A Transfer Learning Model for Identification of Biomarkers

A Transfer Learning Model for Identification of Biomarkers in Chest X-Rays of COVID 19 Patients Mirtha Lucas Advisors: Dr Daniela Raicu, Dr. Jacob Furst May, 2020
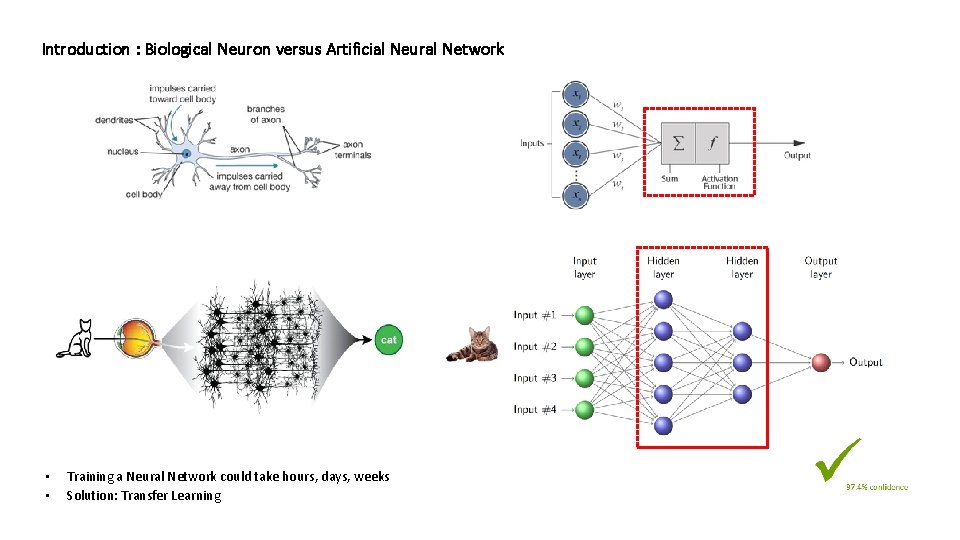
Introduction : Biological Neuron versus Artificial Neural Network • • Training a Neural Network could take hours, days, weeks Solution: Transfer Learning
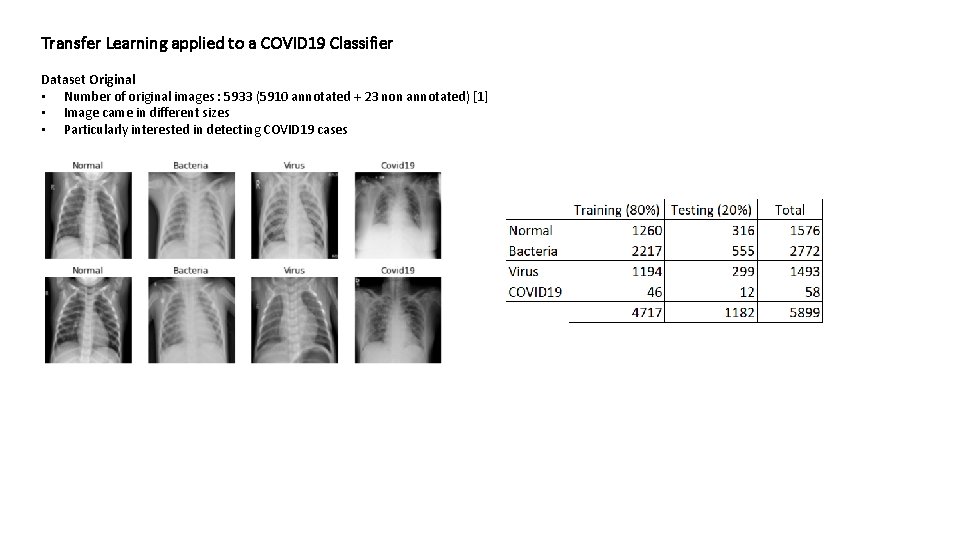
Transfer Learning applied to a COVID 19 Classifier Dataset Original • Number of original images : 5933 (5910 annotated + 23 non annotated) [1] • Image came in different sizes • Particularly interested in detecting COVID 19 cases

Transfer Learning applied to a COVID 19 Classifier Approach • NN with Transfer Learning technique • VGG 19 has 19 (trainable) layers, pretrained on Image. Net images and able to classify images into 1000 categories [2] • We use the base model of VGG 19, containing 16 (trainable) layers, last 3 layers were removed
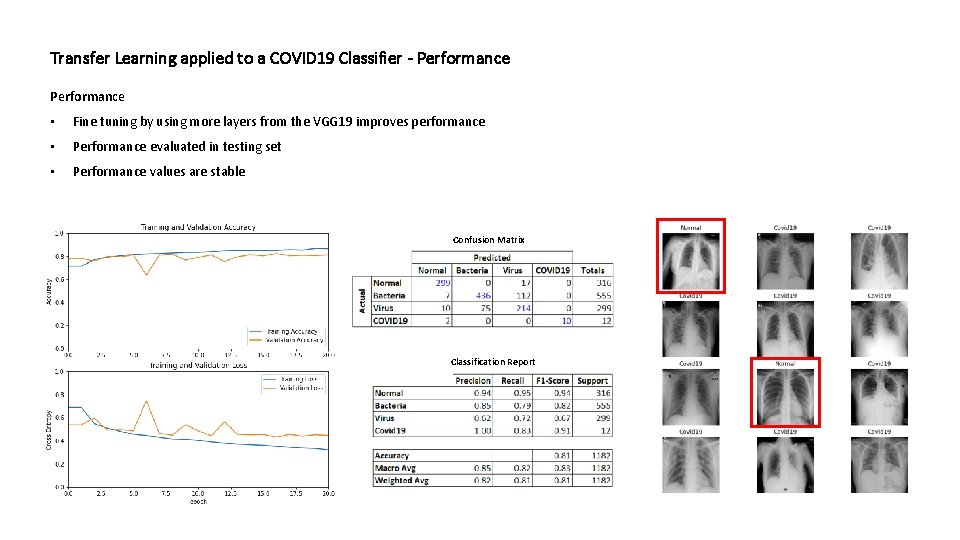
Transfer Learning applied to a COVID 19 Classifier - Performance • Fine tuning by using more layers from the VGG 19 improves performance • Performance evaluated in testing set • Performance values are stable Confusion Matrix Classification Report
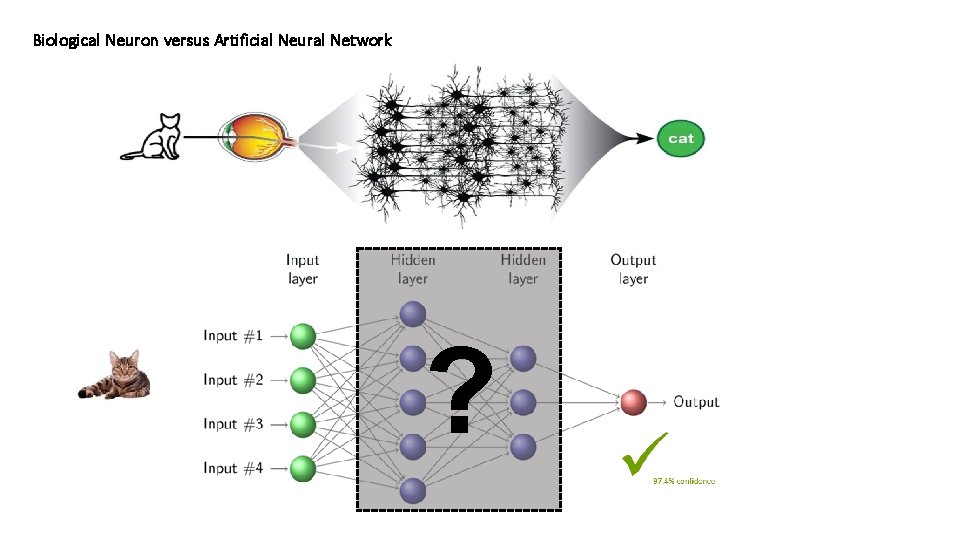
Biological Neuron versus Artificial Neural Network ?
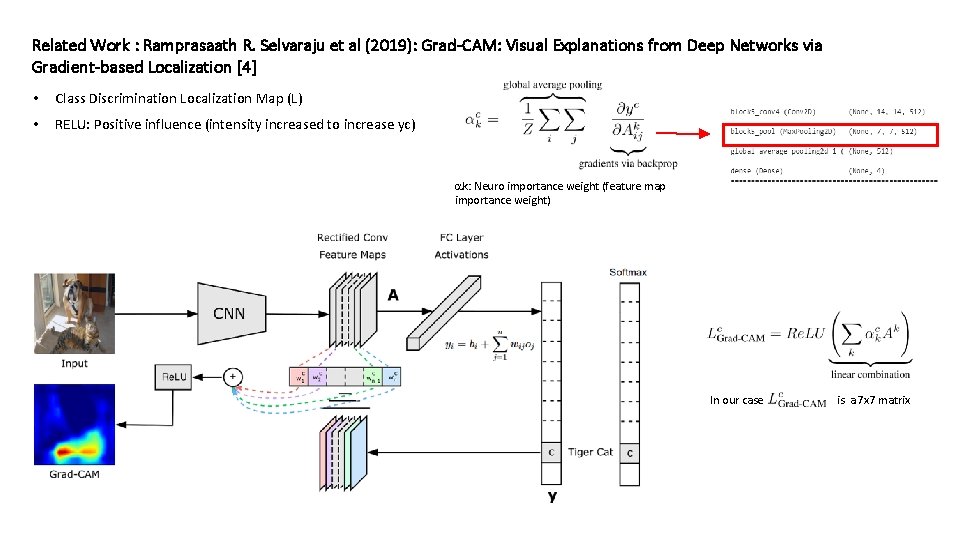
Related Work : Ramprasaath R. Selvaraju et al (2019): Grad-CAM: Visual Explanations from Deep Networks via Gradient-based Localization [4] • Class Discrimination Localization Map (L) • RELU: Positive influence (intensity increased to increase yc) k: Neuro importance weight (feature map importance weight) In our case: Lgrad-CAM is a 7 x 7 matrix
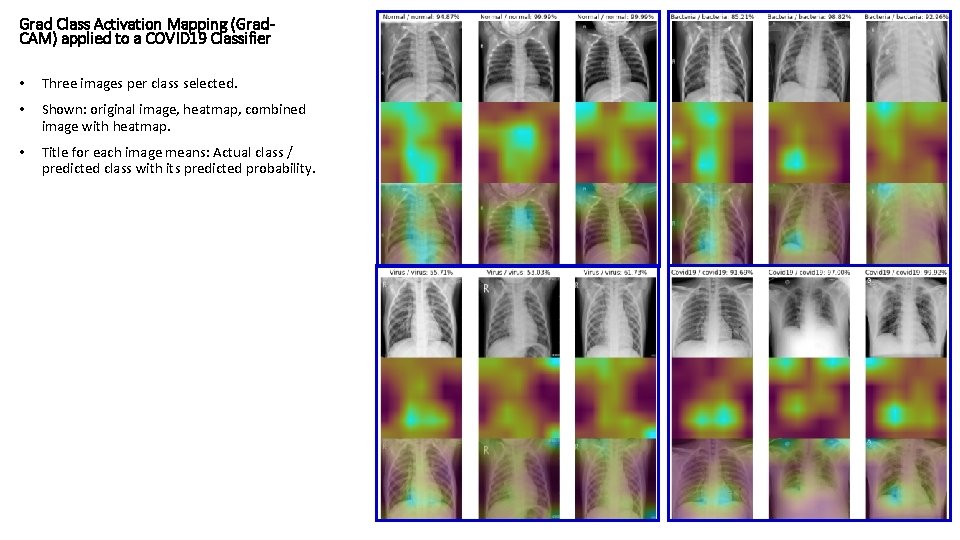
Grad Class Activation Mapping (Grad. CAM) applied to a COVID 19 Classifier • Three images per class selected. • Shown: original image, heatmap, combined image with heatmap. • Title for each image means: Actual class / predicted class with its predicted probability.
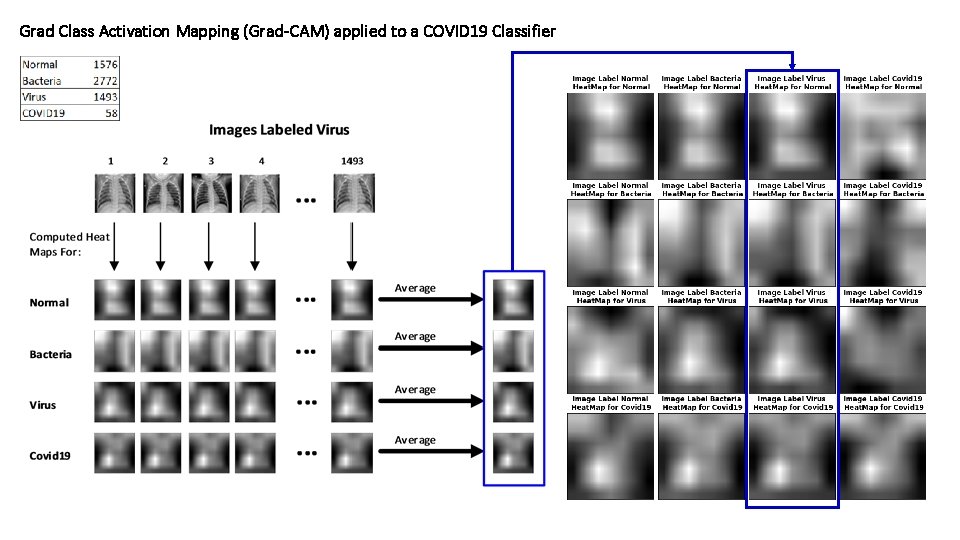
Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier

Average. HM Covid 19 Average. HM Virus Average. HM Bacteria Average. HM Normal Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier • • • Average the gray scale heat maps per each class, along the images of each class. Region of interest for each class The color mosaic indicates regions where each class pays more attention than the rest in order to make a prediction.
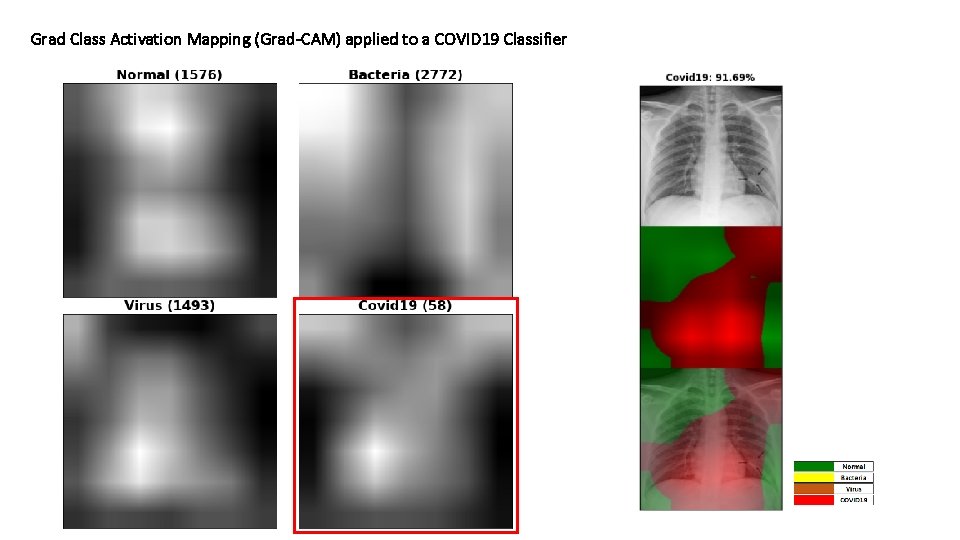
Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier
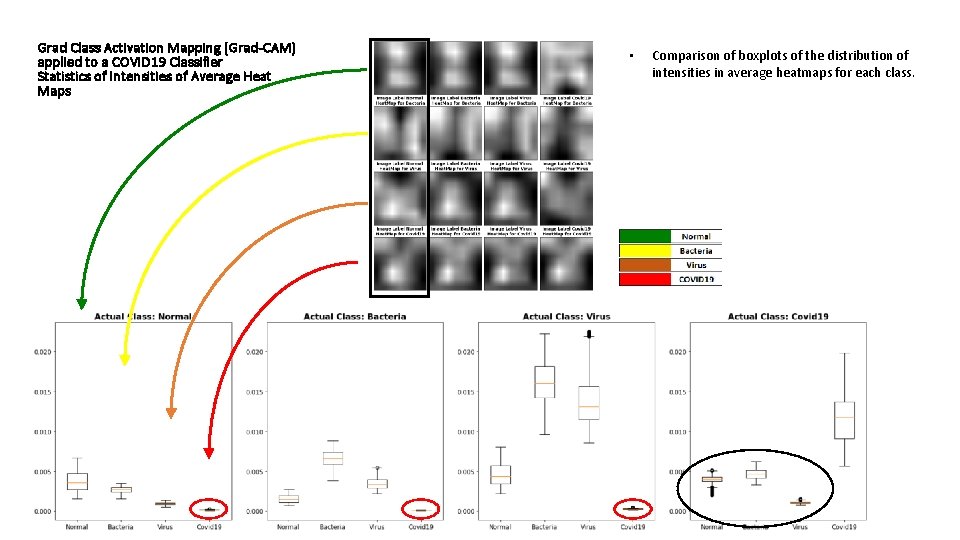
Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier Statistics of Intensities of Average Heat Maps • Comparison of boxplots of the distribution of intensities in average heatmaps for each class.
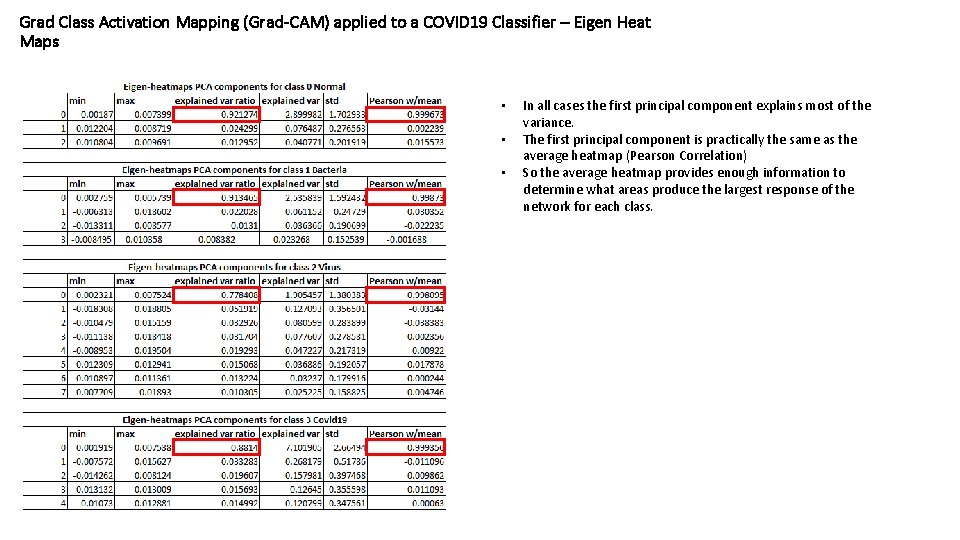
Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier – Eigen Heat Maps • • • In all cases the first principal component explains most of the variance. The first principal component is practically the same as the average heatmap (Pearson Correlation) So the average heatmap provides enough information to determine what areas produce the largest response of the network for each class.

Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier – Eigen Heat Maps vs Average Heat Map Eigen Heatmaps COVID 19 Virus Bacteria Normal PC 1 PC 2 PC 3 • • The first PC are very similar to the average heat maps. The following PC don’t show much variation, although still contribute to the final heatmap (minimally) Average Gray Scale Heatmaps
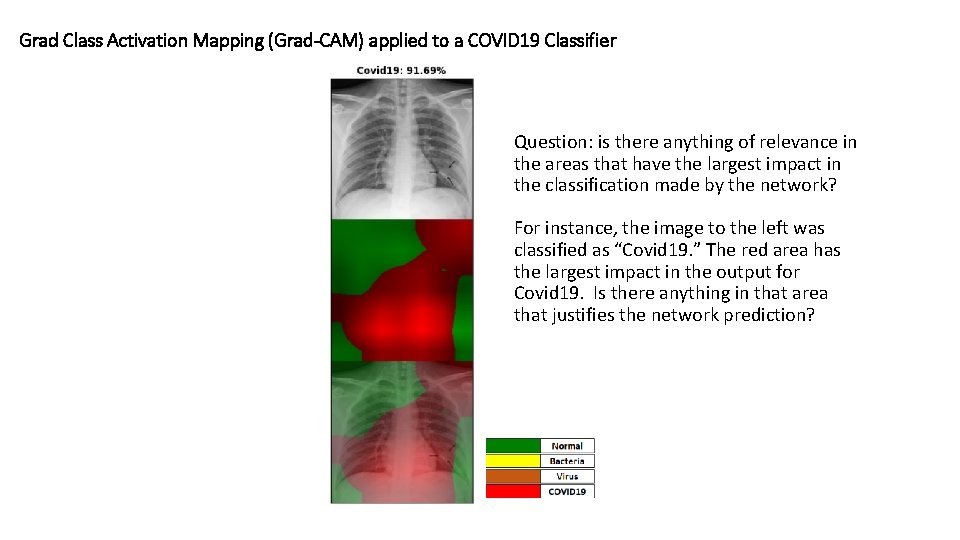
Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier Question: is there anything of relevance in the areas that have the largest impact in the classification made by the network? For instance, the image to the left was classified as “Covid 19. ” The red area has the largest impact in the output for Covid 19. Is there anything in that area that justifies the network prediction?
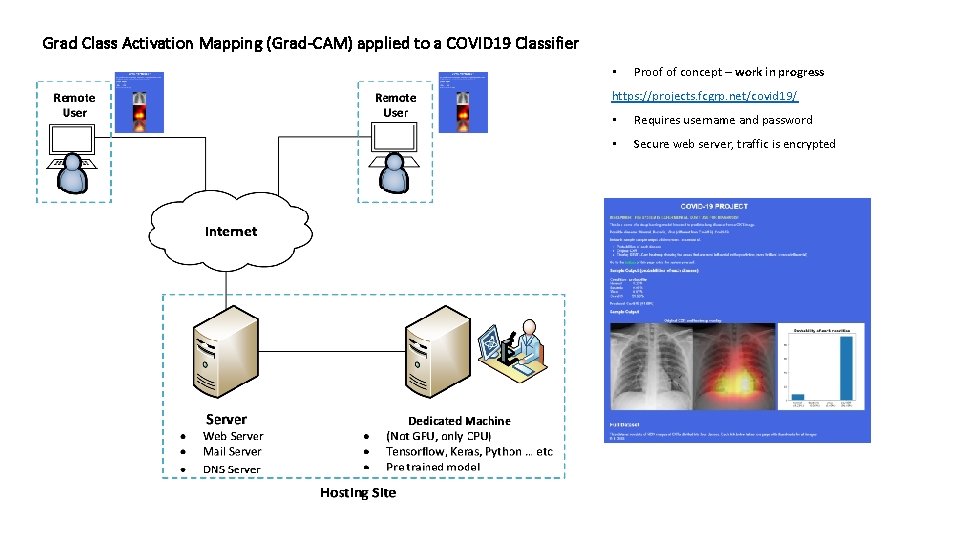
Grad Class Activation Mapping (Grad-CAM) applied to a COVID 19 Classifier • Proof of concept – work in progress https: //projects. fcgrp. net/covid 19/ • Requires username and password • Secure web server, traffic is encrypted
![References [1] Joseph Paul Cohen, Paul Morrison, Lan Dao (2020). COVID-19 Image Data Collection. References [1] Joseph Paul Cohen, Paul Morrison, Lan Dao (2020). COVID-19 Image Data Collection.](http://slidetodoc.com/presentation_image_h/d1643ee601b27019cf95c2d05848aea1/image-17.jpg)
References [1] Joseph Paul Cohen, Paul Morrison, Lan Dao (2020). COVID-19 Image Data Collection. ar. Xiv: 2003. 11597 [eess. IV], https: //arxiv. org/abs/2003. 11597 [2] Karen Simonyan, Andrew Zisserman (2015). Very Deep Convolutional Networks for Large-Scale Image Recognition. ar. Xiv: 1409. 1556 [cs. CV], https: //arxiv. org/abs/1409. 1556 [3] Ioannis D. Apostolopoulos, Tzani Bessiana (2020). Covid-19: Automatic detection from X-Ray images utilizing Transfer Learning with Convolutional Neural Networks. ar. Xiv: 2003. 11617 [eess. IV], https: //arxiv. org/abs/2003. 11617 [4] Ramprasaath R. Selvaraju et al (2019). Grad-CAM: Visual Explanations from Deep Networks via Gradient-based Localization (2019), ar. Xiv: 1610. 02391 [cs. CV], https: //arxiv. org/abs/1610. 02391 [5] Li et al (2020). COVID-Mobile. Xpert: On-Device COVID-19 Screening using Snapshots of Chest X-Ray (2020), ar. Xiv: 2004. 03042 [eess. IV], https: // ar. Xiv: 2004. 03042 v 2
- Slides: 17