: monotonic in time, gives trend to steady state

Convergence If the method is convergent Define the departure between solutions with N and M gridpoints respectively: where and “dis” stands for a suitable distance In N-dimensional space

Convergence If the method is convergent to order p: Coarse grid: Fine grid: Since 3 in the C-grid is the same point as 5 in the F-grid, we have:

Richardson extrapolation If the method is convergent the value at each given space location should become independent of the grid resolution (Richardson extrapolation) as In practice, one fixes a tolerance parameter TOL, and simply asks that
Codlet adr 1. f
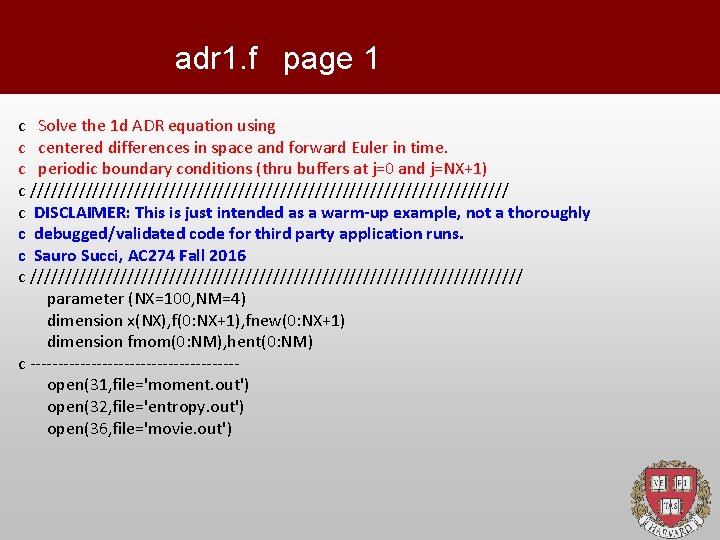
adr 1. f page 1 c Solve the 1 d ADR equation using c centered differences in space and forward Euler in time. c periodic boundary conditions (thru buffers at j=0 and j=NX+1) c /////////////////////////////////// c DISCLAIMER: This is just intended as a warm-up example, not a thoroughly c debugged/validated code for third party application runs. c Sauro Succi, AC 274 Fall 2016 c //////////////////////////////////// parameter (NX=100, NM=4) dimension x(NX), f(0: NX+1), fnew(0: NX+1) dimension fmom(0: NM), hent(0: NM) c -------------------open(31, file='moment. out') open(32, file='entropy. out') open(36, file='movie. out')

adr 1. f page 2 c baseline parameters and input data Nsteps = 1000 sizex = 1. 0 dif = 1. 0 vel = 10. 0 chr = 0. 0 cap = 0. 0 c chemical reaction is df/dt = chr*f*(1 -cap*f) c by setting cap=0 we recover linear growth/decay dx = sizex/float(nx-1) c CFL numbers fix the timestep cfl = 0. 5 dtd = 1. 0 if(dif. ne. 0. 0) dtd = dx*dx/dif dta = 1. 0 if(vel. ne. 0. 0) dta = dx/vel dtc = 1. 0 if(chr. ne. 0. 0) dtc = 1. 0/chr

adr 1. f page 3 c pick up the safest dt (play with them) dt = cfl*min(dta, dtd, dtc) write(6, *) dx, dtd, dta, dtc, dt alfa = vel*dt/dx delta = dif*dt/(dx*dx) chi = chr*dt c initial conditions: unnormalized (play with them) wid = 4 sigma = wid*dx ! play with width do j=1, NX x(j) = dx*float(j-1) xj = (x(j)-sizex/2)/sigma f(j) = exp(-0. 5*xj*xj) end do

adr 1. f page 4 c time-stepper ----------------do it=1, Nsteps f(0) = f(nx) f(nx+1) = f(1) do j=1, NX c build the tridiagonal coefficients a = delta+0. 5*alfa b = delta-0. 5*alfa c = 1. 0 -a-b+chi*(1. 0 -cap*f(j)) fnew(j) = a*f(j-1)+c*f(j)+b*f(j+1) end do c prepare for new timestep do j=1, NX f(j) = fnew(j) end do

adr 1. f page 5 c diagnostics: standard moments, tail moment and entropies if(mod(it, 10). eq. 1) then do m=0, NM fmom(m) = 0. 0 hent(m) = 0. 0 end do xmom = 0. 0 do j=1, NX fj = f(j)*dx fmom(0) = fmom(0) + fj fmom(1) = fmom(1) + fj*x(j) fmom(2) = fmom(2) + fj*x(j) hent(2) = hent(2) - fj*fj hent(3) = hent(3) - fj*fj*fj hent(4) = hent(4) - fj*fj*fj*fj

adr 1. f page 6 c probe the tail, should grow exponentially in time if(x(j). gt. 0. 99*sizex) xmom=xmom+fj c output for gnuplot movies write(36, *) j, f(j) end do c blank lines for gnuplot movies write(36, '(bn)') c monitor the moments in time fmean = fmom(1)/fmom(0) fvar = fmom(2)/fmom(0)-fmean*fmean write(31, *), it, fmom(0), fmean, fvar, xmom/fmom(0) write(32, *), it, hent(2), hent(3), hent(4) endif write(6, *) 'completed time step', it c end evolution --------------------------end do stop end
adr 1. f page 7 Run and Enjoy! Actual codlet: /PART 1 Codes/ADR/adr 1. f
- Slides: 18