EIGENSYSTEMS SVD PCA Big Data Seminar Dedi Gadot

EIGENSYSTEMS, SVD, PCA Big Data Seminar, Dedi Gadot, December 14 th, 2014

EIGVALS AND EIGVECS

Eigvals + Eigvecs •
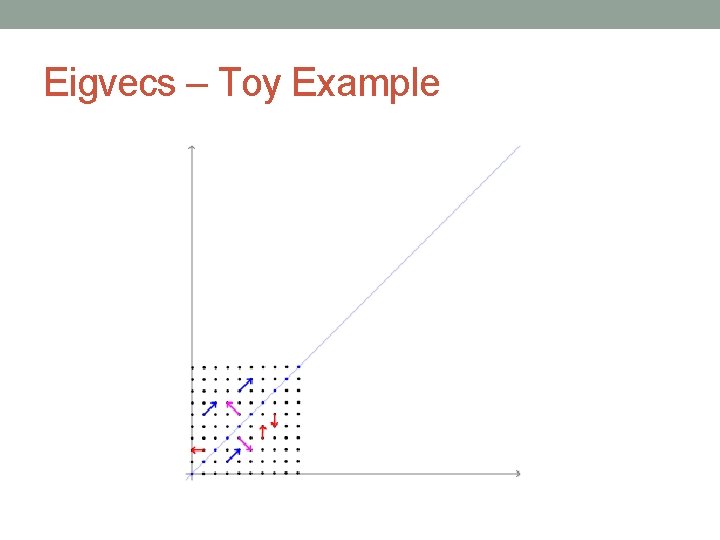
Eigvecs – Toy Example

Geometric Transformations

SVD

SVD • Singular Value Decomposition • A factorization of a given matrix to its components: M = UΣV∗ • When: • M – an m x n real or complex matrix • U – an m x m unitary matrix, called the left singular vectors • V – an n x n unitary matrix, called the right singular vectors • Σ – an m x n rectangular diagonal matrix, called the singular values

Applications and Intuition • If M is a real, square matrix – • U, V can be referred to as rotation matrices and Σ as a scaling matrix M = UΣV∗

Applications and Intuition • The columns of U and V are orthonormal bases • Singular vectors (of a square matrix) can be interpreted as the semiaxes of an ellipsoid in n-dimensional space • SVD can be used to solve homogeneous linear equations • Ax=0, A is a square matrix x is the right singular vector which corresponds to a singular value of A which is zero • Low rank matrix approximation • Take Σ of M and leave only the r largest singular values, rebuild the matrix using U, V and you’ll get a low rank approximation of M • …

SVD and Eigenvalues • Given an SVD of M the following two relations hold: • The columns of V are eigenvectors of M*M • The columns of U are eigenvectors of MM* • The non-zero elements of Σ are the square roots of the non- zero eigenvalues of M*M or MM*

PCA

PCA • Principal Components Analysis • PCA can be thought as fitting an n-dimensional ellipsoid to the data, such that each axis of the ellipsoid represents a principal component, i. e. an axis of maximal variance
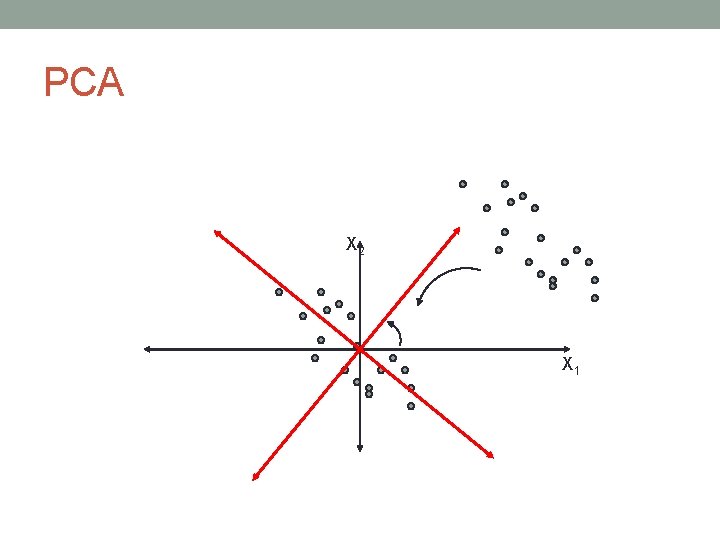
PCA X 2 X 1

PCA – the algorithm • Step A – subtract the mean of each data dimension, thus move all data-points to be centered around the origin • Step B – calculate the covariance matrix of the data

PCA – the algorithm • Step C – calculate the eigenvectors and the eigenvalues of the covariance matrix • The eigenvectors of the covariance matrix are orthonormal (see below) • The eigenvalues tell us the ‘amount of variance’ of the data along each specific new dimension/axis (eigenvector)

PCA – the algorithm • Step D – sort the eigenvalues in descending order • Eigvec #1, which is correlated with Eigval #1, is the 1 st principal component – i. e. the (new) axis with highest variance • Step E (optional) – take only ‘strong’ Principal Components • Step F – project the original data on the newly created base (the PCs, the eigenvectors) to get a rotated, translated coordinate system correlated with highest variance per each axis
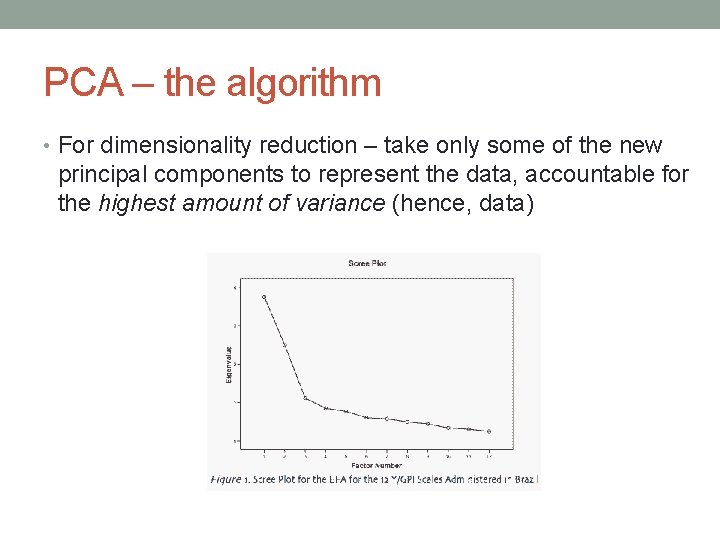
PCA – the algorithm • For dimensionality reduction – take only some of the new principal components to represent the data, accountable for the highest amount of variance (hence, data)
- Slides: 17