Molecular Dynamics Sauro Succi History Molecular Dynamics MD














![Force Calculation: Direct Summation for (i=1; i<=N-1; i++) { F[i]=0. 0; for (j=i+1, j<=N, Force Calculation: Direct Summation for (i=1; i<=N-1; i++) { F[i]=0. 0; for (j=i+1, j<=N,](https://slidetodoc.com/presentation_image_h/283ab853f659d9e6aeb5ccf6700d653d/image-24.jpg)
![Force Calculation: short range for (i=1; i<=N; i++) { F[i]=0. 0; for (j=i+1, j<=N, Force Calculation: short range for (i=1; i<=N; i++) { F[i]=0. 0; for (j=i+1, j<=N,](https://slidetodoc.com/presentation_image_h/283ab853f659d9e6aeb5ccf6700d653d/image-25.jpg)
![Force Calculation: Linked List for (i=1; i<=N; i++) { g=floor(x[i]/rc); for (k=1, k<=npc[g], k++{ Force Calculation: Linked List for (i=1; i<=N; i++) { g=floor(x[i]/rc); for (k=1, k<=npc[g], k++{](https://slidetodoc.com/presentation_image_h/283ab853f659d9e6aeb5ccf6700d653d/image-27.jpg)















- Slides: 56

Molecular Dynamics Sauro Succi

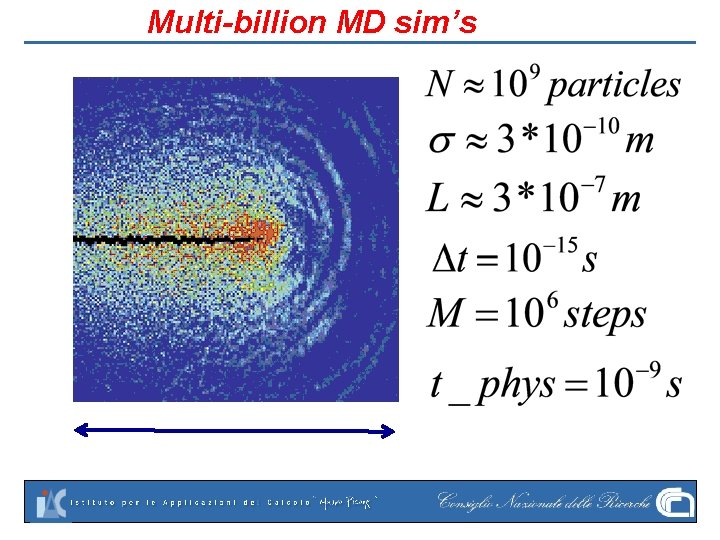
History Molecular Dynamics (MD) began in the mid 50’s with the famous experiments of Alder-Wainwright. They showed violation of Boltzmann’s Molecular Chaos assumption using hundreds of rigid disks. They went on by computing a phase transition in this “virtual-fluid”. Incredible growth ever since: nowadays Multibillion sims can be performed. Still very short of real—life scales, especially in time.

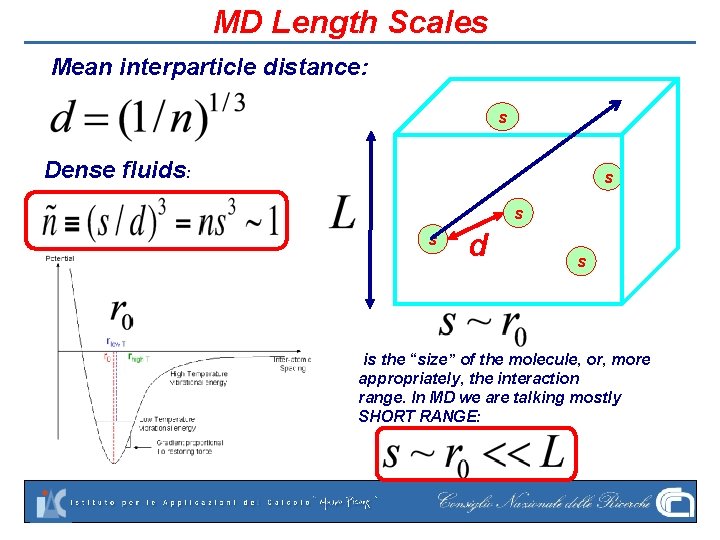
MD Length Scales Mean interparticle distance: s Dense fluids: s s s d s is the “size” of the molecule, or, more appropriately, the interaction range. In MD we are talking mostly SHORT RANGE:

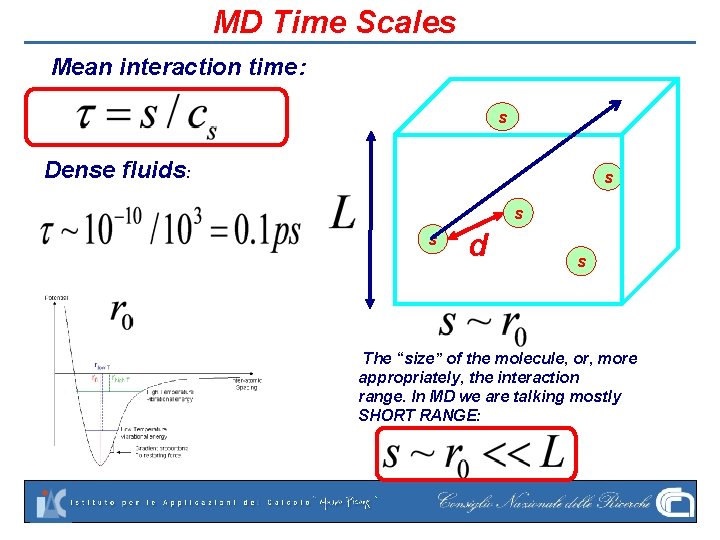
MD Time Scales Mean interaction time: s Dense fluids: s s s d s The “size” of the molecule, or, more appropriately, the interaction range. In MD we are talking mostly SHORT RANGE:

MD Energy Scales Typical energy in thermal units: 6 -12 Lennard-Jones potential s Standard conditions, T=300 K

Classical N-body problem Newton equations for Avogadro’s number of molecules… Potential forces 2 -body 3 -body

Central/Directional Forces Electrostatic forces are aligned (central)

Third Law: Momentum Conservation This implies global momentum conservation:

Energy Conservation

Molecular Dynamics Distinguished Potentials

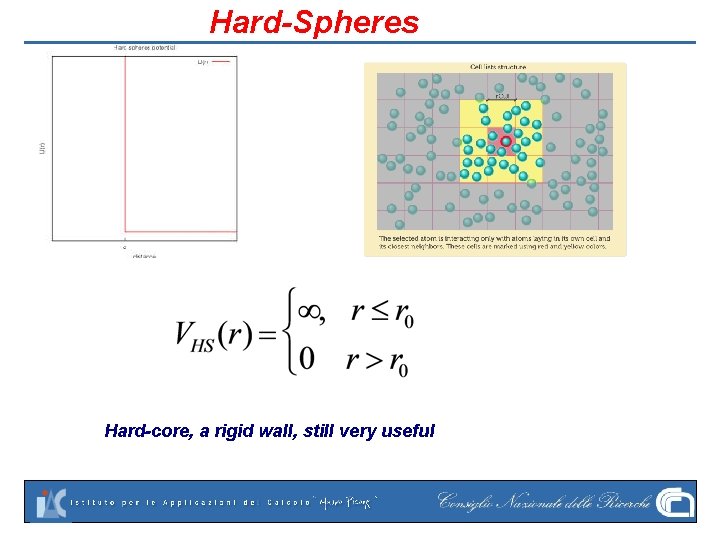
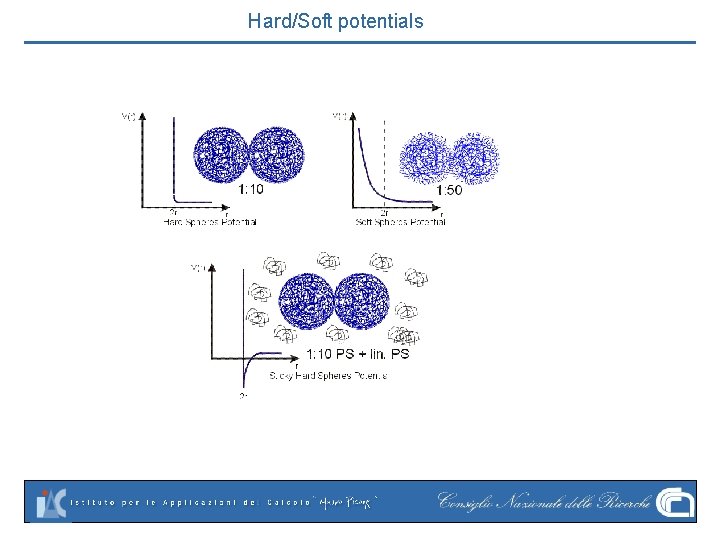
Hard-Spheres Hard-core, a rigid wall, still very useful

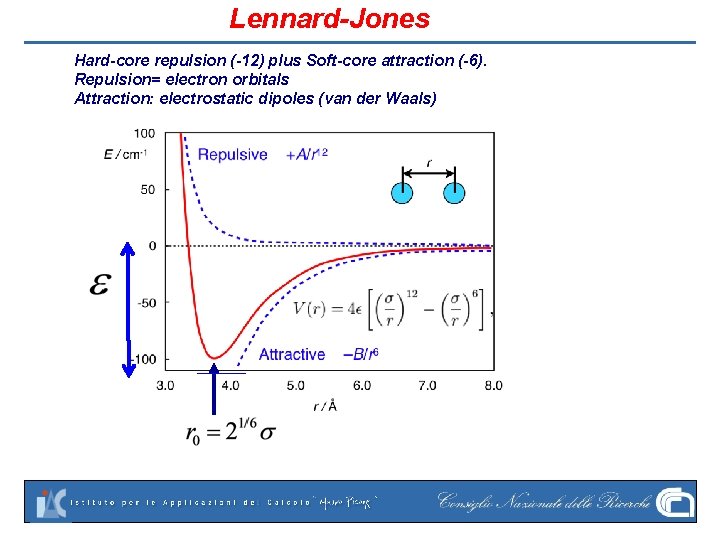
Lennard-Jones Hard-core repulsion (-12) plus Soft-core attraction (-6). Repulsion= electron orbitals Attraction: electrostatic dipoles (van der Waals)

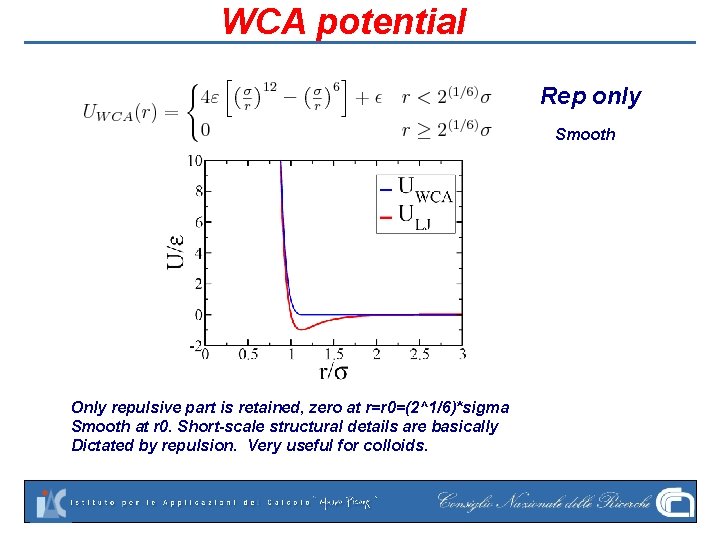
WCA potential Rep only Smooth Only repulsive part is retained, zero at r=r 0=(2^1/6)*sigma Smooth at r 0. Short-scale structural details are basically Dictated by repulsion. Very useful for colloids.

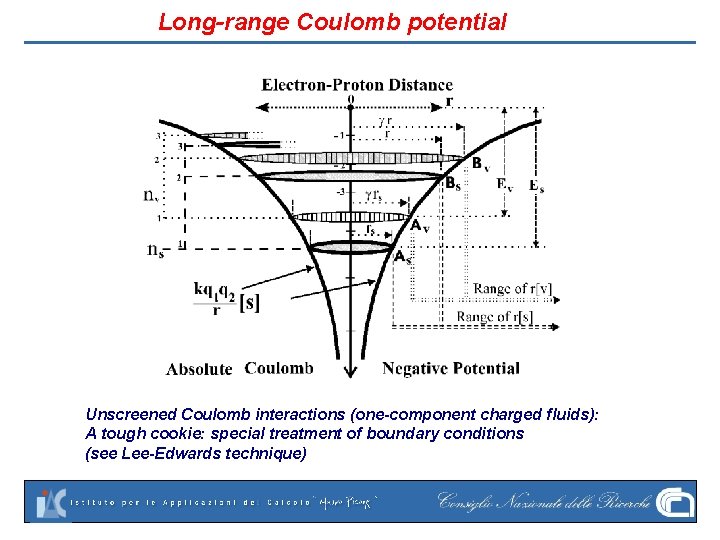
Long-range Coulomb potential Unscreened Coulomb interactions (one-component charged fluids): A tough cookie: special treatment of boundary conditions (see Lee-Edwards technique)

Running the MD simulation 1. Initialization 2. Time integration 3. Data Analysis

Initial Configuration: Positions Particles should be placed at a mean distance d above sigma: Divide the system in B=(L/rc)^3 boxes and place the particle within the box (random or center, no big deal). The cutoff range rc should be several (3 -5) sigmas

Initial Configuration: Velocities Sample from a Gaussian with zero average and variance k. T/m Sample from a gaussian (standard from libraries) See also Box-Mueller algorithm Where u 1 and u 2 are uniform rn in [0, 1]

Time-evolution In principle we have “just” to integrate Newton’s equations. It is of utmost importance to secure time-reversibility, i. e. Liouville theorem: Symplectic integrators. In addition, forces should be computed as efficiently as possible because they take most CPU time.

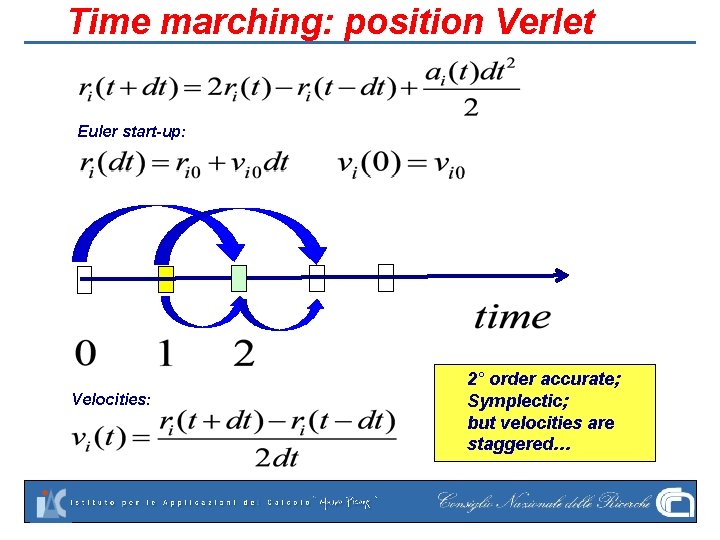
Time marching: position Verlet Euler start-up: Velocities: 2° order accurate; Symplectic; but velocities are staggered…

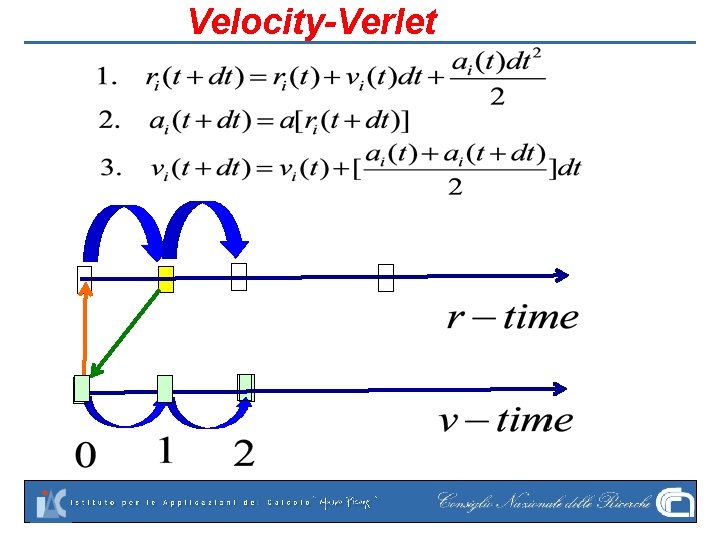
Velocity-Verlet

VV is symplectic Let Velocity-Verlet: Let q. e. d VV is pretty popular: 2° order, simplectic, time-aligned

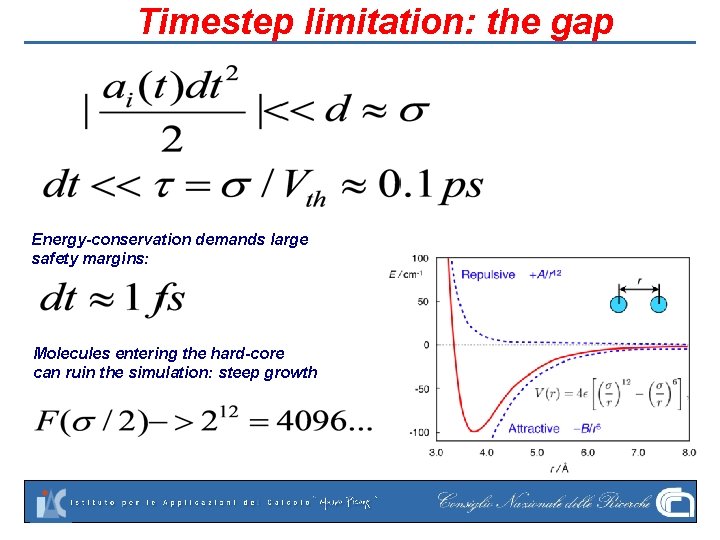
Timestep limitation: the gap Energy-conservation demands large safety margins: Molecules entering the hard-core can ruin the simulation: steep growth

Force computation This is the most CPU time consuming section of MD simulations, scaling in principle like N*N. However, short-range potentials can be dealt with more economic techniques: 1. Direct N-body summation 2. Cut-off summation 3. Linked lists
![Force Calculation Direct Summation for i1 iN1 i Fi0 0 for ji1 jN Force Calculation: Direct Summation for (i=1; i<=N-1; i++) { F[i]=0. 0; for (j=i+1, j<=N,](https://slidetodoc.com/presentation_image_h/283ab853f659d9e6aeb5ccf6700d653d/image-24.jpg)
Force Calculation: Direct Summation for (i=1; i<=N-1; i++) { F[i]=0. 0; for (j=i+1, j<=N, j++{ xij = x[i]-x[j]; yij = y[i]-y[j]; zij = z[i]-z[j]; rij=pow(xij*xij+yij*yij+zij*zij, 0. 5) Fij = -a/rij**13 +b/rij**7 F[i]+=Fij; F[j]-=Fij }
![Force Calculation short range for i1 iN i Fi0 0 for ji1 jN Force Calculation: short range for (i=1; i<=N; i++) { F[i]=0. 0; for (j=i+1, j<=N,](https://slidetodoc.com/presentation_image_h/283ab853f659d9e6aeb5ccf6700d653d/image-25.jpg)
Force Calculation: short range for (i=1; i<=N; i++) { F[i]=0. 0; for (j=i+1, j<=N, j++{ But N(N-1)/2 branches… xij = x[i]-x[j]; yij = y[i]-y[j]; zij = z[i]-z[j]; rij=pow(xij*xij+yij*yij+zij*zij, 0. 5) if(rij<=rcut) { Fij = -a/rij**13 +b/rij**7 F[i]+=Fij }} conditional

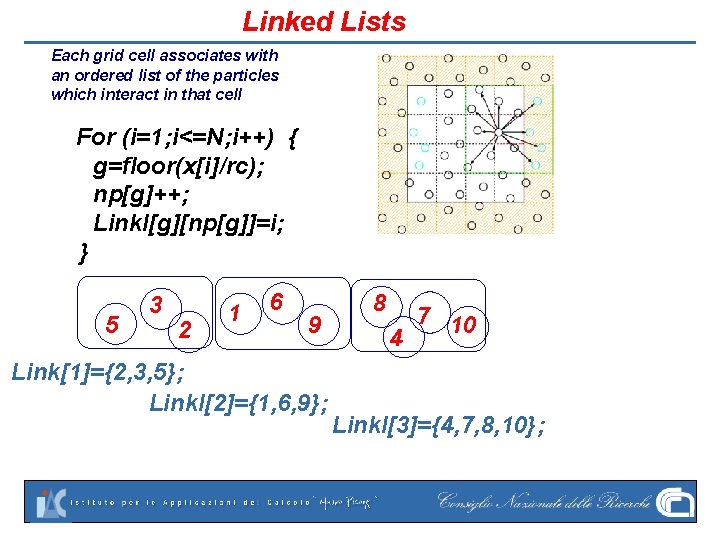
Linked Lists Each grid cell associates with an ordered list of the particles which interact in that cell For (i=1; i<=N; i++) { g=floor(x[i]/rc); np[g]++; Linkl[g][np[g]]=i; } 5 3 2 1 6 9 Link[1]={2, 3, 5}; Linkl[2]={1, 6, 9}; 8 4 7 10 Linkl[3]={4, 7, 8, 10};
![Force Calculation Linked List for i1 iN i gfloorxirc for k1 knpcg k Force Calculation: Linked List for (i=1; i<=N; i++) { g=floor(x[i]/rc); for (k=1, k<=npc[g], k++{](https://slidetodoc.com/presentation_image_h/283ab853f659d9e6aeb5ccf6700d653d/image-27.jpg)
Force Calculation: Linked List for (i=1; i<=N; i++) { g=floor(x[i]/rc); for (k=1, k<=npc[g], k++{ j = Linkl[g][k]; xij= x[i]-x[j]; Fij = -a/xij**13 +b/xij**7 F[i]+=Fij; F[j]-=Fij }}

Data Analysis

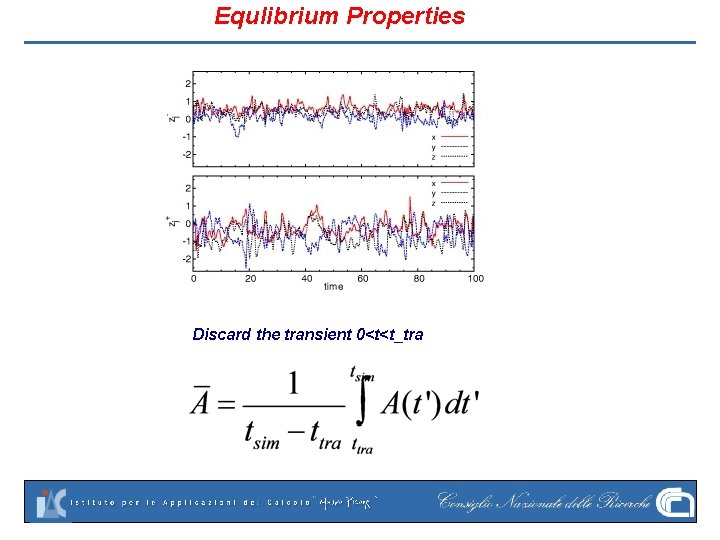
Equlibrium Properties Discard the transient 0<t<t_tra

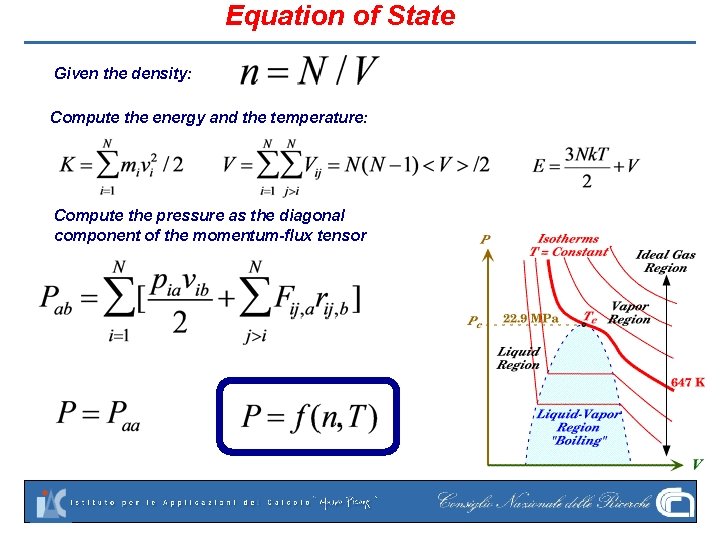
Equation of State Given the density: Compute the energy and the temperature: Compute the pressure as the diagonal component of the momentum-flux tensor

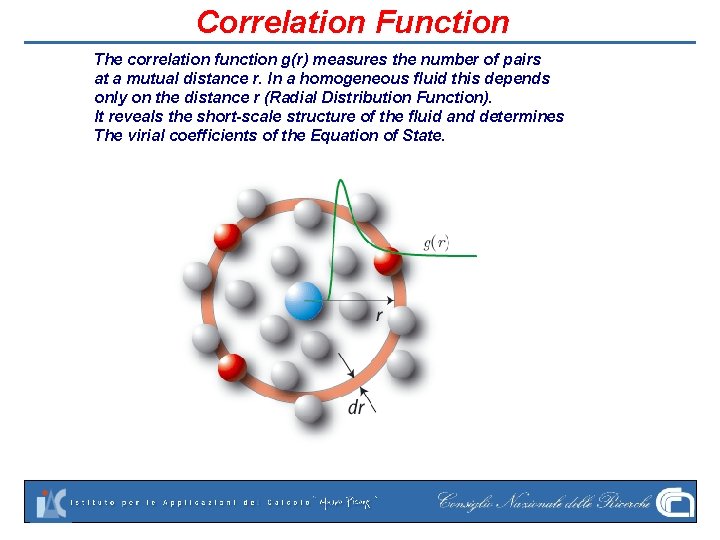
Correlation Function The correlation function g(r) measures the number of pairs at a mutual distance r. In a homogeneous fluid this depends only on the distance r (Radial Distribution Function). It reveals the short-scale structure of the fluid and determines The virial coefficients of the Equation of State.

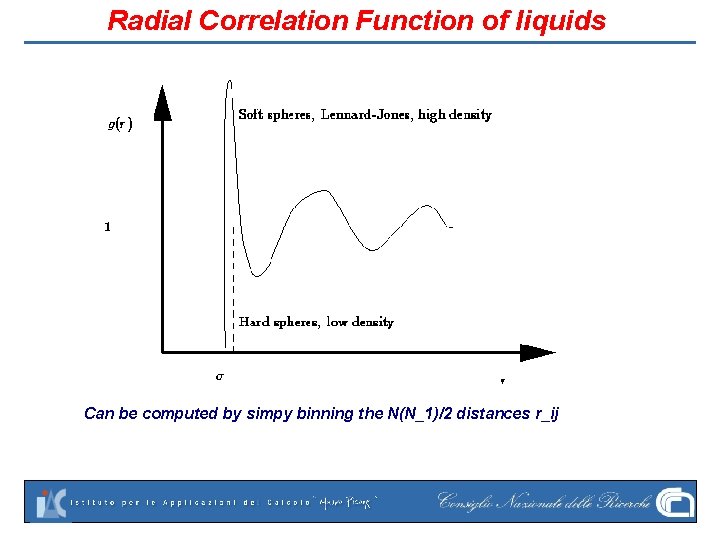
Radial Correlation Function of liquids Can be computed by simpy binning the N(N_1)/2 distances r_ij

Ensembles

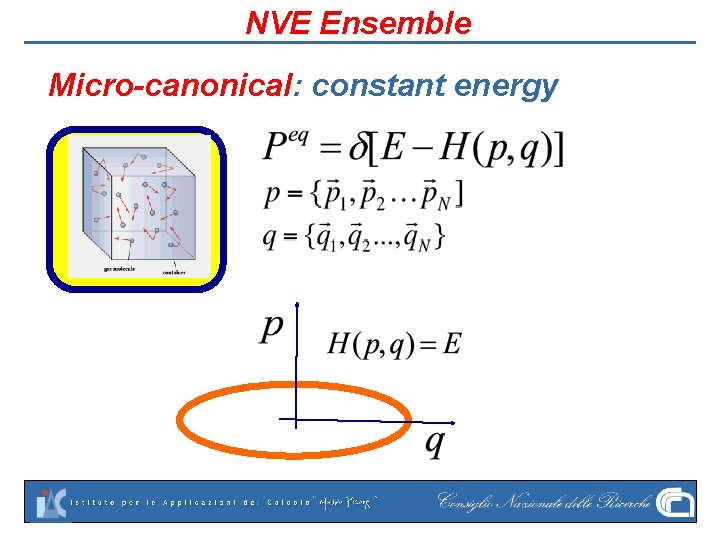
NVE Ensemble Micro-canonical: constant energy

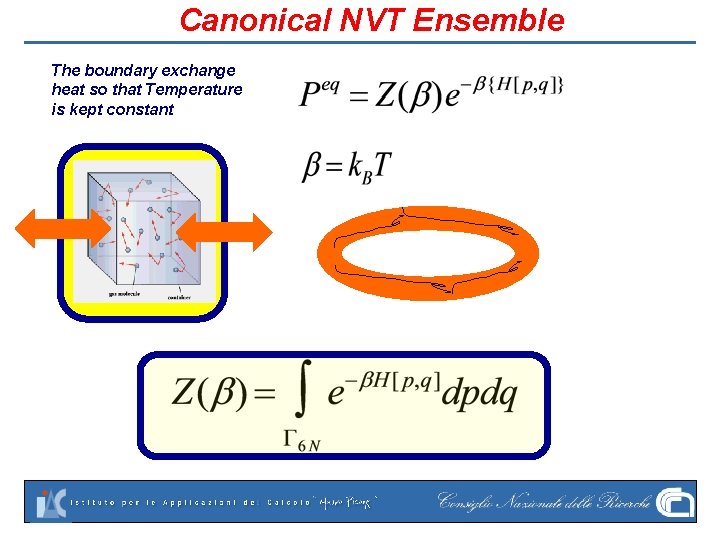
Canonical NVT Ensemble The boundary exchange heat so that Temperature is kept constant

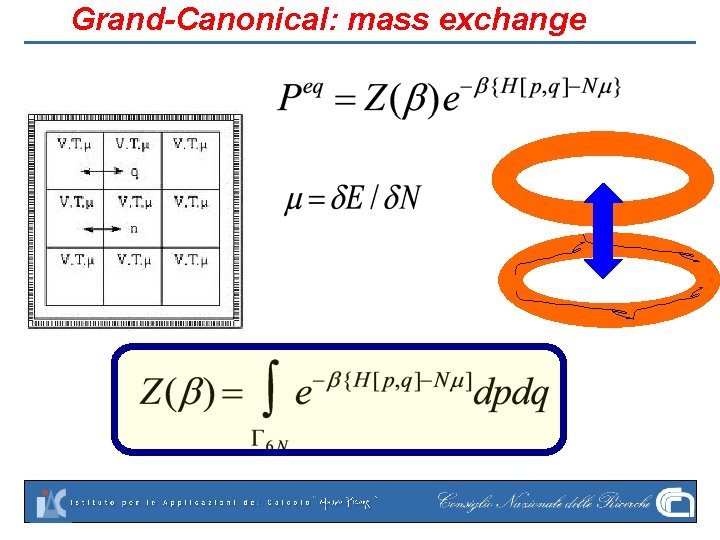
Grand-Canonical: mass exchange

Thermostats

Isokinetic Ensemble (T=const) This defines the kinetic temperature: The dynamics does not conserve KE, hence one must rescale each velocity to keep the reference Temperature constant: However this does not allow any temperature fluctuation, So it does not sample the canonical NVT ensemble

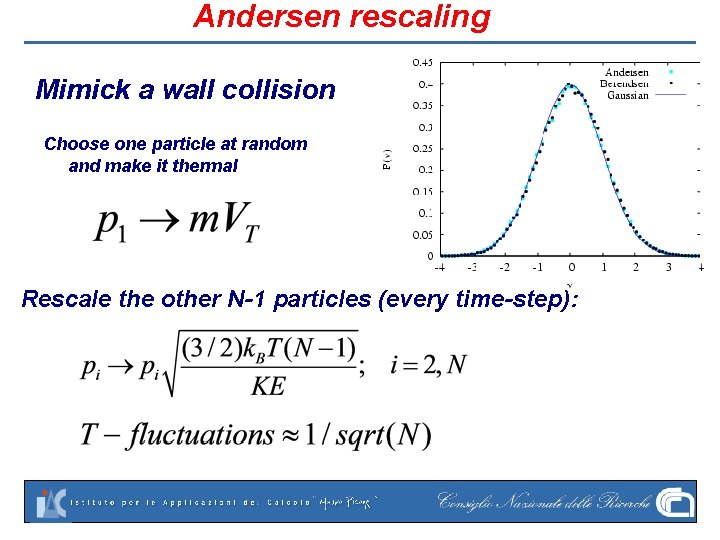
Andersen rescaling Mimick a wall collision Choose one particle at random and make it thermal Rescale the other N-1 particles (every time-step):

Canonical NVT

Velocity-Verlet-Nose’-Hoover

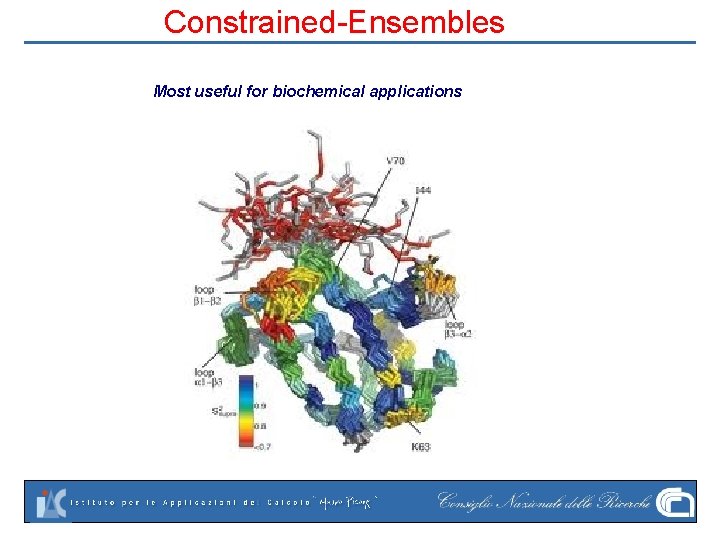
Constrained-Ensembles Most useful for biochemical applications

Fixed-length bonds Holonomic Constraint: the bond length is constant in time The l-th and m-th particles participate to constraint k

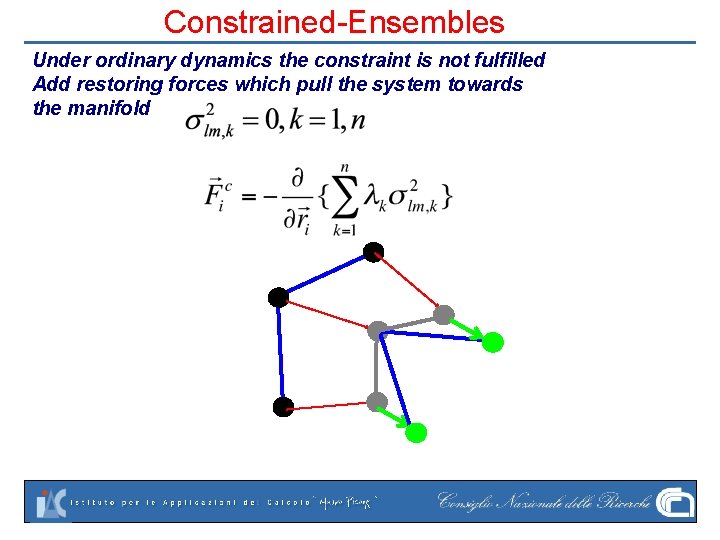
Constrained-Ensembles Under ordinary dynamics the constraint is not fulfilled Add restoring forces which pull the system towards the manifold

SHAKE algorithm 1. Move with ordinary forces to unconstrained positions 2. Apply the constraint force 3. Solve the n nonlin eqs for the n Lagrangian multipliers This is usually done by a nonlinear (Newton-Raphson) iterator

Boundary Conditions

Solid Walls NVE: Particles impinging on the well are reinjected along a random direction and conserved magnitude NVT: Drawn from a Maxwellian at temperature T

MD: state of the Art Many community codes: CHARMM (Chemistry at Harvard Macromolecular Mechanics) GROMACS (GROeningen Machine for Advanced Chemical Simulations) AMBER (Assisted Model Building with Energy refinement) LAMPS (Large scale Atomic/Molecular Massively Parallel Simulator) Special purpose computers ANTON (D. Shaw Research)

Multi-billion MD sim’s

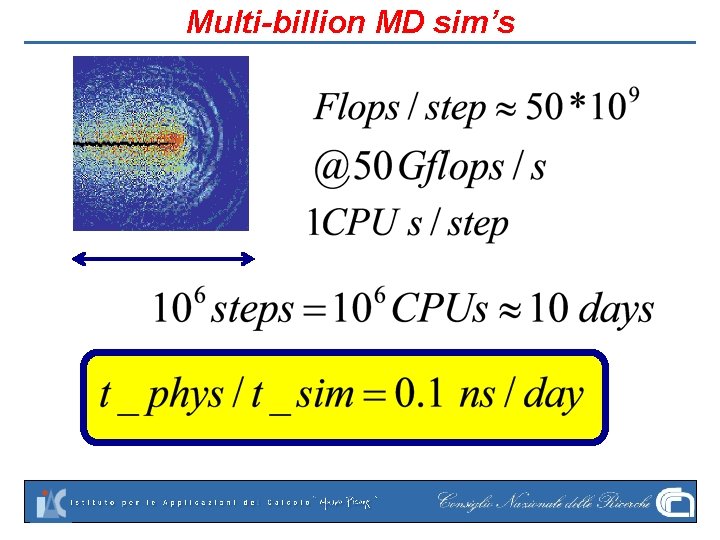
Multi-billion MD sim’s

ANTON: Shaw Research Current academic state-of-the art: 100 ns/day for O(100 K ) molecules ANTON: 20, 000 ns/day (small systems, 25 Katoms with protein though)

Assignements 1. Write your own MD code with solid walls as thermostat (see codlet md. f) 2. Check energy conservation 3. Compute the pressure in the box as the momentum per unit area exchanged with the walls Pressure=F/A, F=d. P/dt 4. Repeat 2. for many values of the density and construct the equation of state P=P(rho, T)

End of the Lecture

Constraint Forces

Hard/Soft potentials

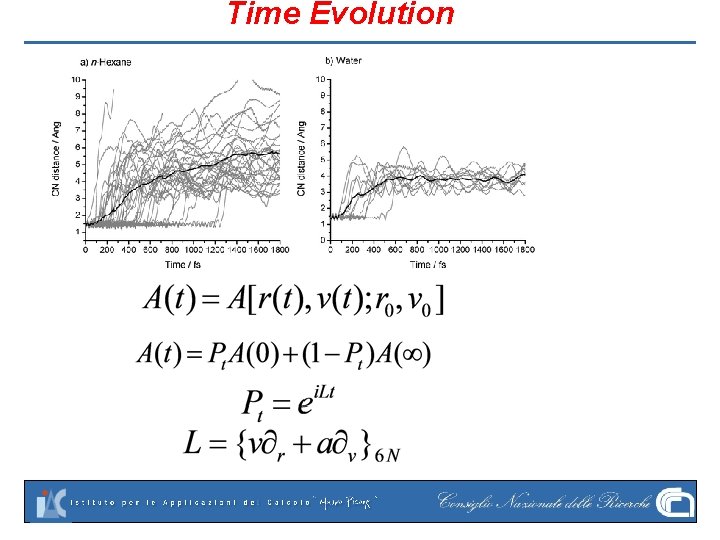
Time Evolution