Overview of Bioconductor Aedn Culhane aedinjimmy harvard edu

Overview of Bioconductor Aedín Culhane aedin@jimmy. harvard. edu http: //bcb. dfci. harvard. edu/~aedin http: //www. hsph. harvard. edu/research/aedin-culhane

Bioconductor Biannual release (normally April, October) to coincide with R release. Current: Bioconductor 2. 9 (release coincide with R 2. 14) To install use script on Bioconductor Website source("http: //www. bioconductor. org/bioc. Lite. R") bioc. Lite()

Packages Overview Bio. Conductor web site • Bioconductor Bioc. Views Task view Software Annotation Data Experimental Data

What Packages do I need? Specific to you data and analysis pipeline but for examples: • Bioconductor Workshops • Bioconductor Workflows

Main types of Annotation Packages • Gene centric Annotation. Dbi packages: – Organism: org. Mm. eg. db. – Technology/Platform: hgu 133 plus 2. db. – Gene. Sets and Pathway (biology level): GO. db or KEGG. db –. db packages can be queried with sql or accessed using annotation package (totable, get, mget) • Genome centric Genomic. Features packages: – Transriptome level: Tx. Db. Hsapiens. UCSC. hg 19. known. Gene – Generic features: Can generate via Genomic. Features • bioma. Rt: – Query web-based `biomart' resource for genes, sequence, SNPs, and etc. • See http: //www. bioconductor. org/help/course-materials/2011/Bio. C 2011/Lab. Stuff/Annotation. Slides. Bioc 2011. pdf

Bioconductor resources • Mailing List (sign up for daily digest) • Documentation, workshop/course material online – Slides from talks, pdf of tutorials, R code • Help available for each software package – Each package MUST contain vignette (howto) • Other resources ww. Rseek. org www. r-bloggers. com

Vignette • Tutorials, provide worked example of package • Required in Bioconductor packages • Written in Sweave (Leisch, 2002). – LATEX dynamic reports in which R code is embedded and executable – All R code in vignette is checked (and executed) by R CMD check – http: //www. bioconductor. org/docs/vignettes. html library("Biobase") library("GOstats") open. Vignette() # Load package of interest

S 4 classes and Expression. Set • Within Bioconductor, you will encounter packages are structured around S 4 objectoriented programming proposed by John Chambers (developer of S) • A class provides a software abstraction of a real world object. • A method performs an action on a class (Think of a class as a noun, and method as verb)

Object (S 4) • An object is an instance of a class. • Descriptions are stored in slots • slot. Names(ob 1) lists all slots in object, or use str(). • To access slots – ob 1@slotname – slotname(ob 1), or – slot(ob 1, “slotname")

Example: Expression. Set > ALL library(ALL) data(ALL) slot. Names(ALL) ALL@pheno. Data(ALL) Expression. Set (storage. Mode: locked. Environment) assay. Data: 12625 features, 128 samples element names: exprs protocol. Data: none pheno. Data sample. Names: 01005 01010. . . LAL 4 (128 total) var. Labels: cod diagnosis. . . date last seen (21 total) var. Metadata: label. Description class(ALL) feature. Data: none ? Expression. Set experiment. Data: use 'experiment. Data(object)' pub. Med. Ids: 14684422 16243790 Annotation: hgu 95 av 2

Method which act on a S 4 class show. Methods(class= "Expression. Set") get. Method("write. exprs", "Expression. Set") Or if you wish to see how the package really works, download and look the source code

Getting Data into R & Bioconductor Aedín Culhane aedin@jimmy. harvard. edu http: //www. hsph. harvard. edu/research/aedin-culhane/

Simple Excel Spread. Sheet data • Simple table – read. table() – read. csv() – scan() • However more datatype specialized. See Technologies on Bioc. Views. – http: //www. bioconductor. org/packages/release/Bioc. Vi ews. html • Large data files. Also see http: //www. revolutionanalytics. com 13

Some common data types • Microarray • SNP • NGS May 2011 14
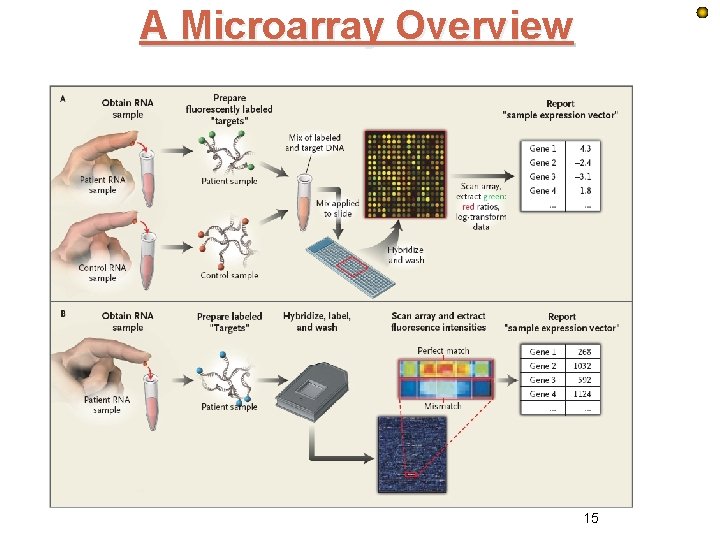
A Microarray Overview 15

Reading Affymetrix Data library(affy) require(affy) # Alternative affybatch <- Read. Affy(celfile. path="[Location of your data]") e. Set<-just. RMA() May 2011 16

Sample R code 17

Expression. Set Class in R May 2011 18

Assessing Data Quality May 2011 19

Public Microarray Data Array. Express • 21997 Studies (622, 617 profiles, ) GEO • 22, 735 Studies (558, 074 profiles) Statistics May 2011

R Code May 2011 21

More on GEOquery require(GEOquery) Let's try to load the GDS 810 dataset which contains data on Alzheimer's disease at various stages of severity. GDS 810<-get. GEO("GDS 810") The get. GEO function returns an object of class GEOData. You can get a description of this class like this: help("GEOData-class") Meta(GDS 810) Columns(GDS 810) head(Table(GDS 810)) May 2011 22

Affy SNP Arrays May 2011 23

Process – Affy SNP Arrays (Oligo package) May 2011 24

Other Arrays • Illumina – Lumi package • 2 color spotted arrays – Limma package • Other arrays – http: //www. bioconductor. org/help/workflows/oli go-arrays/ May 2011 25
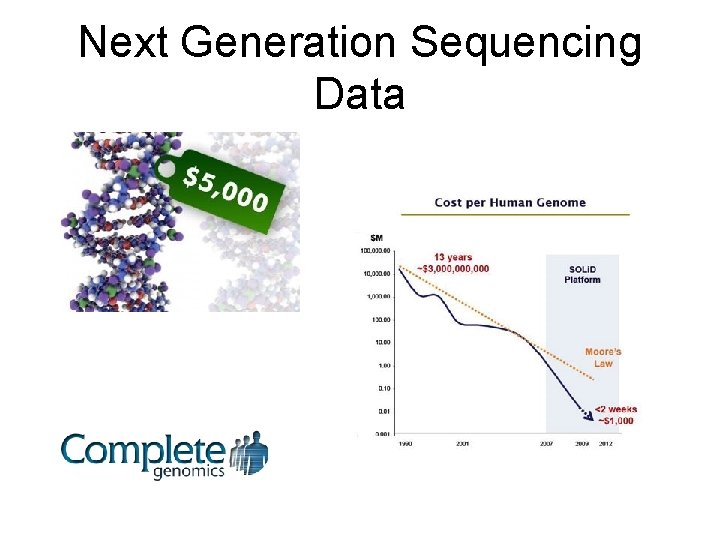
Next Generation Sequencing Data

R Code May 2011 27

Exercise • Install the library GEOquery • Download the dataset GSE 1297 using get. GEO • This data will be downloaded as an e. Set, so to see the expression data and pheno. Data, use p. Data and exprs • Use Array. Quality. Metrics to Assess the data quality of these data May 2011 28

R basics: Getting help • To get help – ? mean – help(mean) • help. search(“mean”) • apropos("mean") • example(mean) • http: //www. bioconductor. org/help/

• With thanks to • www. bioconductor. org/help/course. . . /Bioconductor-Introduction-lab. pdf May 2011 30
- Slides: 30