BLAST What is BLAST Basic Local Alignment Search

BLAST

What is BLAST? • Basic Local Alignment Search Tool • Calculates similarity for biological sequences. • Produces local alignments: only a portion of each sequence must be aligned. • Uses statistical theory to determine if a match might have occurred by chance. 2

BLAST is a heuristic. • A lookup table is made of all the “words” (short subsequences) and “neighboring” words in the query sequence. • The database is scanned for matching words (“hot spots”). • Gapped and un-gapped extensions are initiated from these matches. 3

4
BLAST reports at the NCBI Web page. 5
Formatting Page 6
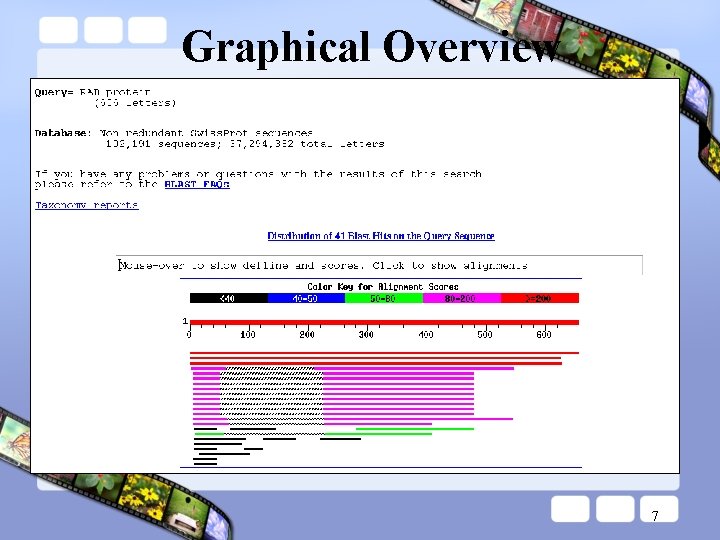
Graphical Overview 7
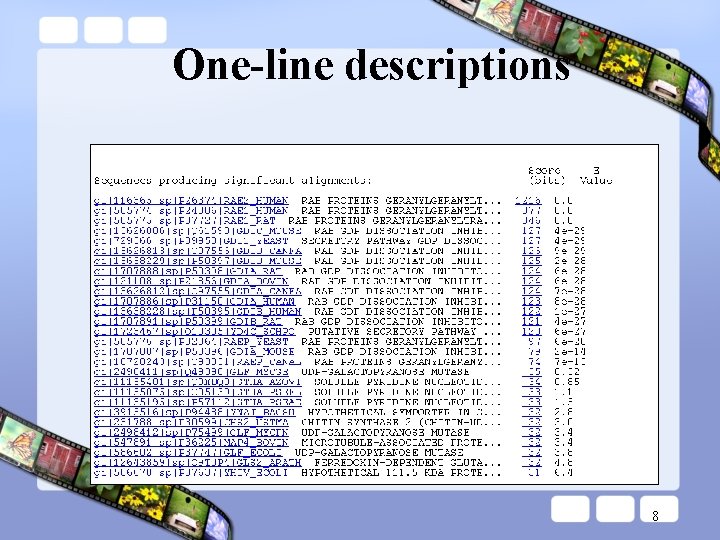
One-line descriptions 8

Expect value • The number of alignments expected by chance with a given score. • The larger the database, the more alignments are expected at a given score • “Errors per Query” or “FALSE Positive rate”. 9

Pair-wise alignments 10

BLOSUM 62 matrix 11
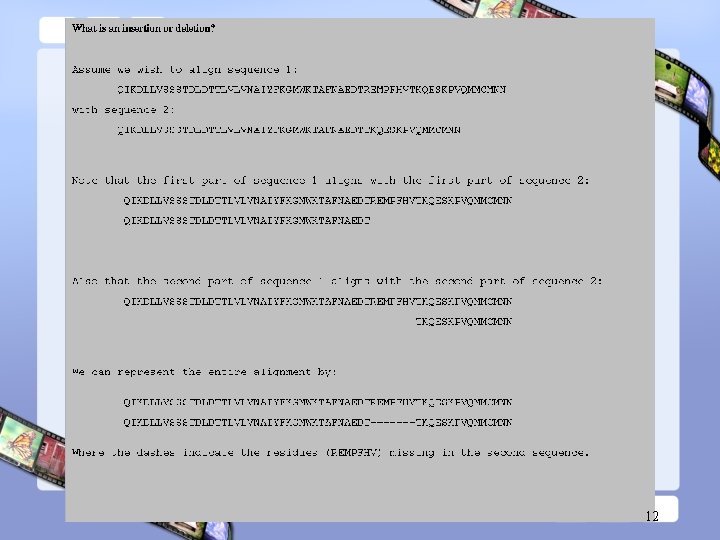
12

Query-anchored alignments 13

Future improvements: Link. Out, taxonomic and structure links. Link to Locus-link Link to Uni. Gene Link to taxonomy 14

BLAST report designed for human readability. • One-line descriptions provide overview designed for human “browsing”. • Redundant information is presented in the report (e. g. , one-line descriptions and alignments both contain expect values, scores, descriptions) so a user does not need to move back and forth between sections. • HTML version has lots of links for a user to explore. • It can change as new features/information becomes available. 15
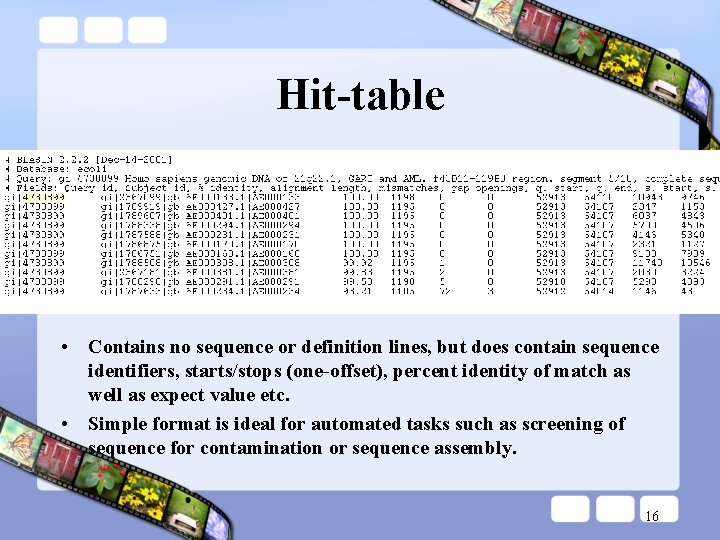
Hit-table • Contains no sequence or definition lines, but does contain sequence identifiers, starts/stops (one-offset), percent identity of match as well as expect value etc. • Simple format is ideal for automated tasks such as screening of sequence for contamination or sequence assembly. 16

17
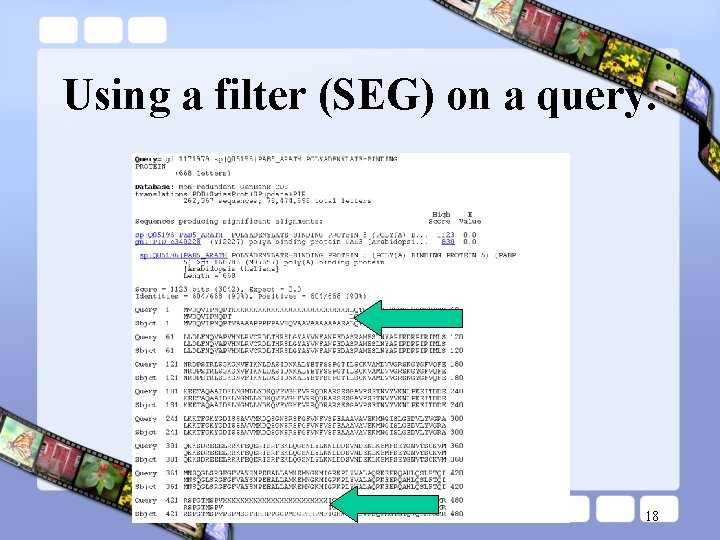
Using a filter (SEG) on a query. 18

What is an ALU • Constitutes about 5% of the human genome. • Short interspersed repeats. • Found in primate genomes. • ALU elements often found in 3’ regions or introns. 19
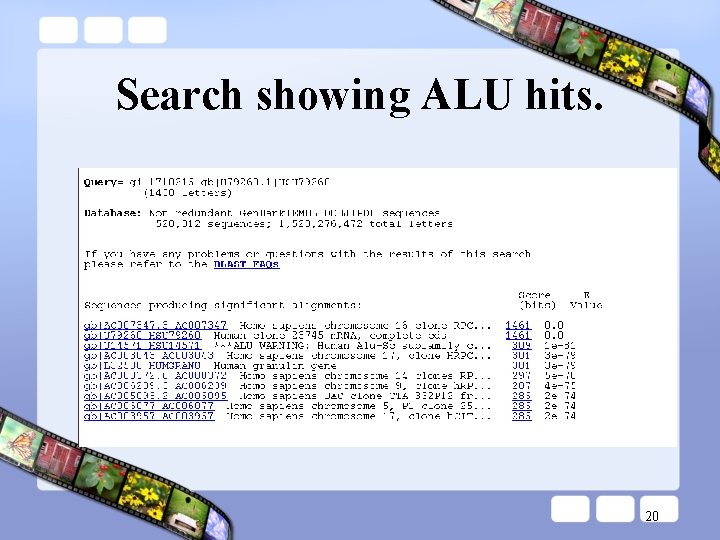
Search showing ALU hits. 20

21

Identifying ALU contaminated regions. • ALU BLAST database on the NCBI Web page. • Repeat Masker: http: //ftp. genome. washington. edu/cgibin/Repeat. Masker 22

Removing ALU contamination • Human repeat filtering on BLAST Web pages. • Repeat. Masker: http: //ftp. genome. washington. edu/cgibin/Repeat. Masker 23

PSI-BLAST • A normal BLASTP (protein-protein) run is performed. • A position-dependent matrix is built using the most significant matches to the database. • The search is rerun using this profile. • The cycle may be repeated until convergence. • The result is a ‘matrix’ tailored to the query. 24
- Slides: 24