1 Cloning and Sequencing Explorer Series Bioinformatics 2

1
Cloning and Sequencing Explorer Series Bioinformatics 2

Instructors Stan Hitomi Coordinator – Math & Science Principal – Alamo School San Ramon Valley Unified School District Danville, CA Kirk Brown Lead Instructor, Edward Teller Education Center Science Chair, Tracy High School and Delta College, Tracy, CA Bio-Rad Curriculum and Training Specialists: Sherri Andrews, Ph. D. sherri_andrews@bio-rad. com Essy Levy, M. Sc. essy_levy@bio-rad. com Leigh Brown, M. A. leigh_brown@bio-rad. com 3

Bioinformatics The application of information technology to molecular biology 4

Questions Concerning your Data 5 Class Data Set • Are our sequences high quality? • Are my sequences similar to GAPDH? • Are any of my sequences primarily cloning vector? Individual Clone Sequences • Do my individual sequences align to give me a single long sequence? • Are there discrepancies between my reads? • Which GAPDH gene did we clone? Annotation of Clone Sequence • What is the intron- exon structure/m. RNA sequence of my clone? • What is the protein sequence of my clone?

Sequence Data Analysis Tools Sequence data storage and analysis tools (i. Finch and Finch TV) Sequence comparison algorithm (NCBI BLAST) Sequence Assembly (CAP 3) m. RNA sequence prediction (BLAST and manual) Protein sequence prediction (EMBL-EBI EMBOSS Transeq) 6

Advanced Preparation • Practice with i. Finch using the guest accounthighly recommended! • Activate your i. Finch account (2 months subscription) • Download Finch. TV onto lab computers • Set up project and folder in i. Finch • Upload sequence data 7

Guest i. Finch Account http: //classroom 1. bio-rad. ifinch. com/Finch Username: BR_guest Password: guest • Example data sets for each stage of process • No uploading of data 8
Your own i. Finch account Each account has a unique URL: http: //Platenumber. ifinch. com/Finch E. g. http: //A 150936. ifinch. com/Finch Instructor’s Username: Platenumber e. g. A 150936 Instructor’s Password: Platenumber e. g. A 150936 Student Username: Platenumber_student e. g. A 150936_student Student Password: Platenumber e. g. A 150936 Once activated- change your passwords! Active for 2 months. 9
Download Finch. TV 10 • www. geospiza. com/finchtv
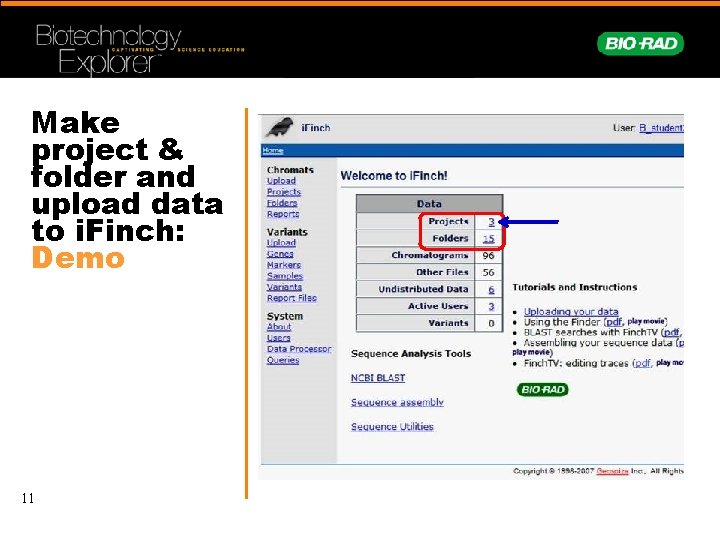
Make project & folder and upload data to i. Finch: Demo 11

Student Activities 1. Review data quality and view sequence traces 2. Use BLAST for preliminary check on which GAPDH was cloned 3. Assemble sequences into a contig 4. Verify which GAPDH gene was cloned 5. Predict intron exon boundaries and generate m. RNA sequence 6. Predict protein sequence 12
Sequence Quality 13
Q 20 values The quality value of a “base call” is: Q= -10 Log 10(Perror) where P is the probability of an error. Thus if the chance that a base call is incorrect is 1/100, P would be 0. 01 and the quality value would be 20 (Q=20). Convention rates sequences by the number of basecalls that have quality values of 20 or higher- a Q 20 value. The quality values of a sequence are calculated automatically by software in i. Finch- a common program for this was developed by the University of Washington and is called “Phred” 14
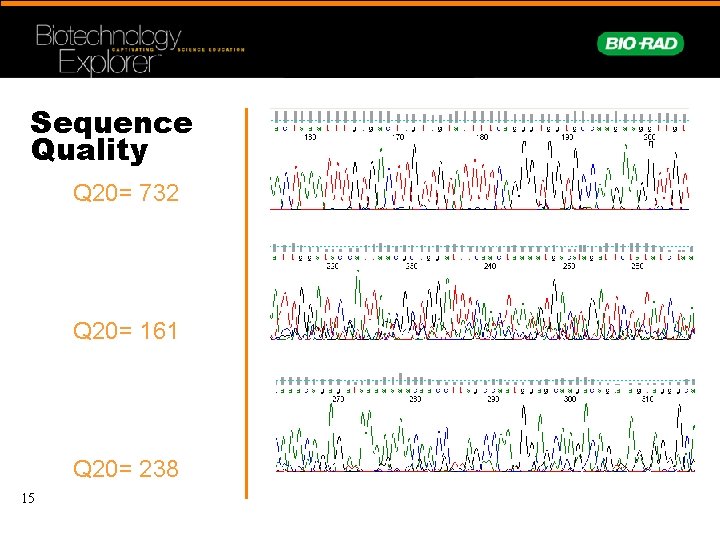
Sequence Quality Q 20= 732 Q 20= 161 Q 20= 238 15

Screen for poor quality sequence, vector, GAPDH family 17
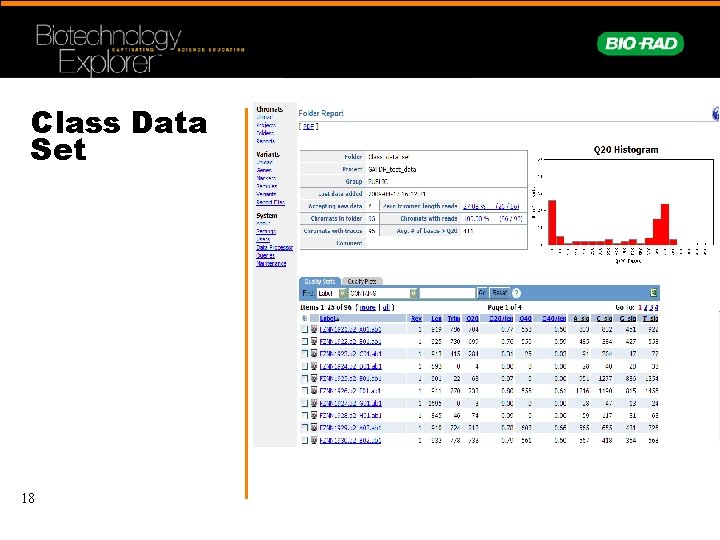
Class Data Set 18

Sort Class Data into Folders 19

Record Data Information 20
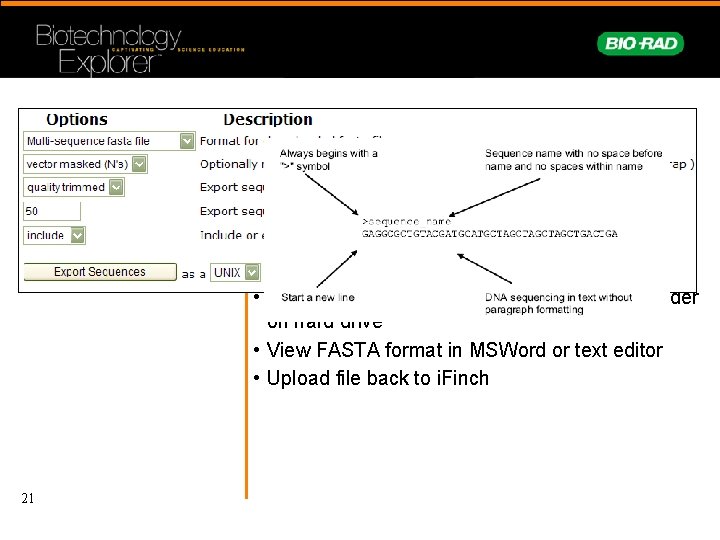
Download sequences for initial screen using BLAST 21 • Open Guest i. Finch account – User: BR-guest, Psw: guest • Click : Folders • Click : Salvia folder • Look at data • Go back to folder report • Click: Download folder data- save to new folder on hard drive • View FASTA format in MSWord or text editor • Upload file back to i. Finch
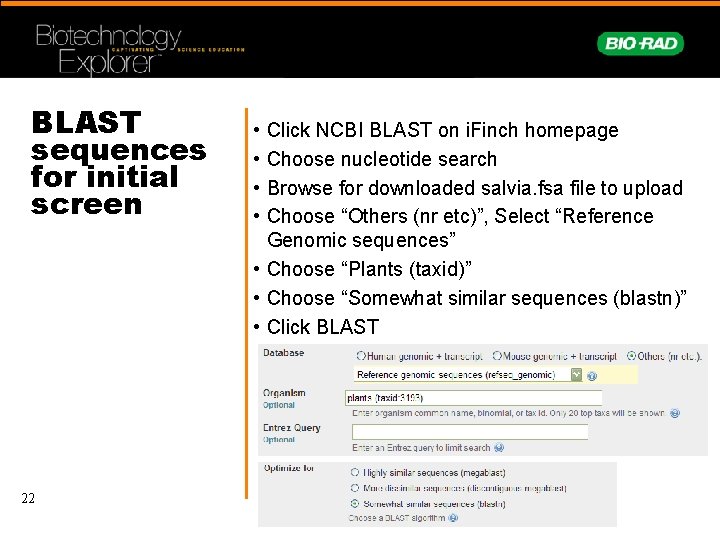
BLAST sequences for initial screen 22 • • Click NCBI BLAST on i. Finch homepage Choose nucleotide search Browse for downloaded salvia. fsa file to upload Choose “Others (nr etc)”, Select “Reference Genomic sequences” • Choose “Plants (taxid)” • Choose “Somewhat similar sequences (blastn)” • Click BLAST

BLAST Results 23 • All 4 sequences were analyzed by BLASTchoose from pull down menu at top of page • Mouse over top bar • Scroll down to list of homologous sequences – E value represents the number of equally good sequence matches to the query sequence that would be expected in a database of the same size containing random sequences. • Scroll down to sequence alignments – Query: Your sequence – Subject: Database matching sequence
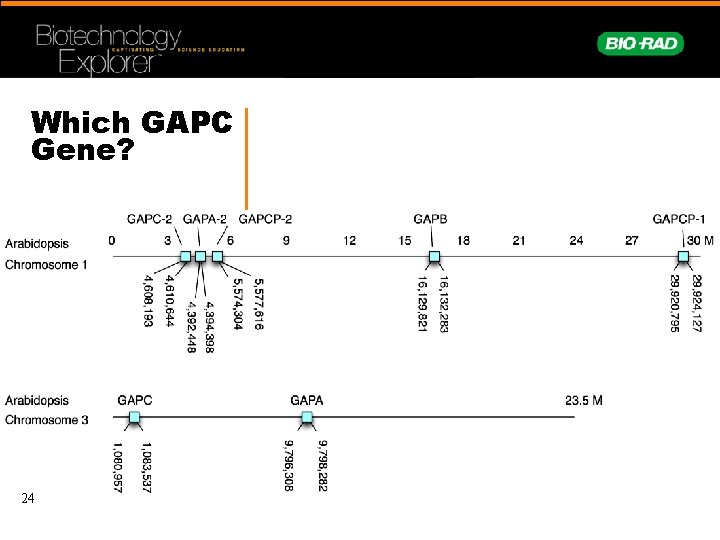
Which GAPC Gene? 24

Break time! 25

Questions Concerning your Data 26 Class Data Set • Are our sequences high quality? • Are my sequences similar to GAPDH? • Are any of my sequences primarily cloning vector? Individual Clone Sequences • Do my individual sequences align to give me a single long sequence? • Are there discrepancies between my reads? • Which GAPDH gene did we clone? Annotation of Clone Sequence • What is the intron- exon structure/m. RNA sequence of my clone? • What is the protein sequence of my clone?
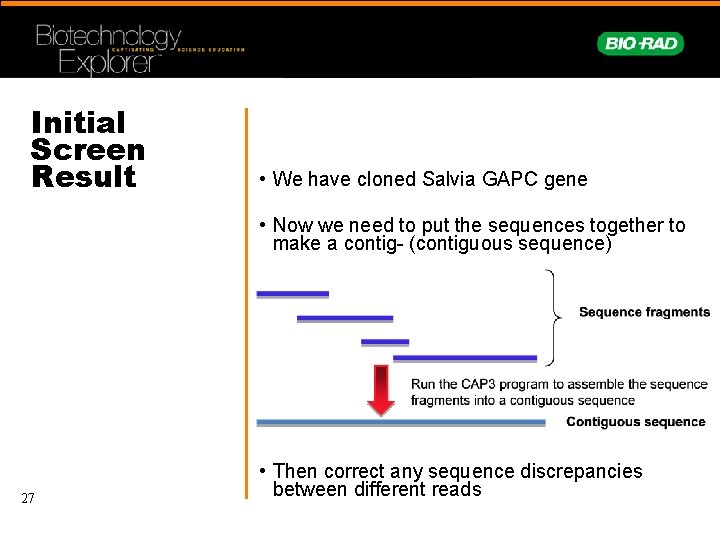
Initial Screen Result • We have cloned Salvia GAPC gene • Now we need to put the sequences together to make a contig- (contiguous sequence) 27 • Then correct any sequence discrepancies between different reads
CAP 3 Program (Contig Assembly Program) 28 • On i. Finch home page click “sequence assembly”

Assembly Results • Your sequence file (your input) • Single sequences (any seqs that could not be assembled) • Contigs (save in FASTA format as “. txt” file) • Assembly details (Save as landscape “. txt file) 29
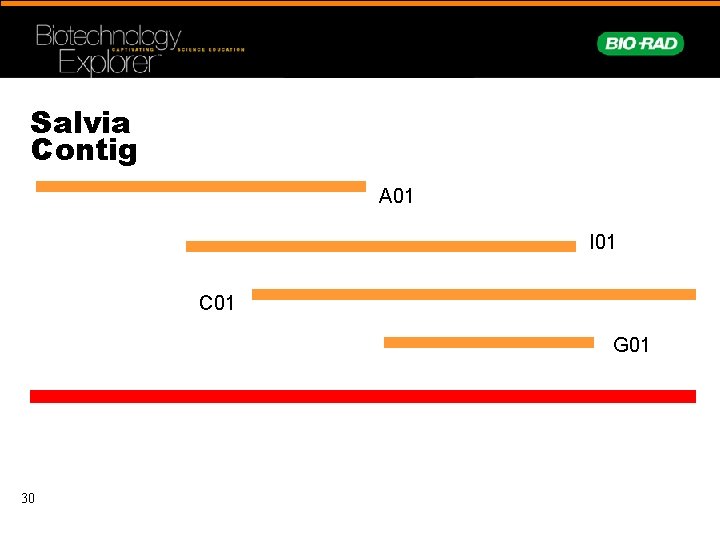
Salvia Contig A 01 I 01 C 01 G 01 30

Check for Discrepancies • Look through assembly file for sequence discrepancies • Open chromatogram files in Finch. TV • Examine actual chromatograms and use personal judgment to determine which base call is correct • Correct Finch. TV file and save back to i. Finch (not available in guest account) noting the changes in the revision history 31 • If the consensus sequence has changed, download folder sequences again like previously and reassemble with CAP 3 program

BLAST search with contig 32 Submit contig FASTA file for BLAST search (same database as before- plant reference genomic database)

Break time! 33

Questions Concerning your Data 34 Class Data Set • Are our sequences high quality? • Are my sequences similar to GAPDH? • Are any of my sequences primarily cloning vector? Individual Clone Sequences • Do my individual sequences align to give me a single long sequence? • Are there discrepancies between my reads? • Which GAPDH gene did we clone? Annotation of Clone Sequence • What is the intron- exon structure/m. RNA sequence of my clone? • What is the protein sequence of my clone?
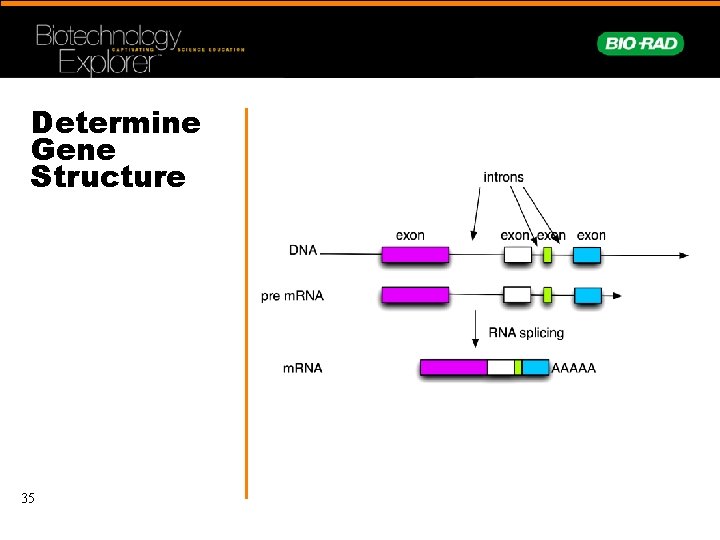
Determine Gene Structure 35
Workflow 36
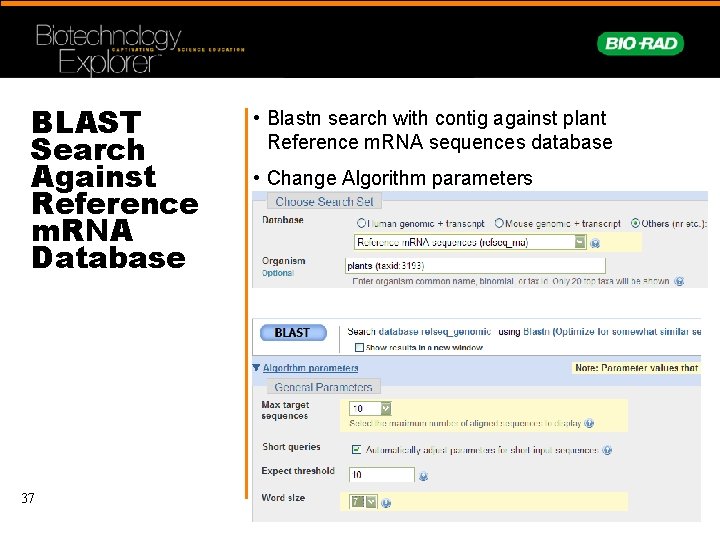
BLAST Search Against Reference m. RNA Database 37 • Blastn search with contig against plant Reference m. RNA sequences database • Change Algorithm parameters
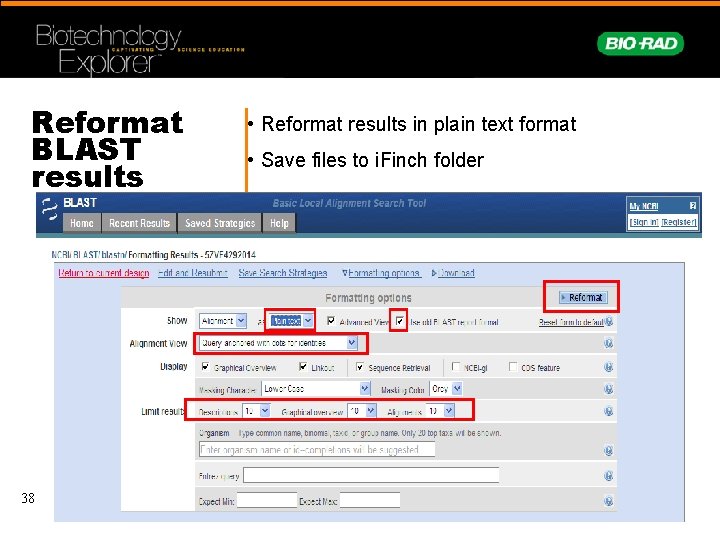
Reformat BLAST results 38 • Reformat results in plain text format • Save files to i. Finch folder

Save Contig File in MSWord • Delete all paragraph marks using find and replace command • Save to hard drive as “. rtf” file with a new name. • Color contig sequence with exons as determined from BLAST results • Put exons together in a first draft of the m. RNA sequence and save to i. Finch folder • Submit draft m. RNA sequence to blastn against plant reference m. RNA database 39
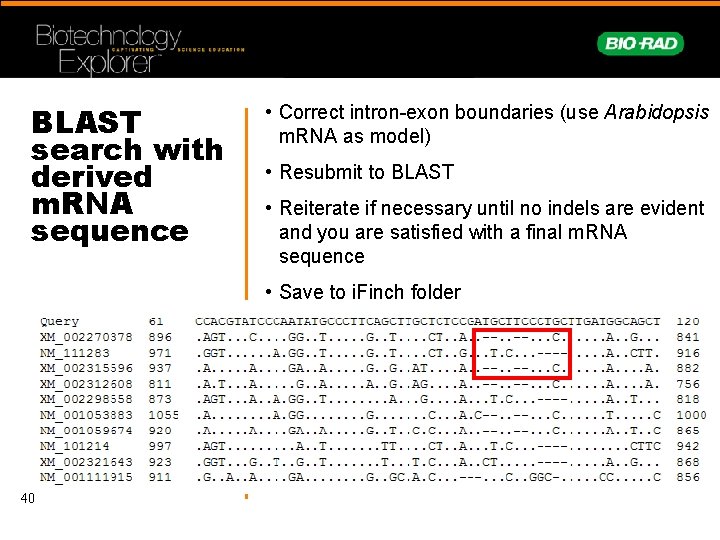
BLAST search with derived m. RNA sequence • Correct intron-exon boundaries (use Arabidopsis m. RNA as model) • Resubmit to BLAST • Reiterate if necessary until no indels are evident and you are satisfied with a final m. RNA sequence • Save to i. Finch folder 40
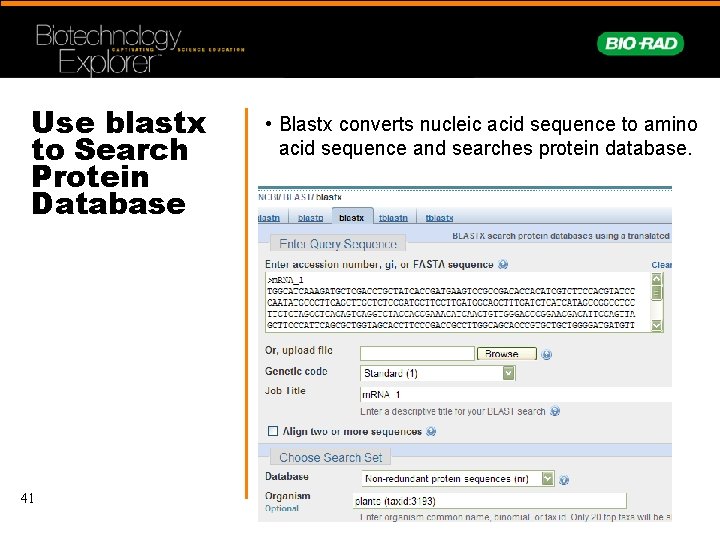
Use blastx to Search Protein Database 41 • Blastx converts nucleic acid sequence to amino acid sequence and searches protein database.
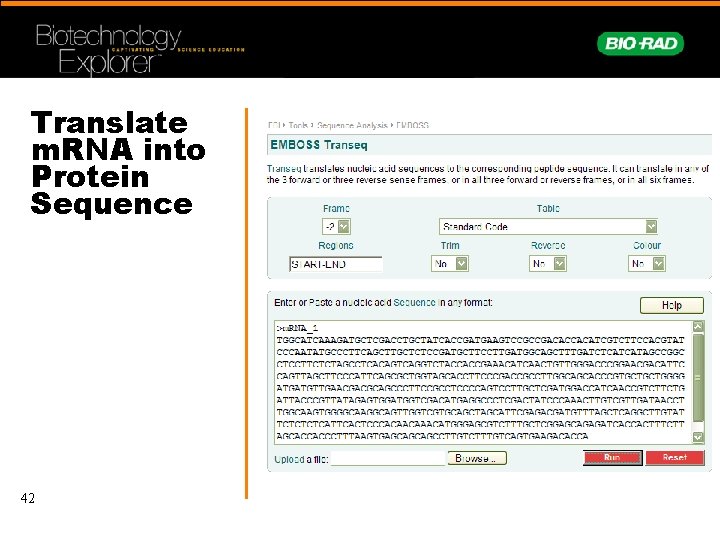
Translate m. RNA into Protein Sequence 42
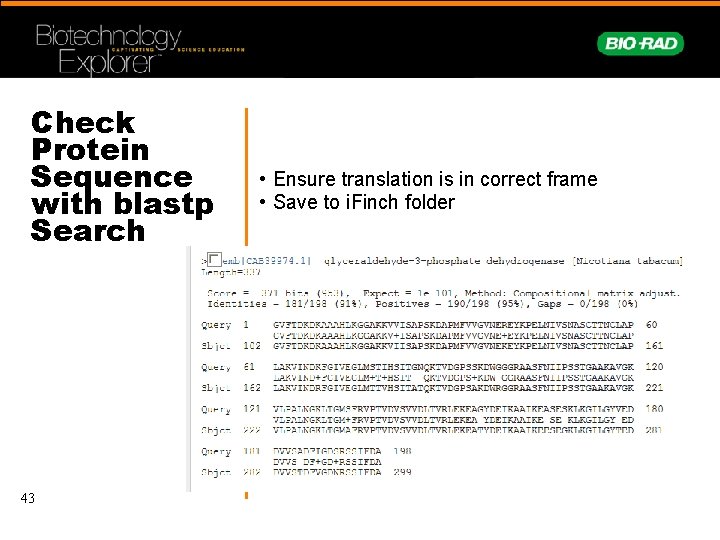
Check Protein Sequence with blastp Search 43 • Ensure translation is in correct frame • Save to i. Finch folder
Congratulations! • You have cloned, sequenced annotated a novel gene. • You could now submit this to Gen. Bank. • Data from additional samples would strengthen the data- for example- assemble sequences from the same gene from different student teams • Download data from i. Finch if you wish to keep it for the long term 44

Webinars • Enzyme Kinetics — A Biofuels Case Study • Real-Time PCR — What You Need To Know and Why You Should Teach It! • Proteins — Where DNA Takes on Form and Function • From plants to sequence: a six week college biology lab course • From singleplex to multiplex: making the most out of your realtime experiments explorer. bio-rad. com Support Webinars 45
- Slides: 44