Chromosome inversions in human populations Marta Ruiz Fernndez











- Slides: 23

Chromosome inversions in human populations Marta Ruiz Fernández Master in Advanced Genetics 17 December 2014

INTRODUCTION • Inversions occur when a chromosome breaks at two points and the segment between the breakpoints is reinserted in the reverse orientation. • Do not change the amount of genetic material Generally viable and show no particular abnormalities at phenotypic level

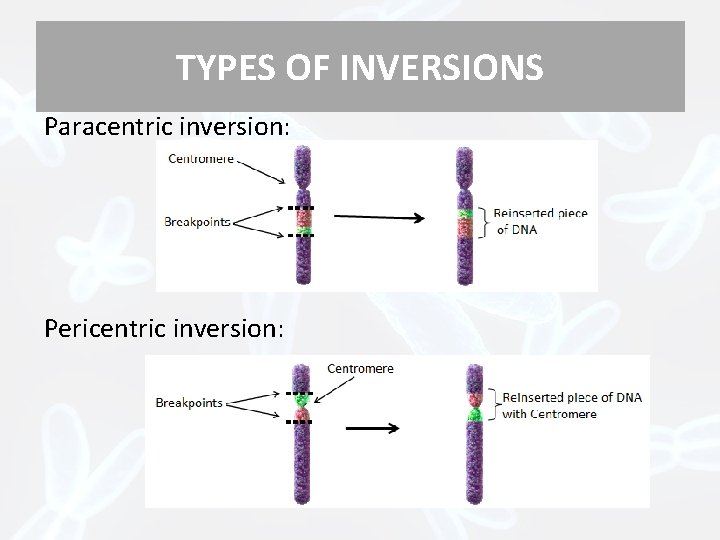
TYPES OF INVERSIONS Paracentric inversion: Pericentric inversion:

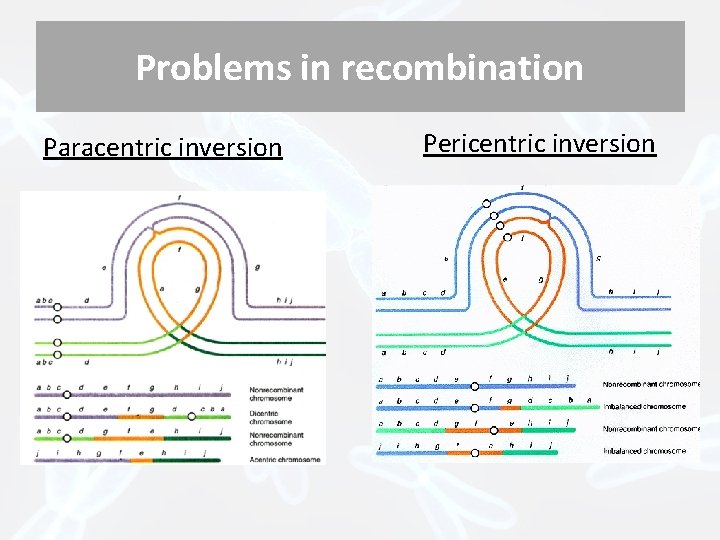
Problems in recombination Paracentric inversion Pericentric inversion

MECHANISMS OF INVERSION • Double strand break is needed for the inversion • “Hotspots” – Usually contains segmental duplications (SD) – Distance between SD = 10 -400 kb – 95 – 97% identity • Mechanisms – Non-allelic homologous recombination (NAHR) – Non-homologous end joining (NHEJ) – Fork stalling and template switching

CONSEQUENCES OF INVERSIONS • Recombination suppressed in heterozygotes – Reducing cross-over within inverted regions – If recombination occurs Not viable cells • Inversions could lead to insertions or deletions • Depending on the breakpoints position: – – Gene disruption Alter gene expression Generate new splicing sites Gene fusion NEUTRAL DELETERIOUS ADAPTATIVE

HOW TO DETECT INVERSIONS • Problems – Inversions do not change genome size – The genome is full of GAPs and the inversions flanked by repetitive sequences International Human Genome Sequencing Consortium (2004) Nature 409: 931 -945.

HOW TO DETECT INVERSIONS • Problems – Inversions do not change genome size – The genome is full of GAPs and the inversions flanked by repetitive sequences • Methods – Cytogenetic (conventional, FISH-based assay) – Pair-end sequencing – Genome assembly

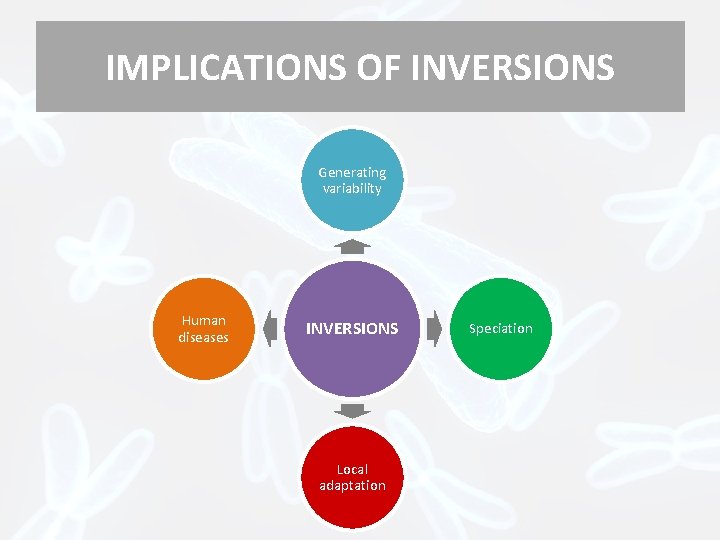
IMPLICATIONS OF INVERSIONS Generating variability Human diseases INVERSIONS Local adaptation Speciation

The role of inversions in human diversity • Hap. Map project Structural Variations

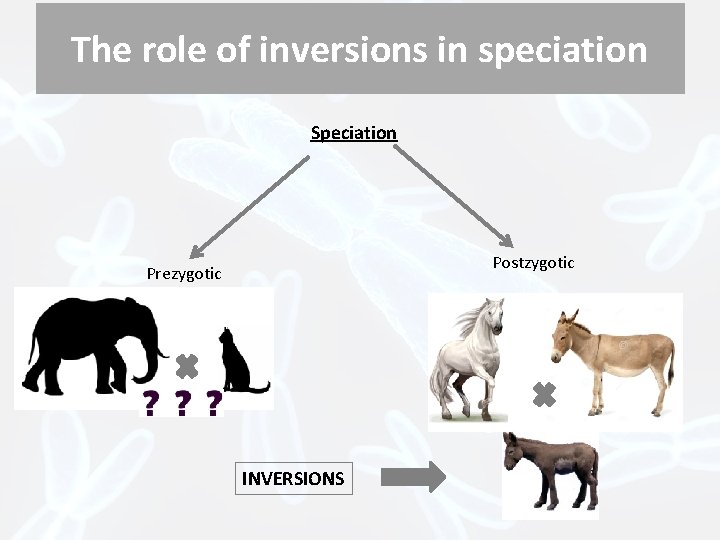
The role of inversions in speciation Speciation Postzygotic Prezygotic INVERSIONS

The role of inversions in local adaptation -Annual -Found in DRY environments in the INLAND -Flowering time BEFORE hot -Perennial -Found in WET and COOL environments in the coast -Flowering time LATER Monkeyflower ( Mimulus guttatus) - Flowering time - Morphological differences

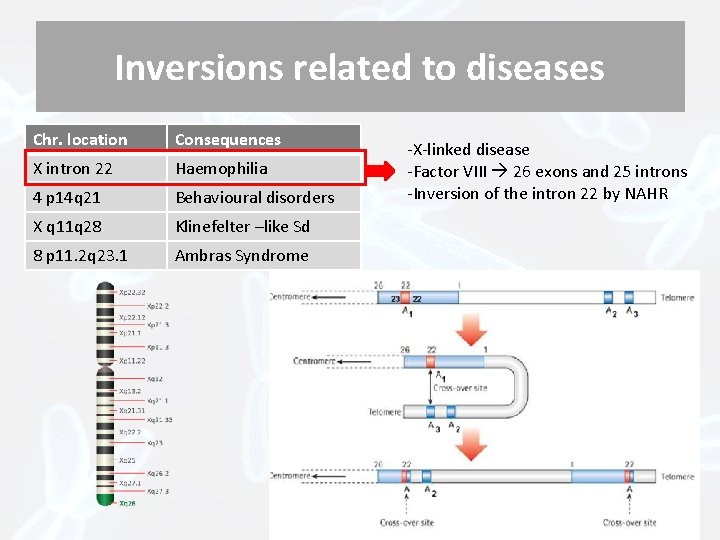
Inversions related to diseases Chr. location Consequences X intron 22 Haemophilia 4 p 14 q 21 Behavioural disorders X q 11 q 28 Klinefelter –like Sd 8 p 11. 2 q 23. 1 Ambras Syndrome -X-linked disease -Factor VIII 26 exons and 25 introns -Inversion of the intron 22 by NAHR

CONCLUSIONS • Several studies have reported many inversions • Way to generate variation between and within species • Diversity of effects depending on: – Where? (Coding/Non-coding, introns, exons) – Is any gene affected? – Does the inversion imply another chromosomal arrangement?

REFERENCES • Antonacci F, Kidd JM, Marques-Bonet T, Ventura M, Siswara P, Jiang Z, et al. Characterization of six human disease-associated inversion polymorphisms. Hum Mol Genet. 2009; 18: 2555– 66. • Gu W, Zhang F, Lupski JR. Mechanisms for human genomic rearrangements. Pathogenetics. 2008; 1: 4. • Bansal V, Bashir A, Bafna V. Evidence for large inversion polymorphisms in the human genome from Hap. Map data 2007: 219– 30. • Lee J, Han K, Meyer TJ, Kim H-S, Batzer M a. Chromosomal inversions between human and chimpanzee lineages caused by retrotransposons. PLo. S One. 2008; 3: e 4047. • Kirkpatrick M. How and why chromosome inversions evolve. PLo. S Biol. 2010; 8. • Lowry DB, Willis JH. A widespread chromosomal inversion polymorphism contributes to a major life-history transition, local adaptation, and reproductive isolation. PLo. S Biol. 2010; 8.

Chromosome inversions in human populations Marta Ruiz Fernández Master in Advanced Genetics 17 December 2014


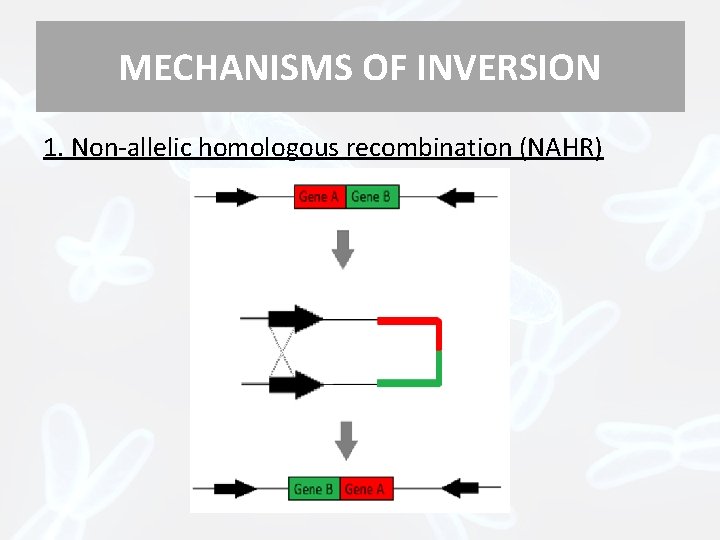
MECHANISMS OF INVERSION 1. Non-allelic homologous recombination (NAHR)

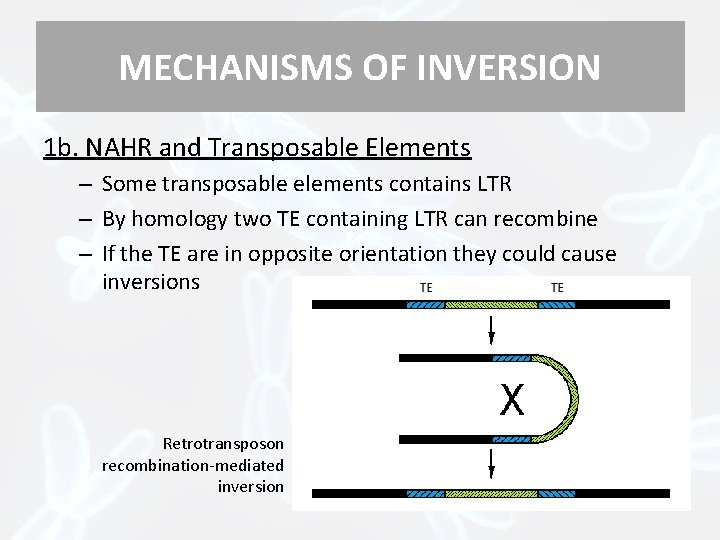
MECHANISMS OF INVERSION 1 b. NAHR and Transposable Elements – Some transposable elements contains LTR – By homology two TE containing LTR can recombine – If the TE are in opposite orientation they could cause inversions Retrotransposon recombination-mediated inversion

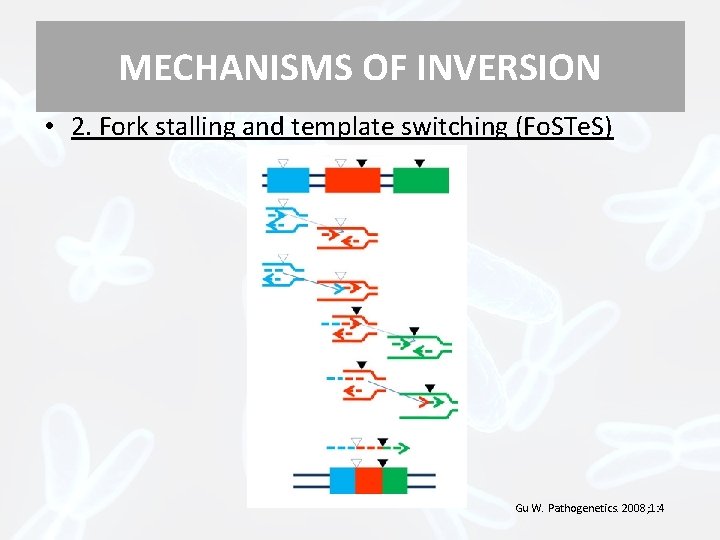
MECHANISMS OF INVERSION • 2. Fork stalling and template switching (Fo. STe. S) Gu W. Pathogenetics. 2008; 1: 4

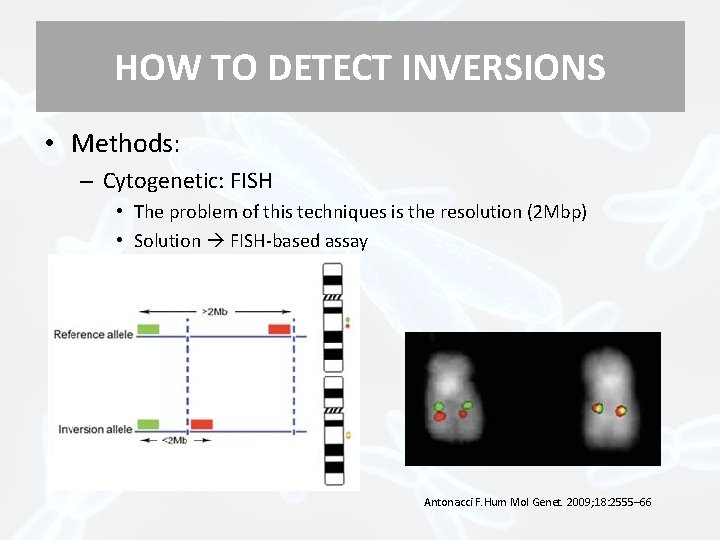
HOW TO DETECT INVERSIONS • Methods: – Cytogenetic: FISH • The problem of this techniques is the resolution (2 Mbp) • Solution FISH-based assay Antonacci F. Hum Mol Genet. 2009; 18: 2555– 66

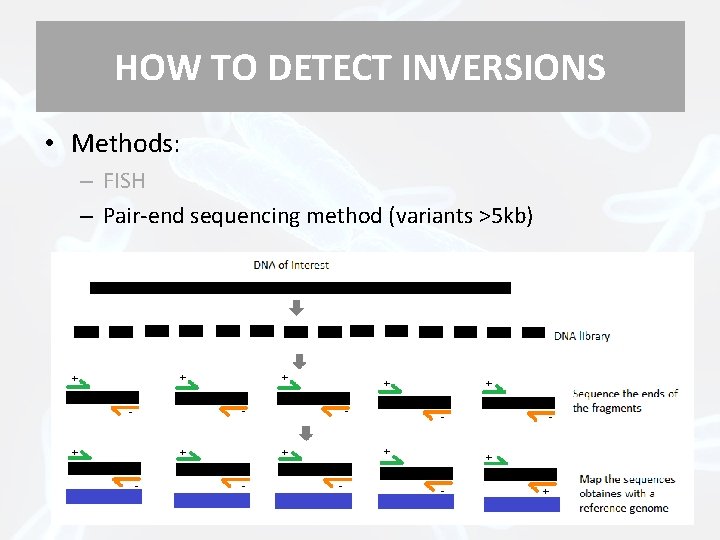
HOW TO DETECT INVERSIONS • Methods: – FISH – Pair-end sequencing method (variants >5 kb) + + - + - +

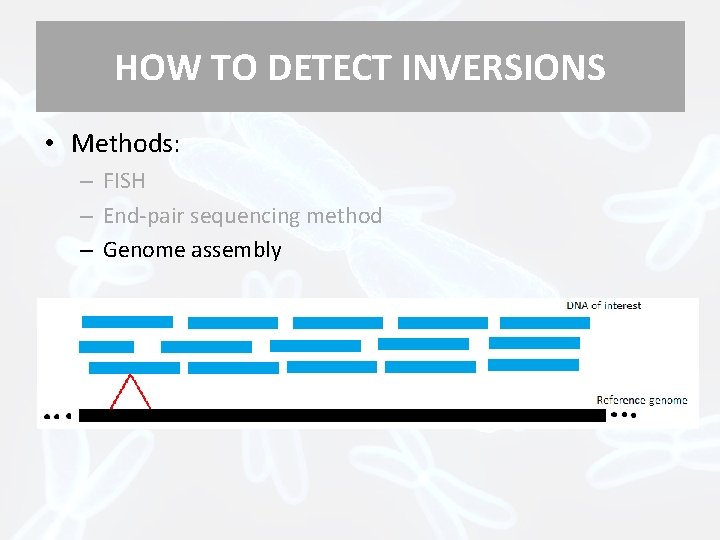
HOW TO DETECT INVERSIONS • Methods: – FISH – End-pair sequencing method – Genome assembly