NHS GMC lab and VR Guidance updates Sam

NHS GMC lab and V@R Guidance updates • Sam Clokie (WMGMC) – GEL 2 MDT (WMGMC and North Thames GMC collaboration) • Emma Baple (GE and South West GMC) – Update on Genomic MDT guidance – Working Part outcomes • Dom Mc. Mullan (WMGMC/ACGS) – Update on progress with RD V@R Guidance doc

WGS Reporting and validation at WMRGL using GEL 2 MDT Samuel Clokie, Ph. D WMRGL | Uo. B

Presentation outline 1. Project history 2. Background on GEL 2 MDT – Project History – User requirements – Design principles 3. 4. 5. 6. Overview of GEL 2 MDT connectivity (APIs etc) GEL 2 MDT Features Example case walkthrough – with screenshots Future plans – Genotyping integration – Hosting – Integration with Genie

Project History • GOSH and BWCH working on similar idea • GOSH had developed a case management web based tool, utilising. csv files • WMRGL had started exploring CIP-API, started GUI development • …work together to produce GEL 2 MDT • Close collaboration has allowed rapid success with equal contribution to GEL 2 MDT

GOSH and WMRGL Bioinformaticians co-design and build GEL 2 MDT Patrick Lombard Ed Stone Helena Ahlfors Samuel Clokie Theo Cole

User Requirements from WMRGL • Need an alternative to Excel • Create a workflow management tool to track patients progress from GEL, through MDT to report • Simple and intuitive to use • Automate data gathering (APIs) • To store discussion/actions about variants and patients during the MDT • To audit the number of cases we are processing every month • Audit the decision making process

Design principles • Web based – locally or nationally hosted – Django based (ORM API) • Allow seamless integration with national bioinformatics tools/ other systems – Python. – OO approach; efficient, readable and extensible – REST API • Extensive unit tests • Backup procedure

GEL 2 MDT features • Highly normalised schema, with postgre. SQL for fast performance • Utilises APIs as much as possible to integrate data sources • Hashes case JSONs, creating a history of changes • Runs VEP for HGVS • Configurable to new JSON file structures • REST API • Documentation (Sphinx) • Available on github
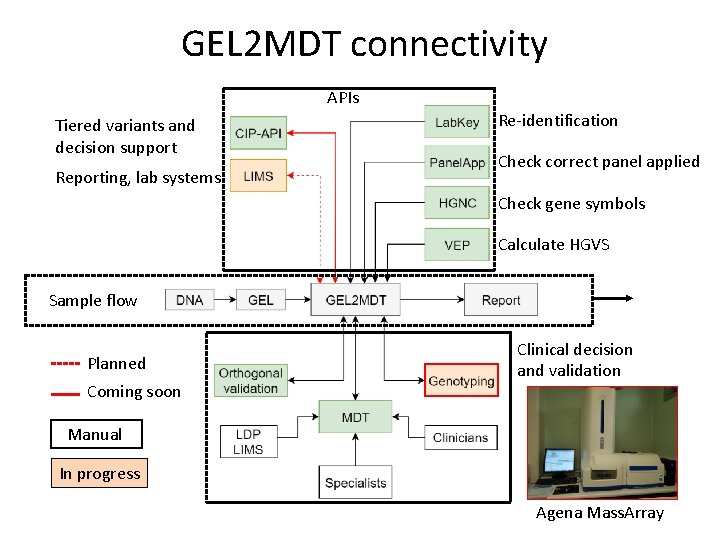
GEL 2 MDT connectivity APIs Tiered variants and decision support Reporting, lab systems Re-identification Check correct panel applied Check gene symbols Calculate HGVS Sample flow Planned Coming soon Clinical decision and validation Manual In progress Agena Mass. Array
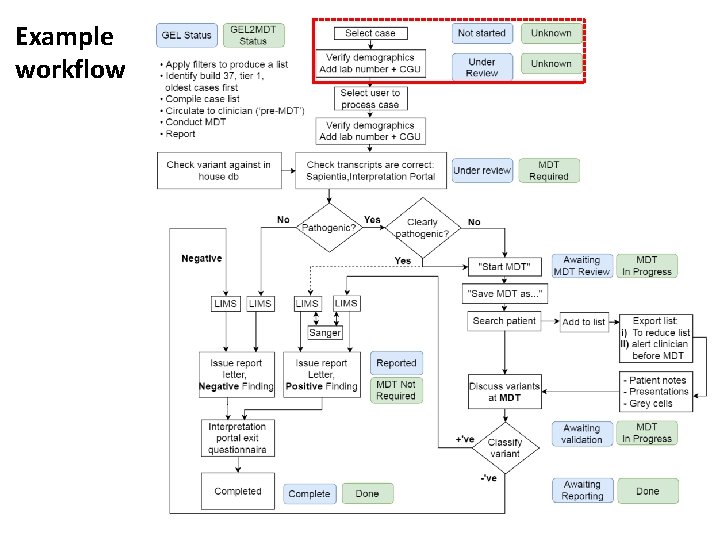
Example workflow
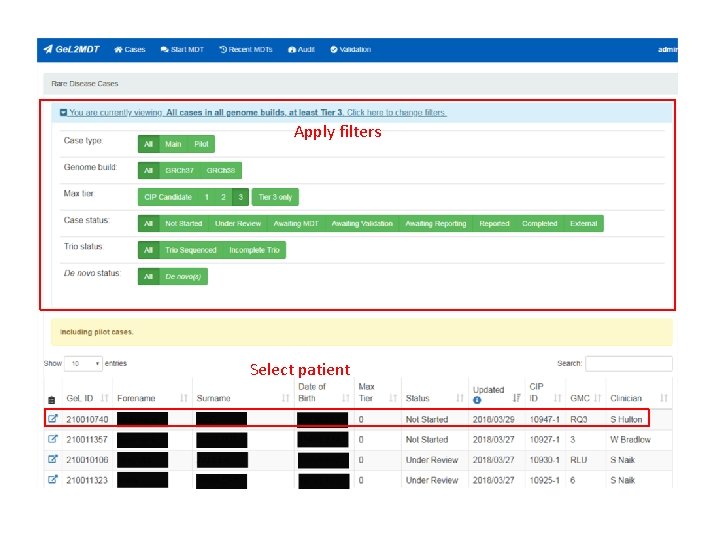
Apply filters Select patient
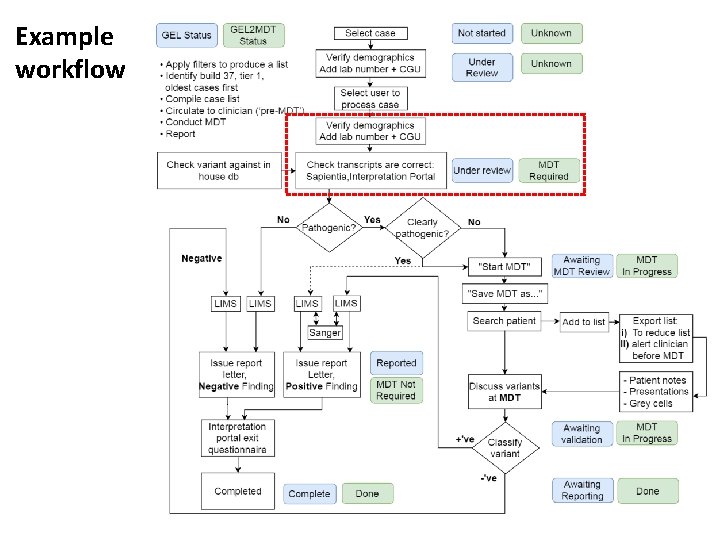
Example workflow
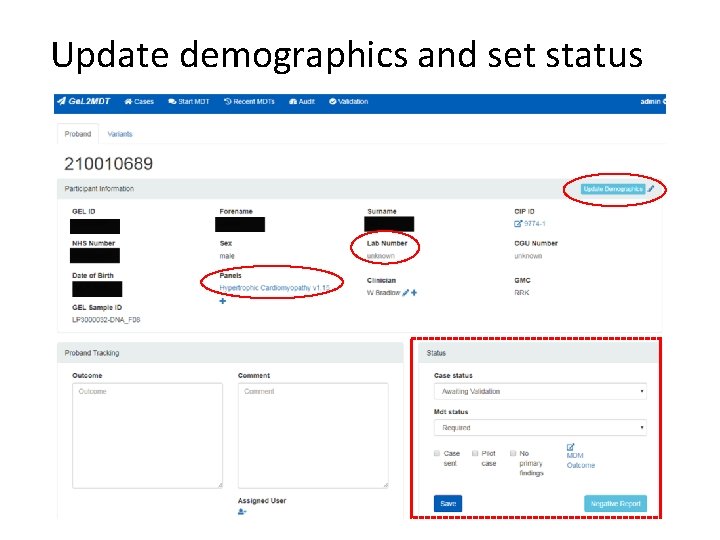
Update demographics and set status
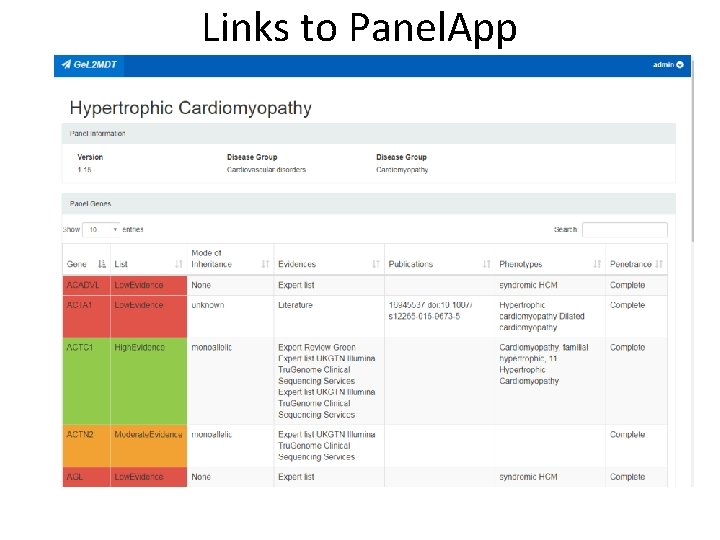
Links to Panel. App
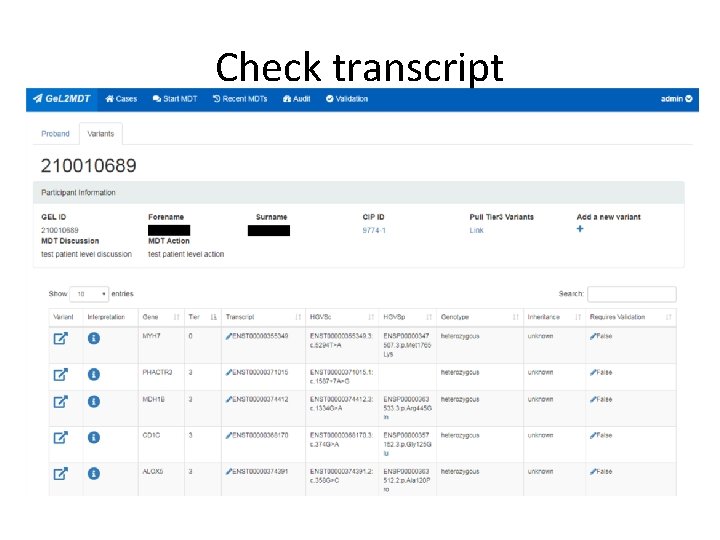
Check transcript
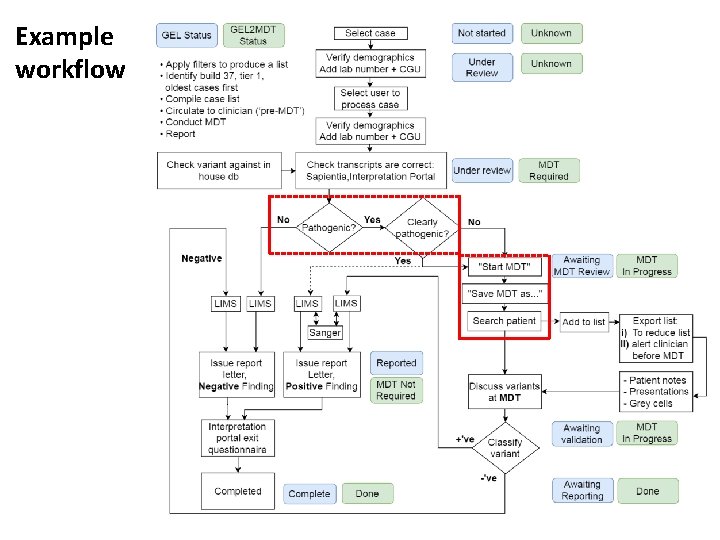
Example workflow
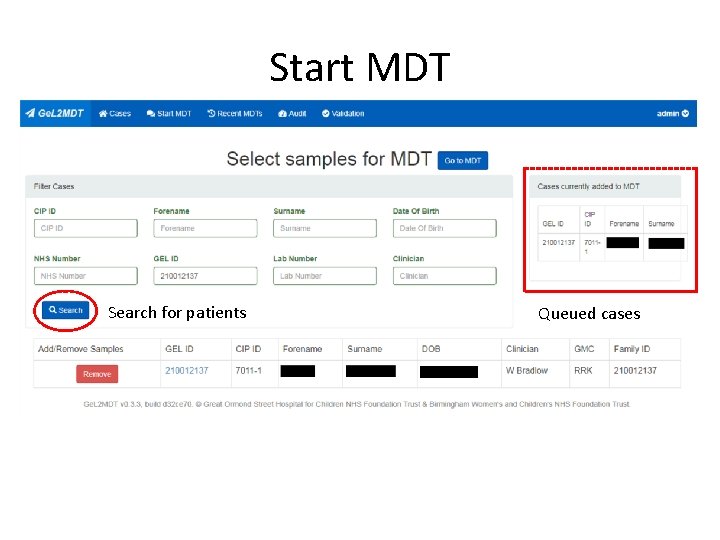
Start MDT Search for patients Queued cases
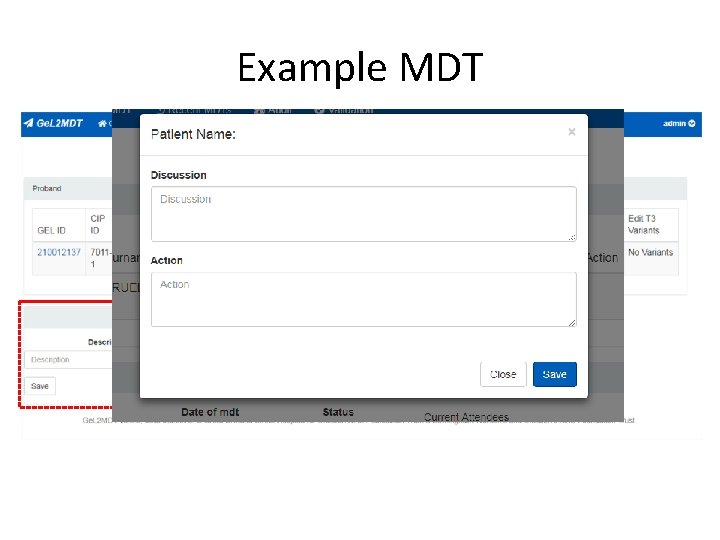
Example MDT
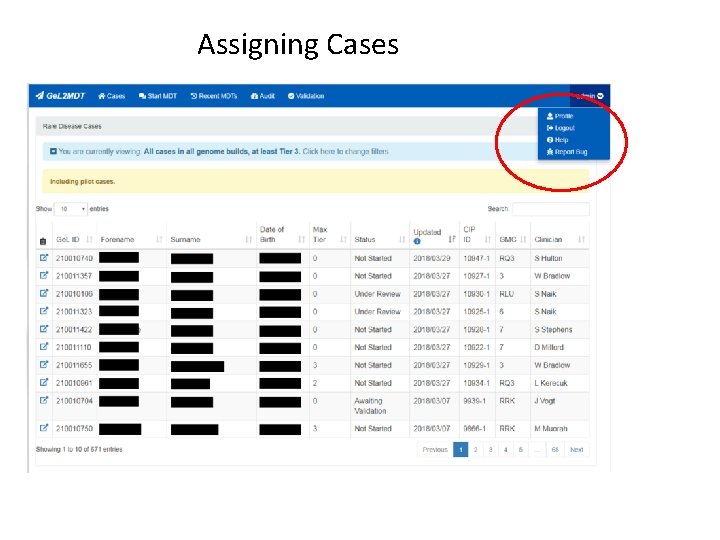
Assigning Cases
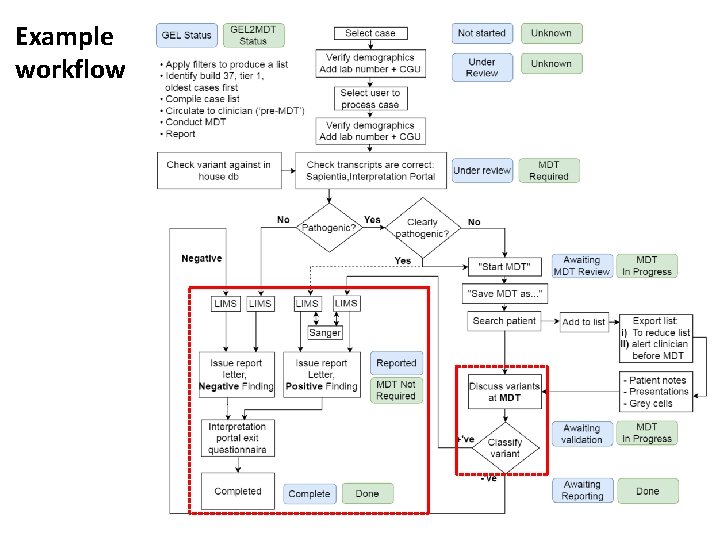
Example workflow
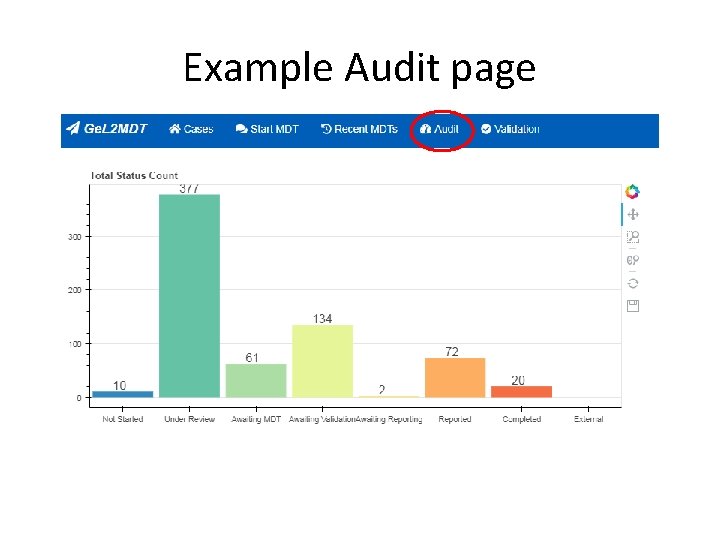
Example Audit page

Summary & Future plans • • • Makes a multi-step pathway manageable CIP-API and support from GEL invaluable Eliminates reliance on excel files, reducing risk Saves £ by reducing time needed per case/calculating manual audits Several ways to use GEL 2 MDT; many linkouts Evolving field, therefore flexible framework is useful Open Source – available on Git Documentation – under construction National hosting: integration with Genie (via API) Genotyping assay integration: Agena platform

Acknowledgments • • Patrick Lombard (GOSH) Theo Cole (BWCH) Ed Stone (BWCH) Helena Ahlfors (GOSH) • • BWCH: Dom Mc. Mullan Kirsten Mackay-Bouford Sarah Turton Gavin Ryan

Questions?
- Slides: 24