Exome sequencing and characterization of 49 960 individuals









- Slides: 19

Exome sequencing and characterization of 49, 960 individuals in the UK Biobank Van Hout et al, Oct 2020 �Genotyping, imputation & sequencing �Whole-Exome Sequencing ◦ What is the ‘exome’? ◦ My experience �UKB 50 k Exomes – “flagship” paper Genetics Forum: 05/11/20 Mesut Erzurumluoglu 1

Genotyping v whole exome sequencing • Genotyping involves inferring genotypes at SNP locations – A typical genotyping array will genotype ~850 k SNPs – with imputation, this figure will rise to 5 -10 million high quality (INFO>0. 8) genotype calls in European samples Image by Kat M Research, Flickr. • whole sequence of a person’s protein coding parts of the genome – the most important region – This will identify >50 k common, rare and ultra-rare coding variants per individual Image courtesy of Wellcome Library, London

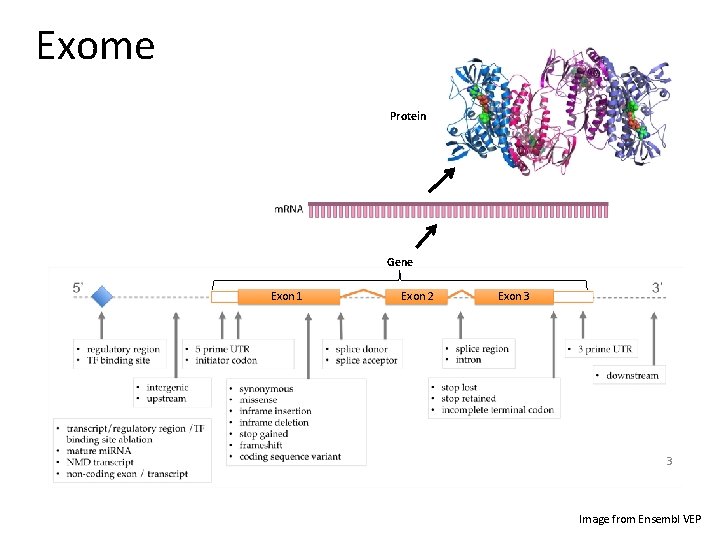
Exome Protein Gene Exon 1 Exon 2 Exon 3 3 Image from Ensembl VEP

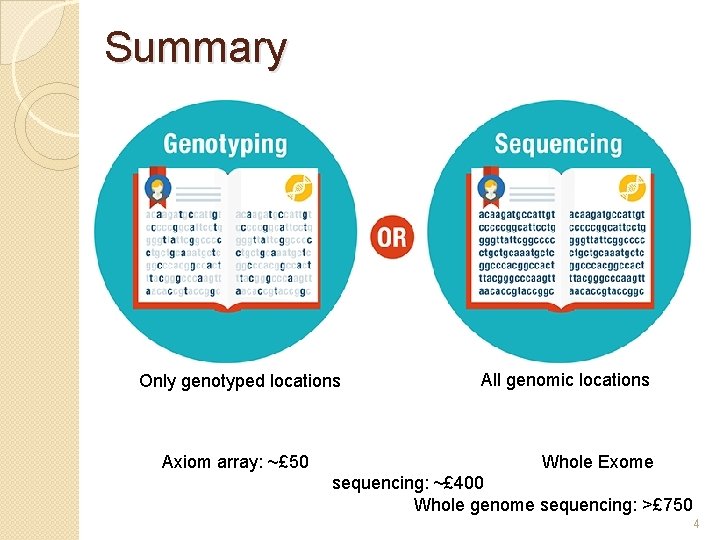
Summary Only genotyped locations Axiom array: ~£ 50 All genomic locations Whole Exome sequencing: ~£ 400 Whole genome sequencing: >£ 750 4

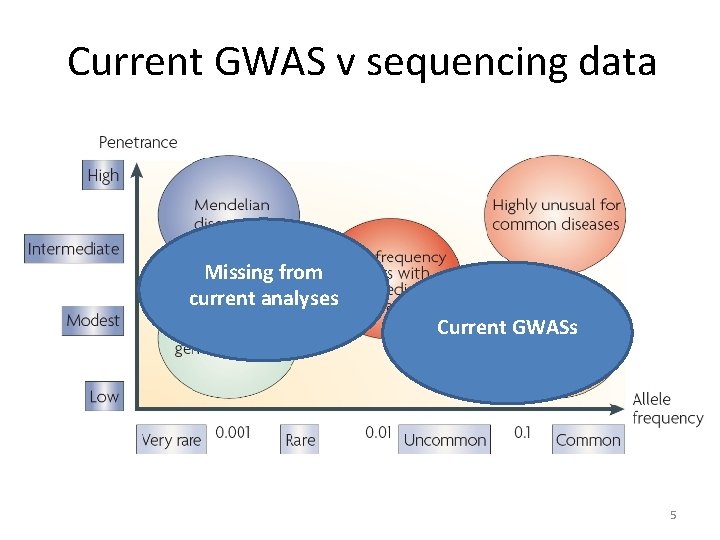
Current GWAS v sequencing data Missing from current analyses Current GWASs 5

9 whole-exomes = Ph. D thesis (2015) Raw DNA -> PCD causal variant c. 925 G>T: p. (E 309*) in CCDC 151 – ref 1 c. 406 C>T: p. (R 136*) in DNAAF 3 1 - Alsaadi & Erzurumluoglu et al, 2014. Nonsense Mutation in CCDC 151 Causes Primary Ciliary Dyskinesia. Human Mutation **Shared as a preprint (2014) on Bio. Rxiv **2 - Erzurumluoglu et al, 2015. Identifying highly-penetrant disease causal mutations using next generation sequencing: Guide to whole process. Bio. Med Res. Int.

UKB 50 K ‘pre-analysis’ stage � Conversion of sequencing data in BCL format to FASTQ format: bcl 2 fastq � Read alignment: bwa 0. 7. 17 � Duplicate marking, stats gathering: picard v 1. 141 � SAM/BAM/CRAM file generation and manipulation: samtools v 1. 7 � Variant calling: We. Call v 1. 1. 2 (Genomics plc) � Sequence Quality Control: Fast. QC 0. 11. 8 � VCF file manipulation and index generation: bcftools v 1. 7 � Ancestry predictions, IBD estimate, pedigree reconstruction: PLINK v 1. 90 Association study � Single variant and burden tests ◦ Quantitative traits: BOLT-LMM_v 2. 3. 2 ◦ Binary outcomes: SAIGE_v 0. 29. 1 7

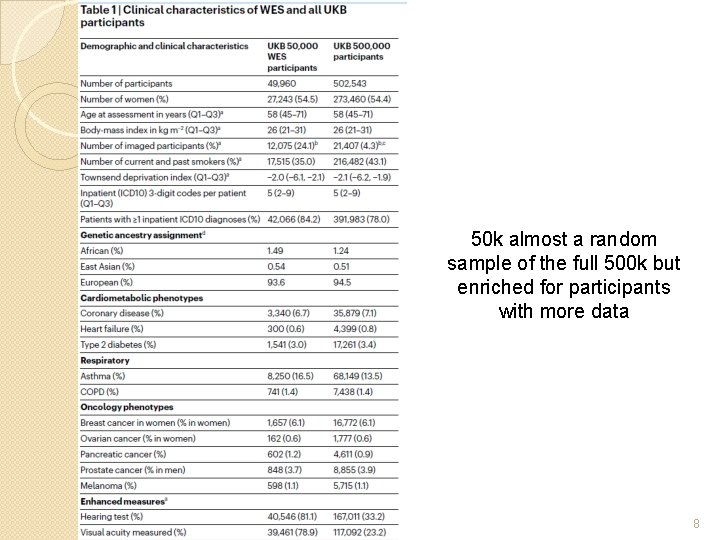
50 k almost a random sample of the full 500 k but enriched for participants with more data 8

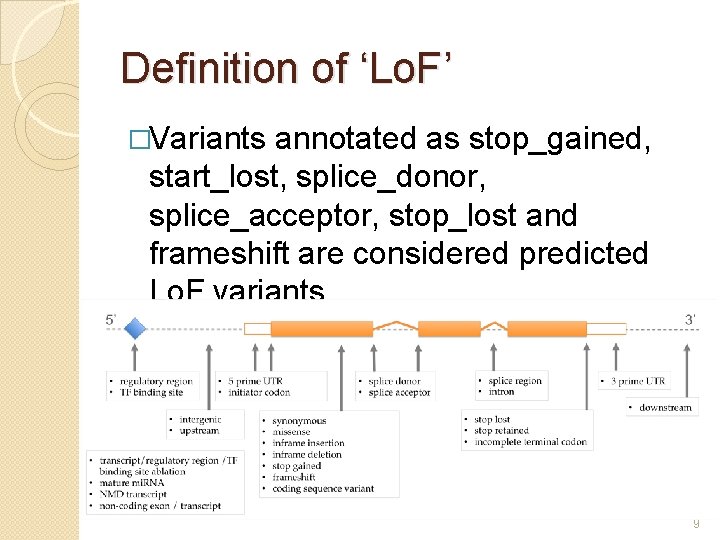
Definition of ‘Lo. F’ �Variants annotated as stop_gained, start_lost, splice_donor, splice_acceptor, stop_lost and frameshift are considered predicted Lo. F variants 9

Main conclusions � N= 49, 960 high-quality exomes � ~4 million coding variants ◦ coverage >20 x at 94. 6% of sites on average (s. d. 2. 1%) ◦ ~98. 6% have a MAF of <1%. ◦ 198, 269 autosomal predicted loss-of-function (Lo. F) variants �>14 -fold increase in SNVs compared to imputed sequence (16. 1 -fold increase for indels) � 17, 718 (>97% of) genes had 1 or more Lo. F variant and 69% of genes had 10 or more � Association study of 1, 730 phenotypes ◦ PIEZO 1 on varicose veins, COL 6 A 1 on corneal resistance, MEPE on bone density, and IQGAP 2 and GMPR on blood cell traits � Prevalence of pathogenic variants of clinical importance (medically actionable variant) is 2% � Penetrance of BRCA 1&2 variants is lower than previous estimates 10

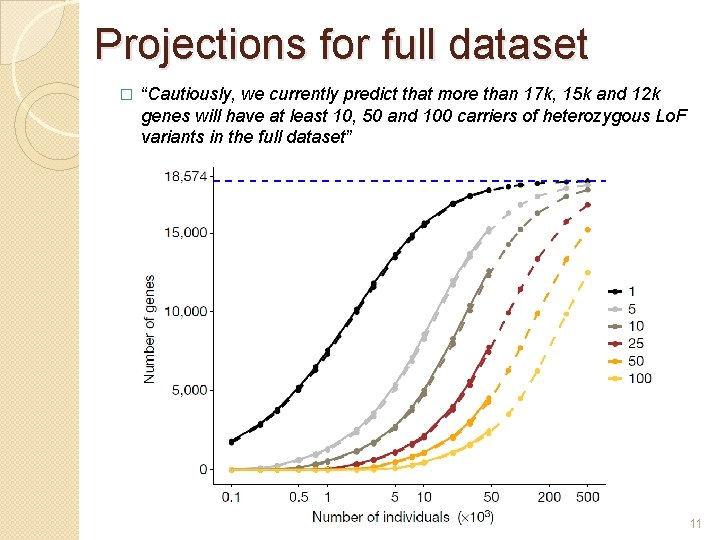
Projections for full dataset � “Cautiously, we currently predict that more than 17 k, 15 k and 12 k genes will have at least 10, 50 and 100 carriers of heterozygous Lo. F variants in the full dataset” 11

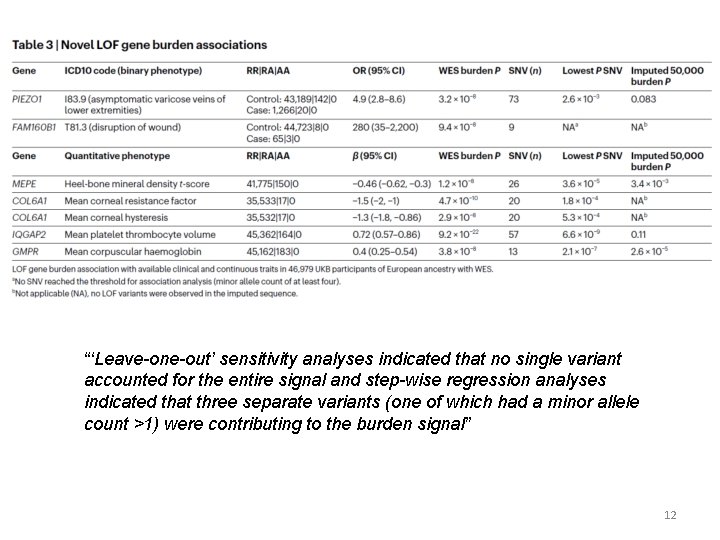
“‘Leave-one-out’ sensitivity analyses indicated that no single variant accounted for the entire signal and step-wise regression analyses indicated that three separate variants (one of which had a minor allele count >1) were contributing to the burden signal” 12


BRCA 1&2 related cancers N= 93 Lo. F variants in BRCA 2 (166 carriers) and 39 Lo. F variants BRCA 1 (59 carriers) 14

Discussion 15

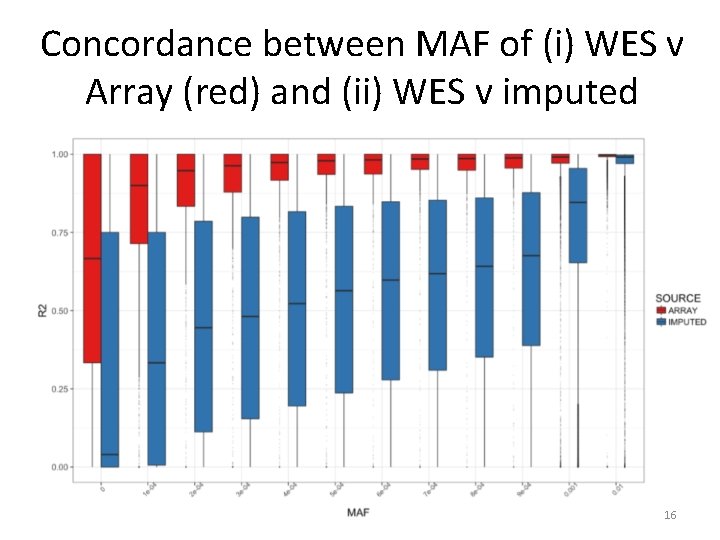
Concordance between MAF of (i) WES v Array (red) and (ii) WES v imputed 16

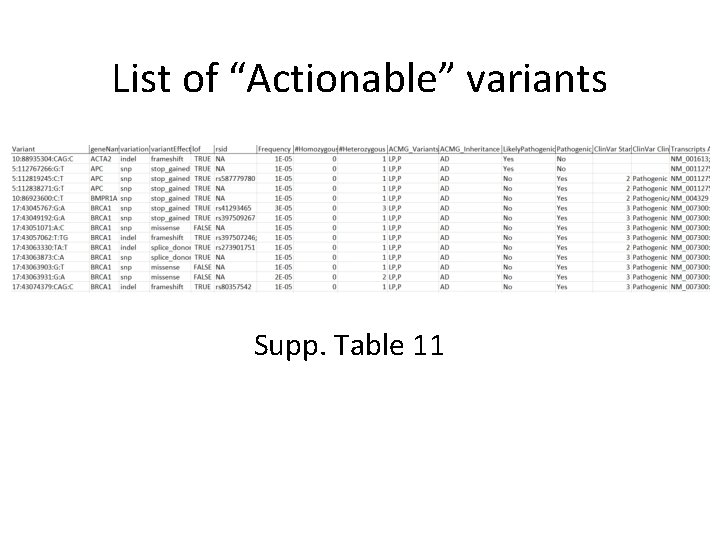
List of “Actionable” variants Supp. Table 11

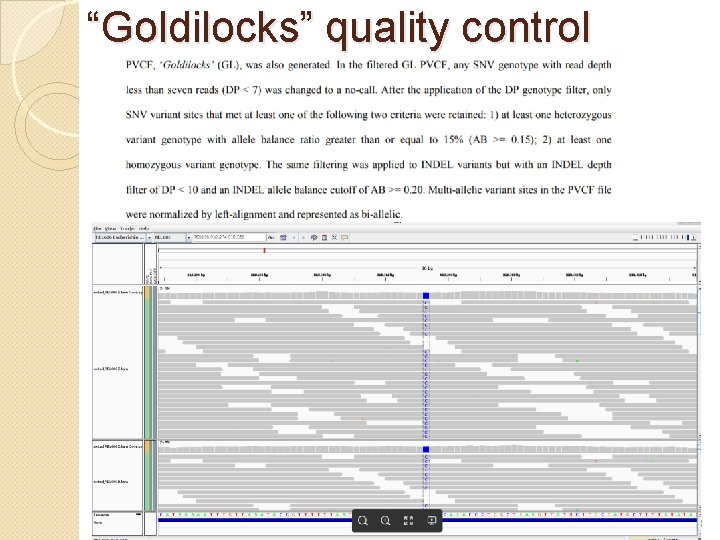
“Goldilocks” quality control 18

Main conclusions � N= 49, 960 high-quality exomes � ~4 million coding variants ◦ coverage >20 x at 94. 6% of sites on average (s. d. 2. 1%) ◦ ~98. 6% have a MAF of <1%. ◦ 198, 269 autosomal predicted loss-of-function (Lo. F) variants �>14 -fold increase in SNVs compared to imputed sequence (16. 1 -fold increase for indels) � 17, 718 (>97% of) genes had 1 or more Lo. F variant and 69% of genes had 10 or more � Association study of 1, 730 phenotypes ◦ PIEZO 1 on varicose veins, COL 6 A 1 on corneal resistance, MEPE on bone density, and IQGAP 2 and GMPR on blood cell traits � Prevalence of pathogenic variants of clinical importance (medically actionable variant) is 2% � Penetrance of BRCA 1&2 variants is lower than previous estimates 19